Summary
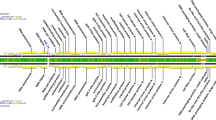
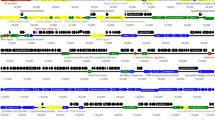
The nucleotide sequence of the circular single-stranded genome of the filamentous Escherichia coli phage I2-2 has been determined and compared with those of the filamentous E. coli phages Ff(M13, fl, or fd) and IKe. The I2-2 DNA sequence comprises 6744 nucleotides; 139 nucleotides less than that of the N- and I2-plasmid-specific phage IKe, and 337 (336) nucleotides more than that of the F-plasmid-specific phage Ff. Nucleotide sequence comparisons have indicated that I2-2, IKe, and Ff have a similar genetic organization, and that the genomes of I2-2 and IKe are evolutionarily more closely related than those of I2-2 and Ff. The studies have further demonstrated that the I2-2 genome is a composite replicon, composed of only two-thirds of the ancestral genome of IKe. Only a contiguous I2-2 DNA sequence of 4615 nucleotides encompassing not only the coat protein and phage assembly genes, but also the signal required for efficient phage morphogenesis, was found to be significantly homologous to sequences in the genomes of IKe and Ff. No homology was observed between the consecutive DNA sequence that contains the origins for viral and complementary strand replication and the replication genes. Although other explanations cannot be ruled out, our data strongly suggest that the ancestor filamentous phage genome of phages I2-2 and IKe has exchanged its replication module during evolution with that of another replicon, e.g., a plasmid that also replicates via the so-called rolling circle mechanism.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Beck E, Zink B (1981) Nucleotide sequence and genome organisation of filamentous bacteriophages fl and fd. Gene 16:35–58
Beck E, Sommer R, Auerswald EA, Kurz C, Zink B, Osterburg G, Schaller H (1978) Nucleotide sequence of bacteriophage fd DNA. Nucleic Acids Res 5:4495–4503
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Bradley DE, Sirgel FA, Coetzee JN, Hedges RW, Coetzee WF (1982) Phages C-2 and J: IncC and Incl plasmid-dependent phages, respectively. J Gen Microbiol 128:2485–2498
Bradley DE, Coetzee JN, Hedges RW (1983) IncI2 plasmids specify sensitivity to filamentous bacteriophage IKe. J Bacteriol 154:505–507
Caro LG, Schnös M (1966) The attachment of the male-specific bacteriophage fl to sensitive strains of Escherichia coli. Proc Natl Acad Sci USA 56:126–132
Cleary JM, Ray DS (1980) Replication of the plasmid pBR322 under the control of a cloned replication origin from the single-stranded DNA phage M13. Proc Natl Acad Sci USA 77:4638–4642
Cleary JM, Ray DS (1981) Deletion analysis of the cloned replication origin region from bacteriophage M13. J Virol 40:197–203
Coetzee JN, Bradley DE, Hedges RW (1982) Phages Iα and I2–2: IncI plasmid-dependent bacteriophages. J Gen Microbiol 128:2797–2804
Dotto GP, Zinder ND (1983) The morphogenetic signal of bacteriophage fl. Virology 130:252–256
Dotto GP, Enea V, Zinder ND (1981) Functional analysis of bacteriophage fl intergenic region. Virology 114:463–473
Dotto GP, Horiuchi K, Zinder ND (1982) Initiation and termination of phage fl plus-strand synthesis. Proc Natl Acad Sci USA 79:7122–7126
Edens L, Konings RNH, Schoenmakers JGG (1978) A cascade mechanism of transcription in bacteriophage M13 DNA. Virology 86:354–367
Enea V, Zinder ND (1982) Interference resistant mutants of phage fl. Virology 122:222–226
Grant RB, Whiteley MH, Shapley AJ (1978) Plasmids of incompatibility group P code for the capacity to propagate bacteriophage IKe. J Bacteriol 136:808–811
Gray CP, Sommer R, Polke C, Beck E, Schaller H (1978) Structure of the origin of DNA replication of bacteriophage fd. Proc Natl Acad Sci USA 75:50–53
Hill DF, Petersen GB (1982) Nucleotide sequence of bacteriophage fl DNA. J Virol 44:32–46
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Khatoon H, Iyer RV, Iyer VN (1972) A new filamentous bacteriophage with sex-factor specificity. Virology 48:145–155
Konings RNH, Luiten RGM, Peelers BPH (1986) Mike, a chimeric filamentous phage designed for the separate production of either DNA strand of pKUN vector plasmids by F+ cells. Gene 46:269–276
Konings, RNH, Verhoeven EJM, Peeters BPH (1987) pKUN, vectors for the separate production of both DNA strands of recombinant plasmids. Methods Enzymol 153:12–34
Marvin DA, Folkhard W (1986) Structure of F-pili: reassessment of the symmetry. J Mol Biol 191:299–300
Meyer TF, Geider K, Kurz C, Schaller H (1979) Cleavage site of bacteriophage fd gene II-protein in the origin of viral strand replication. Nature 278:365–367
Miyata T, Yasunaga T (1980) Molecular evolution of mRNA: a method for estimating evolutionary rates of synonymous and amino acid substitutions from homologous nucleotide sequences and its application. J Mol Evol 16:23–36
Model P, Russel M (1988) Filamentous bacteriophage. In: Calendar R (ed) The bacteriophages, vol 2. Plenum, New York, pp 375–456
Moses PB, Model P (1984) A rho-dependent transcription termination signal in bacteriophage fl. J Mol Biol 172:1–22
Peeters BPH, Peters RM, Schoenmakers JGG, Konings RNH (1985) Nucleotide sequence and genetic organization of the genome of the N-specific filamentous bacteriophage IKe: comparison with the genome of the F-specific filamentous phages M13, fd and fl. J Mol Biol 181:27–39
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor University Press, Cold Spring Harbor NY
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schaller H, Uhlmann A, Geider K (1976) A DNA fragment from the origin of single-strand to double-strand DNA replication of bacteriophage fd. Proc Natl Acad Sci USA 73:49–53
Shine J, Dalgarno L (1974) The 3′-terminal sequence of Escherichia coli 16S ribosomal RNA: complementary to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci USA 71:1342–1346
Smits MA, Jansen J, Konings RNH, Schoenmakers JGG (1984) Initiation and termination signals for transcription in bacteriophage M13. Nucleic Acids Res 12:4071–4081
Staden R (1980) A new computer method for the storage and manipulation of DNA gel reading data. Nucleic Acids Res 8:3673–3694
Staden R (1982) An interactive graphics program for comparing and aligning nucleic acid and amino acid sequences. Nucleic Acids Res 10:2951–2961
Stanisich VA (1974) The properties and host range of male-specific bacteriophages of Pseudomonas aeruginosa. J Gen Microbiol 84:332–342
Van Wezenbeek PMGF, Hulsebos TJM, Schoenmakers JGG (1980) Nucleotide sequence of the filamentous bacteriophage M13 DNA genome: comparison with phage fd. Gene 11:129–148
Watanabe T, Nishida H, Ogata C, Arai T, Sato S (1964) Episome-mediated transfer of drug resistance in Enterobacteriaceae: VII. Two types of naturally occurring R factors. J Bacteriol 88:716–726
Wilbur WJ, Lipman DJ (1983) Rapid similarity searches of nucleic acid and protein data banks. Proc Natl Acad Sci USA 80:726–730
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Zinder ND, Horiuchi K (1985) Multiregulatory element of filamentous bacteriophages. Microbiol Rev 49:101–106
Author information
Authors and Affiliations
Additional information
Offprint requests to: R.N.H. Konings
Rights and permissions
About this article
Cite this article
Stassen, A.P.M., Schoenmakers, E.F.P.M., Yu, M. et al. Nucleotide sequence of the genome of the filamentous bacteriophage I2-2: Module evolution of the filamentous phage genome. J Mol Evol 34, 141–152 (1992). https://doi.org/10.1007/BF00182391
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00182391