Summary
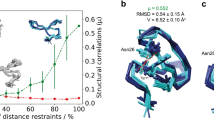
We have systematically examined how the quality of NMR protein structures depends on (1) the number of NOE distance constraints. (2) their assumed precision, (3) the method of structure calculation and (4) the size of the protein. The test sets of distance constraints have been derived from the crystal structures of crambin (5 kDa) and staphylococcal nuclease (17 kDa). Three methods of structure calculation have been compared: Distance Geometry (DGEOM), Restrained Molecular Dynamics (XPLOR) and the Double Iterated Kalman Filter (DIKF). All three methods can reproduce the general features of the starting structure under all conditions tested. In many instances the apparent precision of the calculated structure (as measured by the RMS dispersion from the average) is greater than its accuracy (as measured by the RMS deviation of the average structure from the starting crystal structure). The global RMS deviations from the reference structures decrease exponentially as the number of constraints is increased, and after using about 30% of all potential constraints, the crrors asymptotically approach a limiting value. Increasing the assumed precision of the constraints has the same qualitative effect as increasing the number of constraints. For comparable numbers of constraints/residue, the precision of the calculated structure is less for the larger than for the smaller protein, regardless of the method of calculation. The accuracy of the average structure calculated by Restrained Molecular Dynamics is greater than that of structures obtained by purely geometric methods (DGEOM and DIKF).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Altman, R.B. and Jardetzky, O. (1986)J. Biochem.,100, 1403–1423.
Altman, R.B. and Jardetzky, O. (1989)Methods Enzymol.,177, 218–246.
Altman, R.B., Pachter, R. and Jardetzky, O. (1989) InProtein Structure and Engineering (Ed, Jardetzky, O.) Plenum Press, New York, pp. 79–95.
Altman, R.B., Pachter, R. Carrara, E.A. and Jardetzky, O. (1990)QCPE Bull.,10 (4), Program 596.
Altman, R.B., Pachter, R. and Jardetzky, O. (1992)J. Appl. Magn. Reson., in press.
Arrowsmith, C.H., Czaplicki, J., Iyer, S.B. and Jardetzky, O. (1991a)J. Am. Chem. Soc.,113, 4020–4022.
Arrowsmith, C., Pachter, R., Altman, R. and Jardetzky, O. (1991b)Eur. J. Biochem.,202(2), 53–66.
Blaney, J.M., Crippen, G.M., Dearing, A. and Dixon, J.S. (1990)QCPE, Program 590, Indiana University, Bloomington, IN 47405.
Braun, W. and Gō, N., (1985)J. Mol. Biol. 186, 611–626.
Brünger, A.T. (1990)XPLOR Manual, Yale University, New Haven, CT.
Brünger, A.T., Clore, G.M., Gronenborn, A.M. and Karplus, M. (1986)Proc. Natl. Acad. Sci. USA 83, 3801–3805.
Bystrov, V.F. (1976)Prog. NMR Spectrosc.,10, 41–81.
Carrara, E.A., Brinkley, J.F., Cornelius, C.C., Altman, R.B., Brugge J., Pachter, R., Buchanan, B. and Jardetzky, O. (1990)QCPE Bull. 10(4), Program 596.
Clore, G.M. and Gronenborn, A.M. (1989)J. Magn. Reson.,84, 398–409.
Clore, G.M., Nilges, M., Brunger, A.T., Karplus, M. and Gronenborn, A.M. (1987)FEBS Lett.,213, 269.
Gronenborn, A.G., Wingfield, P.T. and Clore, G.M. (1989)Biochemistry. 28, 5081.
Hendrichson, W.A. and Teeter, M.A. (1981)Nature,290, 107.
Hua, Q.X., Shoelson, S.E., Kochoyan, M. and Weiss, M.A. (1991),Nature 354, 238–241.
Jardetzky, O. (1991), InComputational Aspects of the Study of Biological Macromolecules by NMR Spectroscopy (Eds. Hoch, J.C., Poulson, F.M. and Redfield, C.) Plenum Publ. Corp., New York, pp. 375–390.
Jardetzky, O. and Roberts G.C.K. (1981) InNMR in Molecular Biology, Academic Press, New York, NY, Chapter 5.
Jardetzky, O. and Lane, A.N. (1988) InPhysics of NMR Spectroscopy in Biology and Medicine (Ed. Maraviglia, B.) Ital. Physical Society, North-Holland Physics Publishing, Amsterdam, pp. 267–300.
Keepers, J.W. and James, T. (1984).J. Magn. Reson.,57, 404–426.
Koehl, P., Lefèvre, J.-F. and Jardetzky, O. (1992)J. Mol. Biol. 223, 299–315.
Kuntz, I.D., Crippen, G.M. and Kollman, P.A. (1979)Biopolymers,18, 939–957.
Kuntz, I.D., Thomason, J.F. and Oshiro, C.M. (1989)Methods Enzymol.,177, 159–204.
Lefèvre, J.-F., Lane, A.N. and Jardetzky, O. (1987)Biochemistry,26, 5067–5090.
Lichtarge, O., Jardetzky, O. and Li, C.H. (1987)Biochemistry,26, 5916–5925.
Loll, P.J. and Lattman, E.E. (1989)Proteins: Structure, Function, and Genetics,5, 183–201.
Madrid, M. and Jardetzky, O. (1988)Biochim. Biophys. Acta,953, 61–69.
Madrid, M., Mace, J.E. and Jardetzky, O. (1989)J. Magn. Reson.,83, 267–278.
Metzler, W.J., Hare, D. and Pardi, A. (1989)Biochemistry,28, 7045–7052.
Nilges, M., Clore, G.M. and Gronenborn, A.M. (1988)FEBS Lett. 229, 317–324.
Pachter, R., Altman, R.B. and Jardetzky, O. (1990)J. Magn. Reson.,89, 578–584.
Pardi, A., Billeter, M. and Wüthrich, K. (1984)J. Mol. Biol.,181, 741.
Scheek, R.M., van Gunsteren, W.F. and Kaptein, R. (1989)Methods Enzymol.,177, 204–218.
Van Gunsteren, W.F., Kaptein, R. and Zuiderweg, E.R.P. (1983) InNucleic Acid Conformation and Dynamics (Ed, Olson, W.K.) Report of the NATO/CECAM Workshop, Orsay.
Wüthrich, K. (1986)NMR of Proteins and Nucleic Acids, Wiley, New York.
Wüthrich, K., Billeter, M. and Braun, W. (1983)J. Mol. Biol. 169, 949–961.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Liu, Y., Zhao, D., Altman, R. et al. A systematic comparison of three structure determination methods from NMR data: Dependence upon quality and quantity of data. J Biomol NMR 2, 373–388 (1992). https://doi.org/10.1007/BF01874815
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01874815