Abstract
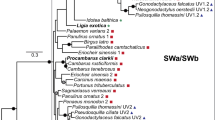
Phylogenetic and physiological methods were used to study the evolution of the opsin gene family in Drosophila. A phylogeny based on DNA sequences from 13 opsin genes including representatives from the two major subgenera of Drosophila shows six major, well-supported clades: The “blue opsin” clade includes all of the Rhl and Rh2 genes and is separated into two distinct subclades of Rhl sequences and Rh2 sequences; the ultraviolet opsin clade includes all Rh3 and Rh4 genes and bifurcates into separate Rh3 and Rh4 clades. The duplications that generated this gene family most likely took place before the evolution of the subgenera Drosophila and Sophophora and their component species groups. Numerous changes have occurred in these genes since the duplications, including the loss and/or gain of introns in the different genes and even within the Rhl and Rh4 clades. Despite these changes, the spectral sensitivity of each of the opsins has remained remarkably fixed in a sample of four species representing two species groups in each of the two subgenera. All of the strains that were investigated had R1-6 (Rhl) spectral sensitivity curves that peaked at or near 480 nm, R7 (Rh3 and Rh4) peaks in the ultraviolet range, and ocellar (Rh2) peaks near 420 nm. Each of the four gene clades on the phylogeny exhibits very conservative patterns of amino acid replacement in domains of the protein thought to influence spectral sen sitivity, reflecting strong constraints on the spectrum of light visible to Drosophila.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Applebury ML, Hargrave PA (1986) Molecular biology of the visual pigments. Vis Res 26:1881–1895
Beverly SM, Wilson AC (1984) Molecular evolution in Drosophila and the higher Diptera II. A time scale for fly evolution. J Mol Evol 21:1–13
Blackman RK, Meselson M (1986) Interspecific nucleotide comparisons used to identify regulatory and structural features of the Drosophila hsp82 gene. J Mol Biol 188:499
Carthew RW, Rubin GM (1990) Seven in absentia, a gene required for specification of R7 cell fate in the Drosophila eye. Cell 63:561–577
Carulli JP, Hard DL (1992) Variable rates of evolution among Drosophila opsin genes. Genetics 132:193–204
Chan T, Lee M, Sakmar TP (1992) Introduction of hydroxyl-bearing amino acids causes bathochromic shifts in rhodopsin. J Biol Chem 276:9478–9480
Colot HV, Hall JC, Rosbash M (1988) Interspecific comparison of the period gene of Drosophila reveals large blocks of non-conserved coding DNA. EMBO J 17:3929–3987
Cowman AF, Zuker CS, Rubin GM (1986) A rhodopsin expressed in only one photoreceptor cell type of the Drosophila eye. Cell 44:705–710
DeSalle R (1992) The phylogenetic relationships of flies in the family Drosophilidae deduced from mtDNA sequences. Mol Phylogenet Evol 1:31–40
Devereux J, Haeberli P, Smithies O (1984) A comprehensive set of sequence programs for the VAX. Nucleic Acids Res 12:387–395
Feiler R, Harris WA, Kirschfeld K, Werhan C, Zuker CS (1988) Targeted misexpression of a Drosophila opsin gene leads to altered visual function. Nature 333:737–741
Feiler R, Bjornson R, Kirschfeld K, Mismer D, Rubin GM, Smith DP, Socolich M, Zuker CS (1992) Ectopic expression of ultravioletrhodopsins in the blue photoreceptor cells of Drosophila: visual physiology and photochemistry of transgenic animals. J Neurosci 12:3862–3868
Fortini ME, Rubin GM (1990) Analysis of cis-acting requirements of the Rh3 and Rh4 genes reveals a bipartite organization to rhodopsin promoters in Drosophila melanogaster. Genes Dev 4:444–463
Fryxell KJ, Meyerowitz EM (1987) An opsin gene that is expressed only in the R7 photoreceptor cell of Drosophila. EMBO J 6:443–451
Grimaldi D (1990) A phylogenetic, revised classification of the Drosophilidae (Diptera). Bull Am Mus Nat Hist 197:1–139
Hall MD, Hoon MA, Ryba NJP, Pottinger JDD, Keen JN, Saibil HR, Findlay JBC (1991) Molecular cloning and primary structure of squid (Loligo forbesi) rhodopsin, a phospholipase C-directed G-protein-linked receptor. Biochem J 274:35–40
Hardie RC (1985) Functional organization of the fly retina. In: Autrum H, Ottoson D, Perl ER, Schmidt RF, Shimazu H, Willis WD (eds) Progress in sensory physiology 5. Springer-Verlag, Berlin, pp 2–79
Harris WA, Stark WS, Walker JA (1976) Genetic dissection of the photoreceptor system in the compound eye of Drosophila melanogaster. J Physiol 256:415–439
Higgins DG, Sharp PM (1988) Clustal: a package for performing multiple sequence alignments on a microcomputer. Gene 73:237–244
Hu KG, Reichert H, Stark WS (1978) Electrophysiological characterization of Drosophila ocelli. J Comp Physiol 126:15–24
Hubbard R, Sperling L (1973) The colors of the visual pigment chromophores. Exp Eye Res 17:581–589
Huber A, Smith DP, Zuker CS, Paulsen R (1990) Opsin of Calliphora peripheral photoreceptors R1–6: homology with Drosophila (Rhl) and post-translational processing. J Biol Chem 265:17906–17910
Kirschfeld K, Francheschini N, Minke B (1977) Evidence for a sensitizing pigment in fly photoreceptors. Nature 269:386–390
Montell C, Jones K, Zuker CS, Rubin GM (1987) A second opsin gene expressed in the ultraviolet sensitive R7 photoreceptor cells of Drosophila melanogaster. J Neurosci 17:1558–1566
Nathans J (1987) Molecular biology of visual pigments. Ann Rev Neurosci 10:163–194
Nathans J (1990a) Determinants of visual pigment absorbance: identification of the retinylidine Schiff's base counterion in bovine Rhodopsin. Biochemistry 29:9746–9752
Nathans J (1990b) Determinants of visual pigment absorbance: role of charged amino acids in the putative transmembrane segments. Biochemistry 29:937–942
Neitz M, Neitz J, Jacobs GH (1991) Spectral tuning of pigments underlying red-green color vision. Science 252:971–974
Neufeld TP, Carthew RW, Rubin GM (1991) Evolution of gene position: chromosomal arrangement and sequence comparison of the Drosophila melanogaster and Drosophila virilis sina and Rh4 genes. Proc Natl Acad Sci USA 88:10203–10207
O'Neil MT, Belote JM (1992) Interspecific comparison of the transformer gene of Drosophila reveals an unusually high degree of evolutionary divergence. Genetics 131:113–128
O'Tousa JE, Baehr E, Martin RL, Hirsch J, Pak WL, Applebury ML (1985) The Drosophila ninaE gene encodes an opsin. Cell 40:839–850
Ovehinnikov VA, Abdulev NG, Zolotrarev AS, Artomonov ID, Bespalov IA, Dergachev AE, Tsuda M (1988) Octopus rhodopsin-amino acid sequence deduced from cDNA. FEBS Lett 232:69–72
Pak WL, Grabowski SR (1978) Physiology of the visual and flight systems. In: Ashburner M, and Wright TRF (eds) The genetics and biology of Drosophila, vol 3a. Academic Press, New York, pp 553–604
Pollock JA, Benzer S (1988) Transcript localization of four opsin genes in the three visual organs of Drosophila. Nature 333:779–782
Sakmar TP, Franke RR, Khorana HG (1991) The role of the retinylidene Schiff base counterion in rhodopsin in determining wavelength absorbance and Schiff base pKa. Proc Natl Acad Sci USA 88:3079–3083
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Sanderson MJ, Doyle JJ (1992) Reconstruction of organismal and gene phylogenies from data on multigene families: concerted evolution, homoplasy, and confidence. Syst Biol 41:4–17
Smith DP, Stamnes MA, Zuker CS (1991) Signal transduction in the visual system of Drosophila. Ann Rev Cell Biol 7:161–190
Stark WS (1975) Spectral selectivity of visual response alterations mediated by interconversions of native and intermediate photopigments in Drosophila. J Comp Physiol 96:343–356
Stark WS (1977) Sensitivity and adaptation in R7, an ultraviolet photoreceptor, in the Drosophila retina. J Comp Physiol 11547–59
Stark WS, Ivanyshyn AM, Greenburg RM (1976) Spectral sensitivities and photopigments in adaptation of fly visual receptors. Naturwissenschaften 63:513–518
Stark WS, Ivanyshyn AM, Greenburg RM (1977) Sensitivity and photopigments of R1–6, a two-peaked photoreceptor, in Drosophila, Calliphora and Musca. J Comp Physiol 121:289–305
Stark WS, Johnson MA (1980) Microspectrophotometry of Drosophila visual pigments: determinations of conversion efficiency in R1–6 receptors. J Comp Physiol 140:275–286
Stark WS, Sapp R, Schilly D (1988) Rhabdomere turnover and rhodopsin cycle: maintenance of retinula cells in D. melanogaster. J Neurocytol 17:499–509
Stark WS, Zitsmann (1976) Isolation of adaptation mechanisms and photopigment spectra by vitamin A deprivation in Drosophila. J Comp Physiol 105:15–27
Stryer L (1986) Cyclic GMP cascade of vision. Ann Rev Neurosci 9:87–119
Swofford D (1992) Phylogenetic analysis using parsimony, version 3.0r. University of Illinois Natural History Survey, Champaign, IL
Winderickx J, Lindsey DT, Sanock E, Teller DY, Motulsky AG, Deeb SS (1992) Polymorphism in red photopigment underlies variation in color matching. Nature 356:431–435
Zhukovsky EA, Oprian DD (1989) Effect of carboxylic acid side chains on the absorption maximum of visual pigments. Science 246:928–930
Zuker CS, Cowman AF, Rubin GM (1985) Isolation and structure of a rhodopsin gene from Drosophila. Cell 40:851–858
Zuker CS, Montell C, Jones K, Laverty T, Rubin GM (1987) A rhodopsin gene expressed in photoreceptor cell R7 of the Drosophila eye: homologies with other signal transducing molecules. J Neurosci 7:1550–1557
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Carulli, J.P., Chen, DM., Stark, W.S. et al. Phylogeny and physiology of Drosophila opsins. J Mol Evol 38, 250–262 (1994). https://doi.org/10.1007/BF00176087
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00176087