Abstract
As sequencing techniques become increasingly efficient, the average length of a sequence is bound to grow. Traditional sequence-comparison algorithms can either compare DNA or protein, but not a mixture, which is actually a common situation. Most obtained DNA sequences contain coding regions, and it is more reliable to compare the coding regions as protein than just as DNA.
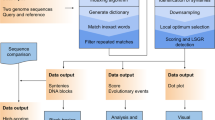
A heuristic algorithm is presented that can compare DNA with both coding and noncoding regions, but that also can compare multiple reading frames and determine which exons are homologous.
A program, GenAl (Genomic Alignment), was developed that implements the algorithm. Its use is demonstrated on two retroviruses.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Doolittle RF (1986) Of URFs and ORFs. University Science Books
Gotoh O (1981) An improved algorithm for matching biological sequences. J Mol Biol 162:705–708
Hein JJ (1994) An algorithm combining DNA and protein alignment. J Theor Biol (in press)
Hein JJ, Støvlbæk J (1993a) A method to analyze aligned genomic DNA sequences. (submitted to J Mol Evol)
Hein JJ, Støvlbæk J (1993b) Multiple genomic alignment. (in preparation)
Hirschberg DS (1975) A linear space algorithm for computing maximal common subsequences. Comm ACM 18(6):341–343
Needlemann SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequences of two proteins. J Mol Biol 48:444–453
Sankoff D (1972) Matching sequences under deletion/insertion constraints. Proc Natl Acad Sci USA 69:4–6
Author information
Authors and Affiliations
Additional information
Correspondence to: J. Hein
Rights and permissions
About this article
Cite this article
Hein, J., Støvlæk, J. Genomic alignment. J Mol Evol 38, 310–316 (1994). https://doi.org/10.1007/BF00176094
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00176094