Abstract
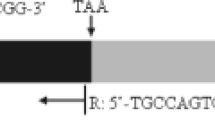
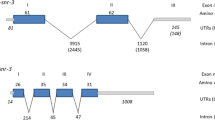
The region of the genome of D. lebanonensis that contains the Adh gene and the downstream Adh-dup gene was sequenced. The structure of the two genes is the same as has been described for D. melanogaster. Adh has two promoters and Adh-dup has only one putative promoter. The levels of expression of the two genes in this species are dramatically different. Hybridizing the same Northern blots with a specific probe for Adh-dup, we did not find transcripts for this gene in D. lebanonensis. The level of Adh distal transcript in adults of D. lebanonensis is five times greater than that of D. melanogaster adults. The maximum levels of proximal transcript are attained at different larval stages in the two species, being three times higher in D. melanogaster late-second-instar larvae than in D. lebanonensis first-instar larvae. The level of Adh transcripts allowed us to determine distal and proximal initiation transcription sites, the position of the first intron, the use of two polyadenylation signals, and the heterogeneity of polyadenylation sites. Temporal and spatial expression profiles of the Adh gene of D. lebanonensis show qualitative differences compared with D. melanogaster. Adh and Adh-dup evolve differently as shown by the synonymous and nonsynonymous substitution rates for the coding region of both genes when compared across two species of the melanogaster group, two of the obscura group of the subgenus Sophophora and D. lebanonensis of the victoria group of the subgenus Scaptodrsophila. Synonymous rates for Adh are approximately half those for Adh-dup, while nonsynonymous rates for Adh are generally higher than those for Adh-dup. Adh shows 76.8% identities at the protein level and 70.2% identities at the nucleotide level while Adh-dup shows 83.7% identities at the protein level and 67.5% identities at the nucleotide level. Codon usage for Adh-dup is shown to be less biased than for Adh, which could explain the higher synonymous rates and the generally lower nonsynonymous substitution rates in Adh-dup compared with Adh. Phylogenetic trees reconstructed by distance matrix and parsimony methods show that Sophophora and Scaptodrosophila subgenera diverged shortly after the separation from the Drosophila subgenus.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Atkinson PW, Mills LE, Starmer WT, Sullivan DT (1988) Structure and evolution of the Adh genes of D. mojavensis. Genetics 120: 713–723
Ayer S, Benyajati C (1990) Conserved enhanced and silencer elements for differential Adh transcription in Drosophila cell lines. Mol Cell Biol 10:3512–3523
Ayer S, Benyajati C (1992) The binding site of a steroid hormone receptor-like protein within the Drosophila Adh adult enhancer is required for high levels of tissue specific alcohol dehydrogenase expression. Mol Cell Biol 12:661–673
Ashburner M (1989) Drosophila. A laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Benyajati C, Spoerel N, Haymerle H, Ashburner M (1983) The messenger RNA for alcohol dehydrogenase in Drosophila melanogaster differs in its 5′ end in different developmental stages. Cell 33:125–133
Benyajati C, Ewel A, McKeon J, Chovav M, Juan E (1992) Characterization and purification of Adh distal promoter factor 2, Adf-2, a cell-specific and promoter-specific repressor in Drosophila. Nucleic Acids Res 20:4481–4489
Brennan MD, Wu CY, Berry AJ (1988) Tissue-specific regulatory differences for the alcohol dehydrogenase genes of Hawaiian Drosophila are conserved in D. melanogaster transformants. Proc Natl Acad Sci USA 85:6866–6869
Brennan MD, Dickinson WJ (1988) Complex developmental regulation of the Drosophila affinidisjuncta alcohol dehydrogenase gene in D. melanogaster. Dev Biol 125:64–74
Bucher P (1990) Weight matrix descriptions of four eukaryotic RNA polymerase II promoter elements derived from 502 unrelated promoter sequences. J Mol Biol 212:563–578
Chirgwin JM, Przybyla AE, Macdonald RJ, Rutter WJ (1970) Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochem J 18:5294–5299
Corbin V, Maniatis T (1989) Role of transcriptional interference in the D. melanogaster Adh promoter switch. Nature 337:279–282
Correa-Rotter R, Mariash CN, Rosenberg ME (1992) Loading and transfer control for Northern hybridization. Biotechniques 12:154–158
David J, Fouillet P, Arens MF (1974) Comparison de la sensibilité ä l'alcool éthilique de six espéces de Drosophila du sous groupe melanogaster. Archs Zool Exp Gén 115:401–410
David JR, Van Herrewege J, Monclus M, Prevosti A (1979) High ethanol tolerance in two distantly related Drosophila species: a probable case of recent convergent evolution. Comp Biochem Physiol 63C:53–56
Dickinson WJ, Rowan RG, Brennan MD (1984) Regulatory gene evolution: adaptive differences in expression of alcohol dehydrogenase in D. melanogaster and D. simulans. Heredity 52:215–225
Fang XM, Wu CY, Brennan MD (1991) Complexity in evolved regulatory variation for alcohol dehydrogenase in Hawaiian Drosophila. J Mol Evol 32:220–226
Fischer JA, Maniatis T (1985) Structure and transcription of the D. mulleri alcohol dehydrogenase genes. Nucleic Acids Res 13:6899–6916
Furia M, Digilio FA, Artiaco D, D'Avino PP, Cavaliere D, Polito LC (1991) Molecular organization of the D. melanogaster Pig-1. J Mol Evol 101:49–54
Felsenstein J (1991) Phylip Version 3.4 University of Washington, Seattle
Geliebter J (1987) Dideoxynucleotide sequencing of RNA and uncloned cDNA. Focus 9.1:5–8
Goldberg DA, Posakony JW, Maniatis T (1983) Correct developmental expression of a cloned alcohol dehydrogenese gene transduced into the Drosophila germ line. Cell 34:59–73
Graves RA, Pandey NB, Chodchoy N, Marzluff WF (1987) Translation is required for regulation of histone mRNA degradation. Cell 48:615–626
Heberlein U, England B, Tjian R (1985) Characterization of Drosophila transcription factors that activate the tandem promoters of the alcohol dehydrogenase gene. Cell 41:965–977
Ikemura T (1985) Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol 2:13–34
Juan E, Gonzalez-Duarte R (1980) Purification and enzyme stability of alcohol dehydrogenase from D. simulans, D. virilis and D. melanogaster Adhs. Biochem J 189:105–110
Juan E, Papaceit M, Quintana A (1990) Nucleotide sequence of the Adh gene of D. lebanonensis. Nucleic Acid Res 18:6420
Kreitman M, Hudson RR (1991) Inferring the evolutionary histories of the Adh and Adh-dup loci in D. melanogaster from patterns of polymorphism and divergence. Genetics 127:565–582
Kimura M (1980) A simple model for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Li WH, Wu CI, Luo CC (1985) A new method for estimating synonymous and nonsynonymous rates of nucleotide substitution considering the relative likelyhood of nucleotide and codon changes. Mol Biol Evol 2:150–174
Marfany G, Gonzalez-Duarte R (1991) The Adh genomic region of D. ambigua: evolutionary trends in different species. J Mol Evol 32:454–462
Moses K, Heberlein U, Ashburner M (1990) The Adh gene promoters of D. melanogaster and D. orena are functionally conserved and share features of sequence structure and nuclease-protected sites. Mol Cell Biol 10:539–548
Okayama H, Kawaichi M, Brownstein M, Lee F, Yokota T, Arai K (1987) High efficiency cloning of full length cDNA. Construction and screening of cDNA expression libraries for mammalian cells. Methods Enzymol 154:3–28
Perkins KK, Dailey GM, Tjian R (1988) In vitro analysis of the Antenapedia P2 promoter: identification of a new Drosophila transcription factor. Genes Dev 2:1615–1625
Quintana A, Juan E (1993) Morphology of mouth hooks and anterior spiracles through the larval development of D. lebanonensis. DIS 72:88
Rowan RG, Dickinson WJ (1988) Nucleotide sequence of the genomic region encoding alcohol dehydrogenase in D. affinidisjuncta. J Mol Evol 28:43–54
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Savakis C, Ashburner M, Willis JH (1986) The expression of the gene coming for alcohol dehydrogenase during the development of D. melanogaster. Dev Biol 114:194–207
Starmer WT, Sullivan DT (1989) A shift in the third-codon-position nucleotide frequency in alcohol dehydrogenease genes in the genus of Drosophila. Mol Biol Evol 6:546–552
Schaeffer SW, Aquadro CF (1987) Nucleotide sequence of the Adh gene region of D. pseudoobscura: evolutionary change and evidence of an ancient gene duplication. Genetics 117:61–73
Sheets MD, Ogg SC, Wickens MP (1990) Point mutations in AAUAAA and the poly A addition site: effects on the accuracy and efficiency of cleavage and polyadenylation in vitro. Nucleic Acid Res 18:5799–5805
Shields DC, Sharp PM, Higgins DG, Wright F (1988) “Silent” sites in Drosophila genes are not neutral: evidence of selection among synonymous codons. Mol Biol Evol 5:704–716
Singh H, Sen R, Baltimore D, Sharp PA (1986) A nuclear factor that binds to a conserved sequence motif in transcriptional control elements of immunoglobulin genes. Nature 319:154–158
Thompson MS, Jacobson JW, Laurie CC (1991) Comparison of alcohol dehydrogenase expression in D. melanogaster and D. simulans. Mol Biol Evol 8:31–48
Villarroya A, Juan E, Egestad B, Jörnvall H (1989) The primary structure of alcohol dehydrogenase from D. lebanonensis. Extensive variation within “short chain” alcohol dehydrogenase lacking zinc. Eur J Biochem 180:191–197
Villarroya A, Juan E (1991) ADH and phylogenetic relationships of D. lebanonensis (Scaptodrosophila). J Mol Evol 32:421–428
Wu CY, Mote J, Brennan M (1990) Tissue-specific expression phenotypes of Hawaiian Drosophila Adh genes in D. melanogaster transformants. Genetics 125:599–610
Author information
Authors and Affiliations
Additional information
Correspondence to: E. Juan
Rights and permissions
About this article
Cite this article
Juan, E., Papaceit, M. & Quintana, A. Nucleotide sequence of the genomic region encompassing Adh and Adh-Dup genes of D. lebanonensis (Scaptodrosophila): Gene expression and evolutionary relationships. J Mol Evol 38, 455–467 (1994). https://doi.org/10.1007/BF00178845
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00178845