Abstract
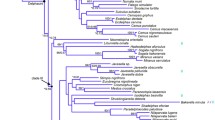
Phylogenetic relationships among eight species of the Drosophila buzzatii species complex (D. mulleri subgroup; D. repleta species group) and D. hamatofila were determined by sequencing the mitochondrial cytochrome oxidase subunit 1, II, and III genes. The species examined included members of the martensis cluster (D. martensis, D. starmeri, D. venezolana), the buzzatii cluster (D. buzzatii, D. serido, D. borborema), and the stalkeri cluster (D. stalkeri, D. richardsoni). The molecular phylogeny was found to be congruent with the chromosomal inversion phylogeny. Analyzing the cytochrome oxidase subunits separately revealed that not all the subunits seem to have the same phylogenetic information content. Parameters are discussed that might explain these differences.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Barrio E, Latorre A, Moya A (1994) Phylogeny of the Drosophila obscura species group deduced from mitochondrial DNA sequences. J Mol Evol 39:478–488
Beckenbach AT, Wei YW, Liu H (1993) Relationships in the Drosophila obscura species group, inferred from mitochondrial cytochrome oxidase 11 sequences. Mol Biol Evol 10:619–634
Benado M, Fontdevilla A, Cerda HG, Garcia G, Ruiz A, Montero C (1984) On the distribution and cactiphilic niche of Drosophila martensis in Venezuela. Biotropica 16:120–124
Brower AVZ (1994) Phylogeny of Heliconius butterflies inferred from mitochondrial DNA sequences (Lepidoptera: Nymphalidae). Mol Phyl Evol 3:159–174
Brown JM, Pellmyr O, Thompson JN, Harrison RG (1994) Mitochondrial DNA phylogeny of the Prodoxidae (Lepidoptera: Incruvarioidea) indicates rapid ecological diversification of Yucca moths. Ann Entomol Soc Amer 87:795–802
Brown WM, Prager EM, Wang A, Wilson AC (1982) Mitochondrial DNA sequences of primates: tempo and mode of evolution. J Mol Evol 18:225–239
Casanova J-L, Pannetier C, Jaulin C, Kourilsky P (1990) Optimal conditions for directly sequencing double-stranded PCR products with Sequenase. Nucl Acids Res 18:4028
Clary DO, Wolstenhome DR (1985) The mitochondrial DNA molecule of Drosophila yakuba: nucleotide sequence, gene organization and genetic code. J Mol Evol 22:252–271
Collins TM, Wimberger PH, Naylor GJP (1994) Compositional bias, character-state bias, and character-state reconstruction using parsimony. Syst Biol 43:482–496
craft J Helm-Bychowski K (1991) Parsimony and phylogenetic inference using DNA sequences: some methodological difficulties. In Miyamoto MM, Cracraft J (eds) Phylogenetic analysis of DNA sequences, Oxford Univ Press, New York, pp 184–220
de Bruijn MHL (1983) Drosophila melanogaster mitochondrial DNA, a novel gene organization and genetic code. Nature 304:234–241
DeSalle R, Freedman T, Prager EM, Wilson AC (1987) Tempo and mode of sequence evolution in mitochondrial DNA of Hawaiian Drosophila. J Mol Evol 26:157–164
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Felsenstein J (1993) PHYLIP: Phylogeny inference Package (ver 3.5c). Univ of Washington, Seattle
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192
Irwin DM, Kocher TD, Wilson AC (1991) Evolution of the cytochrome b gene of mammals. J Mol Evol 32:128–144
Jin L, Nei M (1990) Limitations of the evolutionary parsimony method of phylogenetic analysis. Mol Biol Evol 7:82–102
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kishino H, Hasegawa M (1989) Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in the Hominoidea. J Mol Evol 29: 170–179
Lake JA (1994) Reconstructing evolutionary trees from DNA and protein sequences: paralinear distances. Proc Natl Acad Sci USA: 91: 1455–1459
Liu H, Beckenbach AT (1992) Evolution of the mitochondrial cytochrome oxidase 11 gene among 10 orders of insects. Mol Phyl Evol 1:41–52
Lockhart PJ, Steel MA, Hendy MD, Penny D (1994) Recovering evolutionary trees under a more realistic model of sequence evolution. Mol Biol Evol 11: 605–612
MacIntyre RJ, Collier GE (1986) Protein evolution in the genus Drosophila. In Ashburner M, Carson HL, Thompson IN (eds), The genetics and biology of Drosophila, vol. 3E. Academic Press, London, pp 39–146
Maddison WP, Maddison DR (1992) MacClade: Analysis of phylogeny and character evolution (ver 3.05). Sinauer Associates, Sunderland, Massachusetts
Margush T, McMorris FR (1981) Consensus n-trees. Bull Mathem Biol 43:239–244
Marin I, Ruiz A, Pla C, Fontdevila A (1994) Reproductive relationships among ten species of the Drosophila repleta group from South America and the West Indies. Evolution 47:1616–1624
Martin AP, Kessing BD, Palumbi SR (1990) Accuracy of estimating genetic distance between species from short sequences of mitochondrial DNA. Mol Biol Evol 7:485–488
Mullis K, Faloona F, Scharf S, Saiki R, Horn G, Erlich HA (1987) Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Col Spring Harbor Symp Quant Biol 51:263–273
Olmstead RG, Sweere JA (1994) Combining data in phylogenetic systematics: an empirical approach using three molecular data set in the Solanaceae. Syst Biol 43:467–481
Rohlf FJ (1982) Consensus indices for comparing classifications. Math Biosci 59:131–144
Ruiz A, Wasserman M (1993) Evolutionary cytogenetics of the Drosophila buzzatii species complex. Heredity 70:582–596
Saccone C, Pesole G, Preparata G. (1989) DNA microenvironments and the molecular clock. J Mol Evol 29:407–411
Saccone C, Lanave C, Pesole G (1993) Time and biosequences. J Mol Evol 37:154–159
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA (1988) Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239: 487–491
Saitou N, Imanishi M (1989) Relative efficiencies of the Fitch-Margolish, maximum-parsimony, maximum-likelihood, minimumevolution, and neighbor-joining methods of phylogenetic tree reconstruction in obtaining the correct tree. Mol Biol Evol 6:514–525
Simon C, Franke A, Martin A (1991) The polymerase chain reaction: DNA extraction and amplication. In Hewitt GM, Johnston AWB, Younr JPW (eds) Molecular techniques in taxonomy. NATO ASI Series, Springer-Verlag, Berlin Heidelberg, pp 329–355
Simon C, Frati F, Beckenbach A, Crespi B, Liu H, Flook P (1994) Evolution, weighting and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann Entomol Soc Amer 87:651–701
Sperling FAH, Hickey DA (1994) Mitochondrial DNA sequence variation in the spruce budworm species complex (Choristoneura: Lepidoptera). Mol Biol Evol 11:656–665
Spicer GS (1992) Reevaluation of the phylogeny of the Drosophila virilis species group (Diptera: Drosophilidae). Ann Entomol Soc Amer 85:11–25
Sullivan J, Holsinger KE, Simon C (1995) Among-site rate variation and phylogenetic analysis of 12S rRNA in sigmodontine rodents. Mol Biol Evol 12:988–1001
Swofford DL (1995) PAUP*Star (test ver 4.0). Sinauer, Sunderland, Massachusetts
Swofford DL, Olsen GJ (1990) Phylogeny reconstruction. In: Hillis DM, Moritz C, (eds) Molecular Systematics, Sinauer, Sunderland, Massachusetts, pp 411–501
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Templeton AR (1983) Phylogenetic inference from restriction endonuclease cleavage site maps with particular reference to the evolution of humans and apes. Evolution 37:221–224
Tidon-Sklorz R, Sene FM (1995) Drosophila seriema n. sp.: new member of the Drosophila serido (Diptera: Drosophilidae) superspecies taxon. Ann Entomol Soc Amer 88:139–142
Wasserman M (1982a) Evolution of the repleta group. In Ashburner M, Carson HL, Thompson JN (eds), The genetics and biology of Drosophila, vol. 3B. Academic Press, London, pp 61–139
Wasserman M (1982b) Cytological evolution in the Drosophila repleta species group. In Barker JSF, Starmer WT (eds), Ecological genetics and evolution: the cactus-yeast-Drosophila model system. Academic Press, New York, pp 49–64
Wasserman M (1992) Cytological evolution of the Drosophila repleta species group. In Krimbas CB, Powell JR (eds), Drosophila inversion polymorphism. CRC Press, Boca Raton, pp. 455–552
Yang Z (1994) Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J Mol Evol 39:306–314
Author information
Authors and Affiliations
Additional information
Correspondence to: G. Spicer
Rights and permissions
About this article
Cite this article
Spicer, G.S. Phylogenetic utility of the mitochondrial cytochrome oxidase gene: Molecular evolution of the Drosophila buzzatii species complex. J Mol Evol 41, 749–759 (1995). https://doi.org/10.1007/BF00173155
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00173155