Abstract
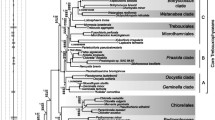
Complete nuclear-encoded small-subunit 18S rRNA (=SSU rRNA) gene sequences were determined for the prasinophyte green alga Mantoniella squamata; the charophycean green algae Chara foetida, Coleochaete scutata, Klebsormidium flaccidum, and Mougeotia scalaris; the bryophytes Marchantia polymorpha, Fossombronia pusilla, and Funaria hygrometrica; and the lycopod Selaginella galleottii to get a better insight into the sequential evolution from green algae to land plants. The sequences were aligned with several previously published SSU rRNA sequences from chlorophytic and charophytic algae as well as from land plants to infer the evolutionary relationships for major evolutionary lineages within the Chlorobionta by distance matrix, maximum parsimony, and maximum likelihood analyses. Phylogenetic trees created by the different methods consistently placed the Charophyceae on the branch leading to the land plants. The Charophyceae were shown to be polyphyletic with the Charales (“charalean” algae) diverging earlier than the Coleochaetales, Klebsormidiales, Chlorokybales, and Zygnematales (“charophycean” algae) which branch from a point closer to the land plants in most analyses. Maximum parsimony and maximum likelihood analyses imply a successive evolution from “charophycean” algae, particularly Coleochaetales, to bryophytes, lycopods, and seed plants. In contrast, distance matrix methods group the bryophytes together with the “charophycean” algae, suggesting a separate evolution of these organisms compared with the club moss and the seed plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Baldauf SL, Palmer JD (1990) Evolutionary transfer of the chloroplast tufA gene to the nucleus. Nature 344:262–265
Baldauf SL, Manhart JR, Palmer JD (1990) Different fates of the chloroplast tufA gene following its transfer to the nucleus in green algae. Proc Natl Acad Sci USA 87:5317–5321
Behn W, Herrmann RG (1977) Circular molecules in the β-satellite DNA of Chlamydomonas reinhardtii. Mol Gen Genet 157:25–30
Bold HC, Wynne MJ (1985) Introduction to the algae. Prentice-Hall, Englewood Cliffs, NJ
Bopp M, Knoop B (1984) Culture methods for bryophytes. In: Vasil I (ed) Cell culture and somatic cell genetics of plants. Academic Press, New York, pp 96–106
Camin JH, Sokal RR (1965) A method for deducing branching sequences in phylogeny. Evolution 19:311–326
Capesius I (1994) New information about the phylogeny of bryophytes from the nuclear encoded 18S rRNA genes. J Plant Physiol (submitted)
Cavalier-Smith T (1993) Kingdom Protozoa and its 18 phyla. Microbiol Rev 57:953–994
Cech TR (1988) Conserved sequences and structures of group I introns: building an active site for RNA catalysis—a review. Gene 73:259–271
Chan YL, Gutell R, Noller HF, Wool IG (1984) The nucleotide sequence of a rat 18S ribosomal ribonucleic acid gene and a proposal for the secondary structure of 18S ribosomal ribonucleic acid. J Biol Chem 259:224–230
Chapman RL, Buchheim MA (1991) Ribosomal RNA sequences: analysis and significance in the phylogeny and taxonomy of green algae. Crit Rev Plant Sci 10:343–368
Chapman RL, Buchheim MA (1992) Green algae and the evolution of land plants: inferences from nuclear-encoded rRNA gene sequences. Biosystems 28:127–137
De Jesus M, Tabatabai F, Chapman DJ (1989) Taxonomic distribution of copper-zinc superoxide dismutase in green algae and its phylogenetic importance. J Phycol 25:767–772
Delwiche CF, Graham LE, Thomson N (1989) Lignin-like compounds and sporopollenin in Coleochaete, an algal model for land plant ancestry. Science 245:399–401
Devereux R, Loeblich III AR, Fox GE (1990) Higher plant origins and the phylogeny of green algae. J Mol Evol 31:18–24
De Wachter R, Neefs J-M, Goris A, Van de Peer Y (1992) The gene coding for small ribosomal RNA in the basidiomycete Ustilago maydis contains a group I intron. Nucleic Acids Res 20:1251–1257
Domozych DS, Stewart KD, Mattox KR (1980) The comparative aspects of cell wall chemistry in the green algae (Chlorophyta). J Mol Evol 15:1–12
Eckenrode VK, Arnold J, Meagher RB (1985) Comparison of the nucleotide sequence of soybean 18S rRNA with the sequences of other small-subunit rRNAs. J Mol Evol 21:259–269
Elwood HJ, Olsen GJ, Sogin ML (1985) The small subunit ribosomal RNA gene sequences from the hypotrichous ciliates Oxytricha nova and Stylonychia pustulata. Mol Biol Evol 2:399–410
Felsenstein J (1978) Cases in which parsimony and compatibility methods will be positively misleading. Syst Zool 27:401–410
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Felsenstein J (1988) Phylogenies from molecular sequences: inference and reliability. Annu Rev Genetics 22:521–565
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package); Version 3.5c. University of Washington, Seattle
Fitch WM, Margoliash E (1967) Construction of phylogenetic trees. Science 155:279–284
Gonzalez IL, Schmickel RD (1986) The human 18S ribosomal RNA gene: Evolution and stability. Am J Hum Genet 38:419–427
Graham LE (1993) Origin of land plants. John Wiley & Sons, New York
Graham LE, Kaneko Y (1991) Subcellular structures of relevance to the origin of the land plants (embryophytes) from green algae. Crit Rev Plant Sci 10:323–342
Graham LE, Wilcox LW (1983) The occurrence and phylogenetic significance of putative placental transfer cells in the green alga Coleochaete. Am J Bot 70:113–120
Graham LE, Delwiche CF, Mishler BD (1991) Phylogenetic connections between the “green algae” and the “bryophytes.” Adv Bryol 4:213–244
Gunderson JH, Elwood H, Ingold A, Kindle K, Sogin ML (1987) Phylogenetic relationships between chlorophytes, chrysophytes, and oomycetes. Proc Natl Acad Sci USA 84:5823–5827
Halanych KM (1991) 5S ribosomal RNA sequences inappropriate for phylogenetic reconstruction. Mol Biol Evol 8:249–253
Hendriks L, Van de Peer Y, Van Herck M, Neefs J-M, De Wachter R (1990) The 18S ribosomal RNA sequence of the sea anemone Anemonia sulcata and its evolutionary position among other eukaryotes. FEBS Lett 269:445–449
Hori H, Osawa S (1987) Origin and evolution of organisms as deduced from 5S ribosomal RNA sequences. Mol Biol Evol 4:445–472
Hoshaw RW, McCourt RM, Wang JC (1990) Phylum Conjugaphyta. In: Margolis L, Corliss JO, Melkonian M, Chapman DJ (eds) Handbook of Protoctista, Jones and Barlett, Boston, pp 119–131
Hotchkiss Jr AT, Gretz MR, Hicks KB, Brown Jr RM (1989) The composition and phylogenetic significance of the Mougeotia (Charophyceae) cell wall. J Phycol 25:646–654
Hultman T, Bergh S, Moks T, Uhlen M (1991) Bidirectional solid-phase sequencing of in vitro-amplified plasmid DNA. Biotechniques 10:84–93
Huss VAR, Sogin ML (1989) Primary structure of the Chlorella vulgaris small subunit ribosomal RNA coding region. Nucleic Acids Res 17:1255
Huss VAR, Sogin ML (1990) Phylogenetic position of some Chlorella species within the Chlorococcales based upon complete small-subunit ribosomal RNA sequences. J Mol Evol 31:432–442
Huss VAR, Dörr R, Grossmann U, Kessler E (1986) Deoxyribonucleic acid reassociation in the taxonomy of the genus Chlorella. I. Chlorella sorokiniana. Arch Microbiol 145:329–333
Kimura M (1980) A simple method for estimating evolutionary rates of base substitution through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Krishnan S, Bamabas S, Barnabas J (1990) Interrelationships among major protistan groups based on a parsimony network of 5S rRNA sequences. Biosystems 24:135–144
Li W-H, Wolfe KH, Sourdis J, Sharp PM (1987) Reconstruction of phylogenetic trees and estimation of divergence times under nonconstant rates of evolution. Cold Spring Harb Symp Quant Biol 52:847–856
Liesack W, Söller R, Stewart T, Haas H, Giovannoni S, Stackebrandt E (1992) The influence of tachytelically (rapidly) evolving sequences on the topology of phylogenetic trees—intrafamily relationships and the phylogenetic position of Planctomycetaceae as revealed by comparative analysis of 16S ribosomal RNA sequences. Syst Appl Microbiol 15:357–362
Manhart JR, Palmer JD (1990) The gain of two chloroplast tRNA introns mark the green algal ancestors of land plants. Nature 345: 268–270
Mankin AS, Skryabin KG, Rubtsov PM (1986) Identification of ten additional nucleotides in the primary structure of yeast 18S rRNA. Gene 44:143
Mattox KR, Stewart KD (1984) Classification of the green algae: a concept based on comparative cytology. In: Irvin DEG, John DM (eds) Systematics of the green algae. Academic Press, London, pp 29–72
McCarroll R, Olsen GJ, Stahl YD, Woese CR, Sogin ML (1983) Nucleotide sequence of the Dictyostelium discoideum small-subunit ribosomal ribonucleic acid inferred from the gene sequence: Evolutionary implications. Biochemistry 22:5858–5868
McFadden GI, Melkonian M (1986) Use of Hepes buffer for microalgal culture media and fixation for electron microscopy. Phycologia 25:551–557
Medlin L, Elwood HJ, Stickel S, Sogin M (1988) The characterization of enzymatically amplified eukaryotic 16S-like rRNA-coding regions. Gene 71:491–499
Melkonian M (1990) Phylum Chlorophyta: class Prasinophyceae. In: Margulis L, Corliss JO, Melkonian M, Chapman DJ (eds) Handbook of Protoctista. Jones and Barlett, Boston, pp 600–607
Mishler BD, Churchill SP (1985) Transition to a land flora: phylogenetic relationships of the green algae and bryophytes. Cladistics 1:305–328
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 84:321–4325
Nairn CJ, Ferl RJ (1988) The complete nucleotide sequence of the small-subunit ribosomal RNA coding region for the cycad Zamia pumila: phylogenetic implications. J Mol Evol 27:133–141
Neefs J-M, De Wachter R (1990) A proposal for the secondary structure of a variable area of eukaryotic small ribosomal subunit RNA involving the existence of a pseudoknot. Nucleic Acids Res 18: 5695–5704
Ragan MA, Parsons TJ, Sawa T, Straus NA (1994) 18S ribosomal DNA sequences indicate a monophyletic origin of Charophyceae. J Phycol 30:490–500
Rathgeber J, Capesius I (1990) Nucleotide sequence of the intergenic spacer and the 18S ribosomal RNA gene from mustard (Sinapis alba). Nucleic Acids Res 18:1288
Raubeson LA, Jansen RK (1992) Chloroplast DNA evidence on the ancient evolutionary split in vascular land plants. Science 255: 1697–1699
Rouvière P, Mandelco L, Winkler S, Woese CR (1992) A detailed phylogeny for the Methanomicrobiales. System Appl Microbiol 15:363–371
Rubtsov PM, Musakhanov MM, Zakharyev VM, Krayev AS, Skryabin KG, Bayev AA (1980) The structure of the yeast ribosomal RNA genes. I. The complete nucleotide sequence of the 18S ribosomal RNA gene from Saccharomyces cerevisiae. Nucleic Acids Res 8: 5779–5794
Saitou N, Nei M (1987) The neighbor joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schönbohm E (1963) Untersuchungen über die Starklichtbewegung des Mougeotia-Chloroplasten. Zeitschr Bot 51:233–276
Schuster RM (1984) Evolution, phylogeny and classification of the Hepaticae. In: Schuster RM (ed) New manual of bryology. The Hatori Botanical Laboratory, Nichinan, Miyazaki, Japan, pp 1071–1092
Sensen CW, Banrevi A, Capesius I (1994) The sequence of the 18S rDNA gene from Pinus wallichiana. Plant Mol Biol 24:259
Sensen CW, Hinkle GJ, Sogin ML, Melkonian M (1992) The sequence of the 18S rDNA gene from Spermatozopsis similis Preisig et Melkonian, 1894 (Chlorophyceae). Plant Mol Biol 20:1209
Sogin ML (1989) Evolution of eukaryotic microorganisms and their small subunit ribosomal RNAs. Am Zool 29:487–499
Steele KP, Holsinger KE, Jansen RK, Taylor DW (1991) Assessing the reliability of 5S rRNA sequence data for phylogenetic analysis in green plants. Mol Biol Evol 8:240–248
Stewart C-B (1993) The powers and pitfalls of parsimony. Nature 361:603–607
Stewart WN (1983) Paleobotany and the evolution of plants. Cambridge University Press, Cambridge, pp 394–395
Surek B, Beemelmanns U, Melkonian M, Bhattacharya D (1994) Ribosomal RNA sequence comparisons demonstrate an evolutionary relationship between Zygnematales and charophytes. Plant Syst Evol 191:171–181
Swofford DL (1993) PAUP: phylogenetic analysis using parsimony, version 3.1.1. Illinois Natl Hist Surv, Champaign, IL
Tajima F (1993) Simple methods for testing the molecular evolutionary clock hypothesis. Genetics 135:599–607
Takaiwa F, Oono K, Sugiura M (1984) The complete nucleotide sequence of a rice 17S rRNA gene. Nucleic Acids Res 12:5441–5448
Van de Peer Y, De Baere R, Cauwenberghs J, De Wachter R (1990) Evolution of green plants and their relationship with other photosynthetic eukaryotes as deduced from 5S ribosomal RNA sequences. Plant Syst Evol 170:85–96
Wilcox LW, Fuerst PA, Floyd GL (1993) Phylogenetic relationships of four ‘charophycean’ green algae inferred from complete nuclear-encoded small subunit rRNA gene sequences. Am J Bot 80:1028–1033
Author information
Authors and Affiliations
Additional information
Correspondence to.: V.A.R. Huss
Rights and permissions
About this article
Cite this article
Kranz, H.D., Mikš, D., Siegler, ML. et al. The origin of land plants: Phylogenetic relationships among charophytes, bryophytes, and vascular plants inferred from complete small-subunit ribosomal RNA gene sequences. J Mol Evol 41, 74–84 (1995). https://doi.org/10.1007/BF00174043
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00174043