Abstract
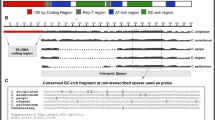
A comprehensive sequence analysis of three early chorion genes (6F6.1, 6F6.2, 6F6.3) which form a small subfamily is presented. Two main features characterize this subfamily: (1) the 6F6 gene copies are β-branch genes and, unlike typical chorion genes which are organized in divergent gene pairs, they are unpaired, and (2) they are not clustered in genetic locus Ch3 but are dispersed in Ch1-2, which is about 3 to 4 centiMorgans away and contains middle and late chorion genes. Sequence comparisons show that members of this subfamily exhibit high identity values in their major coding region (94–96%) and that similarities also extend, but to a lesser degree, into their noncoding regions. The putative 6F6 promoter regions have no significant similarities with the corresponding regions of other early β-genes but quite surprisingly share common elements with middle and late genes. The main difference among the 6F6 gene introns is the presence of inserted sequences: the insert into 6F6.2 (“IR”; 248 bp) is flanked by a 102–103-bp inverted repeat, while those into 6F6.1 (“FIB”; 184 bp) and 6F6.3 (“HOPE”; 951 bp) are carried by a partial Bm1 element. HOPE has features of a non-LTR retrotransposable element. Preliminary experiments indicate that the copy number of IR and HOPE in the Bombyx mori genome is about 5,000 and 20,000, respectively. The great similarity of 6F6 genes cannot be accounted for by selective pressure but rather appears to be the result of gene-conversion-like events, which are supposed to operate frequently in middle and late chorion genes but not in other known early β-genes. Using the relative position and orientation of the 6F6 gene copies, it is possible to propose an evolutionary scheme for the formation of chorion locus Ch1-2.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Adams DS, Eickbush TH, Herrera RJ, Lizardi PM (1986) A highly reiterated family of transcribed oligo(A)-terminated, interspersed DNA elements in the genome of Bombyx mori. J Mol Biol 187: 465–478
Berg JM (1986) Potential metal-binding domains in nucleic acid binding proteins. Science 232:485–487
Burke WD, Eickbush TH (1986) The silkmoth late chorion locus. I. Variation within two paired multigene families. J Mol Biol 190: 343–356
Burke WD, Eickbush DG, Xiong Y, Jakubczak J, Eickbush TH (1993) Sequence relationship of retrotransposable elements RI and R2 within and between divergent insect species. Mol Biol Evol 10: 163–185
Eickbush TH (1992) Transposing without ends: the non-LTR retrotransposable elements. New Biol 4:430–440
Eickbush TH, Burke WD (1985) Silkmoth chorion gene families contain patchwork patterns of sequence homology. Proc Natl Acad Sci USA 82:2814–2818
Eickbush TH, Burke WD (1986) The silkmoth late chorion locus. II. Gradients of gene conversion in two paired multigene families. J Mol Biol 190:357–366
Eickbush TH, Kafatos FC (1982) A walk in the chorion locus of {iyBombyx mori}. Cell 29:633–643
Eickbush TH, Rodakis GC, Lecanidou R, Kafatos FC (1985) A complex set of early chorion DNA sequences from Bombyx mori. Dev Biol 112:368–376
Gage LP (1974) The Bombyx mori genome: analysis by DNA reassociation kinetics. Chromosoma 45:27–42
Goldsmith MR (1989) Organization and developmental timing of the Bombyx mori chorion gene clusters in strain c108. Dev Genet 10: 16–23
Green LM, Berg JM (1989) A retroviral Cys-Xaa2-Cys-Xaa4 His-Xaa4-Cys peptide binds metal ions: spectroscopic studies and a proposed three-dimensional structure. Proc Natl Acad Sci USA 86:4047–4051
Henikoff S (1987) Exonuclease III generated deletions for DNA sequence analysis. Promega Notes No. 8
Hibner BL, Burke WD, Lecanidou R, Rodakis GC, Eickbush TH (1988) Organization and expression of three genes from the silkmoth early chorion locus. Dev Biol 125:423–431
Hibner BL, Burke WD, Eickbush TH (1991) Sequence identity in an early chorion multigene family is the result of localized gene conversion. Genetics 128:595–606
Jakubczak JL, Burke WD, Eickbush TH (1991) Retrotransposable elements R1 and R2 interrupt the rRNA genes of most insects. Proc Natl Acad Sci USA 88:3295–3299
Kafatos FC (1980) Recombinant DNA studies on the structure, evolution, and developmental expression of structural gene families. Am J Trop Med Hyg 29(5)Suppl:1111–1116
Kafatos FC, Mitsialis SA, Nguyen HT, Spoerel N, Tsitilou SG, Mazur GD (1987a) Evolution of structural genes and regulatory elements for the insect chorion. In: Raff R, Raff EC (eds) Development as an evolutionary process, vol 8. Alan R Liss, New York, pp 161–178
Kafatos FC, Spoerel N, Mitsialis SA, Nguyen HT, Romano C, Lingappa JR, Mariani BD, Rodakis GC, Lecanidou R, Tsitilou SG (1987b) Developmental control and evolution in the chorion gene families of insects. Adv Genet 24:223–242
Lecanidou R, Eickbush TH, Rodakis GC, Kafatos FC (1983) Novel B family sequence from an early chorion cDNA library of Bombyx mori. Proc Natl Acad Sci USA 80:1955–1959
Lecanidou R, Rodakis GC, Eickbush TH, Kafatos FC (1986) Evolution of a silkmoth chorion gene superfamily: gene families CA and CB. Proc Natl Acad Sci USA 83:6514–6518
Lecanidou R, Rodakis GC (1992) Three copies of the early gene 6F6 are interspersed in and around the late chorion gene cluster of Bombyx mori. J Mol Evol 34:304–314
Martin SL (1991) LINEs. Carr Opin Genet Dev 1:505–508
Mitsialis SA, Kafatos FC (1985) Regulatory elements controlling chorion gene expression are conserved between flies and moths. Nature 317:453–456
Mitsialis SA, Spoerel N, Leviten M, Kafatos FC (1987) A short defined DNA region is sufficient for developmentally correct expression of moth chorion genes in Drosophila. Proc Natl Acad Sci USA 84: 7987–7991
Mori S, Izumi S, Tomino S (1991) Complete nucleotide sequences of major plasma protein genes of Bombyx mori. Biochim Biophys Acta 1090:129–132
Pearson WR, Lipman DJ (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci USA 85:2444–2448
Pustell J, Kafatos FC (1984) A convenient and adaptable package of computer programs for DNA and protein sequence management, analysis and homology determination. Nucleic Acids Res 12:643–655
Pustell J, Kafatos FC (1986) A convenient and adaptable microcomputer environment for DNA and protein sequence manipulation and analysis. Nucleic Acids Res 14:479–488
Regier JC, Wiegmann BM, Leclerc RF, Friedlander TP (1994) Loss of phylogenetic information in chorion gene families of Bombyx mori by gene conversion. Mol Biol Evol 11:72–87
Rodakis GC, Lecanidou R, Eickbush TH (1984) Diversity in a chorion multigene family created by tandem duplications and a putative gene-conversion event. J Mol Evol 20:265–273
Rodakis GC, Lecanidou R (1992) The possible evolutionary significance of repeat elements near and within an early chorion gene in the late chorion locus of Bombyx mori. J Mol Evol 34:315–323
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Laboratory Press, Cold Spring Harbor, New York
Smith GP (1973) Unequal crossover and the evolution of multigene families. Cold Spring Harbor Symp Quant Biol 38:507
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Spoerel N, Nguyen HT, Eickbush TH, Kafatos FC (1989) Gene evolution and regulation in the chorion complex of Bombyx mori: hybridization and sequence analysis of multiple developmentally middle A/B chorion gene pairs. J Mol Biol 209:1–19
Weiner AM, Deininger PL, Efstratiadis A (1986) Nonviral retroposons: genes, pseudogenes, and transposable elements generated by the reverse flow of genetic information. Ann Rev Biochem 55:631–661
Xiong Y, Sakaguchi B, Eickbush TH (1988) Gene conversions can generate sequence variants in the late chorion multigene families of Bombyx mori. Genetics 120:221–231
Author information
Authors and Affiliations
Additional information
Correspondence to: G.C. Rodakis
Rights and permissions
About this article
Cite this article
Kravariti, L., Lecanidou, R. & Rodakis, G.C. Sequence analysis of a small early chorion gene subfamily interspersed within the late gene locus in Bombyx mori . J Mol Evol 41, 24–33 (1995). https://doi.org/10.1007/BF00174038
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00174038