Abstract
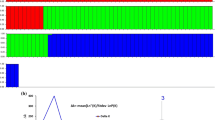
Genetic variability of cultivated and wild barley, Hordeum vulgare ssp. vulgare and spontaneum, respectively, was assessed by RFLP analysis. The material consisted of 13 European varietes, single-plant offspring lines of eight land races from Ethiopia and Nepal, and five accessions of ssp. spontaneum from Israel, Iran and Turkey. Seventeen out of twenty-one studied cDNA and gDNA probes distributed across all seven barley chromosomes revealed polymorphism when DNA was digested with one of four restriction enzymes. A tree based on genetic distances using frequencies of RFLP banding patterns was estimated and the barley lines clustered into five groups reflecting geographical origin. The geographical groups of land-race lines showed less intragroup variation than the geographical groups of spontaneum lines. The group of European varieties, representing large variation in agronomic traits, showed an intermediate level. The proportion of gene diversity residing among geographical groups (FST) varied from 0.19 to 0.94 (average 0.54) per RFLP pattern, indicating large diversification between geographical groups.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Allard RW (1988) Genetic changes associated with the evolution of adaptedness in cultivated plants and their wild progenitors. J Hered 79:225–238
Allard RW, Saghai Maroof MA, Zhang Q, Jørgensen RA (1990) Genetic and molecular organization of ribosomal DNA (rDNA) variants in wild and cultivated barley. Genetics 126:743–751
Brown AHD, Munday J (1982) Population-genetic structure and optimal sampling of land races of barley from Iran. Genetica 58:85–96
Christiansen SK, Giese H (1990) Genetic analysis of the obligate parasitic barley powdery mildew fungus based on RFLP and virulence loci. Theor Appl Genet 79:705–712
Doll H, Brown AHD (1979) Hordein variation in wild (Hordeum spontaneum) and cultivated (H. vulgare) barley. Can J Genet Cytol 21:391–404
Feinberg A, Vogelstein B (1983) A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) version 3.5c. Distributed by the author. Department of Genetics, University of Washington
Fitch WM, Margoliash E (1967) Construction of phylogenetic trees. Science 155:279–284
Gepts P, Clegg MT (1989) Genetic diversity in pearl millet [Pennisetum glaucum (L.) R. Br.]at the DNA sequence level. Heredity 80:203–208
Giese H, Holm-Jensen AG, Jensen HP, Jensen J (1993) Localization of the Laevigatum powdery mildew resistance gene to barley chromosome 2 by the use of RFLP markers. Theor Appl Genet 85:897–900
Giese H, Holm-Jensen AG, Mathiassen H, Kjær B, Rasmussen SK, Bay H, Jensen J (1994) Distribution of RAPD markers on a linkage map of barley. Hereditas 120:267–273
Graner A, Siedler H, Jahoor A, Herrmann RB, Wenzel G (1990) Assessment of the degree and the type of restriction fragment length polymorphism in barley (Hordeum vulgare). Theor Appl Genet 80:826–832
Hartl DL, Clark AG (1989) Principles of population genetics, 2nd edn. Sinauer Associates, Sunderland, Massachusetts, USA
Harlan JR (1979) On the origin of barley. In: Barley: origin, botany, culture, utilization, pests. Agriculture Handbook 338, US Department of Agriculture, pp 10–36
Jana S, Pietrzak LN (1988) Comparative assessment of genetic diversity in wild and primitive cultivated barley in a center of diversity. Genetics 119:981–990
Maniatis T, Fritsch EF, Sambrook I (1982) Molecular cloning: a laboratory manual. Cold Spring Harbour Laboratory, Cold Spring Harbour, New York
Nevo E, Zohary D, Brown AHD, Haber M (1979) Genetic diversity and environmental associations of wild barley, Hordeum spontaneum, in Israel. Evolution 33:815–833
Nevo E, Beiles A, Zohary D (1986) Genetic resources of wild barley in the Near East: structure, evolution and application in breeding. Biol J Linn Soc 27:355–380
Saghai Maroof MA, Allard RW, Zhang Q (1990) Genetic diversity and ecogeographical differentiation among ribosomal DNA alleles in wild and cultivated barley. Proc Natl Acad Sci USA 87:8486–8490
Sharp PJ, Kreis M, Shewry PR, Gale MD (1988) Location of α-amylase sequences in wheat and its relatives. Theor Appl Genet 75:286–290
Sharp PJ, Chao S, Desai S, Gale MD (1989) The isolation, characterization and application in the Triticeae of a set of wheat RFLP probes identifying each homoeologous chromosome arm. Theor Appl Genet 78:342–348
Shin JS, Chao S, Corpuz L, Blake T (1990) A partial map of the barley genome incorporating restriction fragment length polymorphism, polymerase chain reaction, isozyme, and morphological marker loci. Genome 33:803–810
Zhang Q, Saghai Maroof MA, Kleinhofs A (1993) Comparative diversity analysis of RFLPs and isozymes within and among populations of Hordeum vulgare ssp. spontaneum. Genetics 134:909–916
Author information
Authors and Affiliations
Additional information
Communicated by P. M. A. Tigerstedt
Rights and permissions
About this article
Cite this article
Petersen, L., Østergård, H. & Giese, H. Genetic diversity among wild and cultivated barley as revealed by RFLP. Theoret. Appl. Genetics 89, 676–681 (1994). https://doi.org/10.1007/BF00223704
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00223704