Abstract
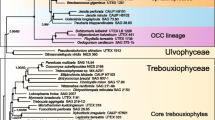
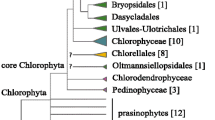
The marine green coccoidal alga Nanochlorum eukaryotum (N.e.) is of small size with an average diameter of 1.5 μm. It is characterized by primitive-appearing biochemical and morphological properties, which are considerably different from those of other green algae. Thus, it has been proposed that N.e. may be an early developed algal form. To prove this hypothesis, DNA of N.e. was isolated by a phenol extraction procedure, and the chloroplast DNA separated by preparative CsCl density-gradient centrifugation. The kinetic complexity of the nuclear and of the chloroplast DNA was evaluated by reassociation kinetics to 3 × 107 by and 9 × 104 bp, respectively. Several chloroplast genes, including the rRNA genes, were cloned on distinct fragments. The order of the rRNA genes corresponds to the common prokaryotic pattern. The 16S rRNA gene comprises 1,548 bases and is separated from the 23S rRNA gene with its 2,920 bases by a short spacer of 460 bases, which also includes the tRNAIle and tRNAAla genes. The 5S rRNA gene has not been found; it must start further than 500 bases downstream from the 3′-end of the 23S rRNA gene. From the chloroplast rRNA sequences, we have deduced secondary structures of the 16S and 23S rRNAs, which are in agreement with standard models. The rRNA sequences were aligned with corresponding chloroplast sequences; phylogenetic relationships were calculated by several methods. From these calculations, we conclude that N.e. is most closely related to Chlorella vulgaris. Therefore, N.e. does not represent an early developed algal species; the primitive-appearing morphological and biochemical characteristics of N.e. must rather be explained by secondary losses.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Andreoli C, Rascio N, Casodoro G (1978) Chlorella nana sp. nov. (Chlorophyceae): a new marine Chlorella. Bot Mar 21:253–256
Archie JW (1989) Phylogenies of plant families: a demonstration of phylogenetic randomness in DNA sequence data derived from proteins. Evolution 43:1796–1800
Assali NE, Martin WF, Sommerville CC, Loiseaux de Goer S (1991) Evolution of the rubisco operon from prokaryotes to algae: structure and analysis of the rbcS gene of the brown alga Pylaiella littoralis. Plant Mol Biol 17:853–863
Assali NE, Mache R, Loiseaux de Goer S (1990) Evidence for a composite phylogenetic origin of the plastid genome of the brown alga Pylaiella littoralis (L.) Kjellm. Plant Mol Biol 15:307–315
Bachellerie JP, Michot B (1989) Evolution of large subunit rRNA structure: the 3′-terminal domain contains elements of secondary structure specific to major phylogenetic groups. Biochimie 71:701–709
Boczar BA, Delaney T, Cattolico RA (1989) The gene for the ribulose-1,5-bisphosphate carboxylase (Rubisco) small subunit protein of the marine chromophyte Olisthodiscus luteus is similar to that of a chemoautotrophic bacterium. Proc Natl Acad Sci USA 86:4996–4999
Bolivar F, Rodriguez RL, Betlach MC, Boyer HW (1977) Construction and characterization of new cloning vehicles I. Ampicillin-resistant-derivatives of the plasmid pMB9. Gene 2:75–93
Britten RJ, Davidson EH (1976) Studies on nucleic acid reassociation kinetics: empirical equations describing DNA reassociation. Proc Natl Acad Sci USA 73:415–419
Britten RJ, Graham DE, Neufeld BR (1974) Analysis of repeating DNA sequences by reassociation. In: Grossman L, Moldave K (eds) Methods in enzymology, vol XXIX. Academic Press, New York, pp 363–418
Cedergren R, Gray MW, Abel Y, Sankoff D (1988) The evolutionary relationship among known life forms. J Mol Evol 28:98–112
Cryer DF, Eccleshall R, Marmur J (1975) Isolation of yeast DNA. In: Prescott DM (ed) Methods in cell biology, vol XII. Academic Press, New York, pp 39–44
Dagert M, Ehrlich SD (1979) Prolonged incubation in calcium chloride improves the competence of E. coli cells. Gene 6:23–28
Dams E, Hendriks L, van de Peer Y, Neefs JF, Smits G, Vandenbempt I, De Wachter R (1988) Compilation of small ribosomal subunit RNA sequences. Nucleic Acids Res 16:r87-r173
Dempsey GP, Lawrence D, Cassie V (1980) The ultrastructure of Chlorella minutissima Fott et Novakova (Chlorophyceae, Chlorococcales). Phycologia 19:13–19
Dörr R, Huss VAR (1990) Characterization of nuclear DNA in 12 species of Chlorella (Chlorococcales, Chlorophyta) by DNA reassociation. Biosystems 24:145–155
Douglas SE (1992) Eukaryote-eukaryote endosymbioses: insights from studies of a cryptomonad alga. Biosystems 28:57–68
Douglas SE, Murphy CA, Spencer DF, Gray MW (1991) Cryptomonad algae are evolutionary chimaeras of two phylogenetically distinct unicellular eukaryotes. Nature 350:148–151
Engberg J, Nielsen H, Lenaers G, Fujitani H, Higashinakagawa T (1990) Comparison of primary and secondary 26S rRNA structures in two Tetrahymena species: evidence for a strong evolutionary and structural constraint in expansion segments. J Mol Evol 39:514–521
Eschbach S, Wolters J, Sitte P (1991) Primary and secondary structure of the nuclear small subunit ribosomal RNA of the Cryptomonad Pyrenomonas salina as inferred from the gene sequence: evolutionary implications. J Mol Evol 32:247–252
Felsenstein J (1989) Phylip manual 3.2. University of California, Berkeley
Felsenstein J (1988) Phylogeny from molecular sequences: inference and reliability. Anna Rev Genet 22:521–565
Freier SM, Kierzek R, Jaeger JA, Sugimoto N, Caruthers MH, Neilson T, Turner DH (1986) Improved free-energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci USA 83:9373–9377
Geisert M, Rose T, Bauer W, Zahn RK (1987) Occurrence of carotenoids and sporopollenin in Nanochlorum eukaryotum. Biosystems 20:133–142
Gillis M, DeLey J, De Cleene M (1970) The determination of molecular weight of bacterial genome DNA from renaturation rates. Eur J Biochem 12:143–153
Giovannoni SJ, Turner S, Olsen GJ, Barns S, Lane DJ, Pace NR (1988) Evolutionary relationship among cyanobacteria and green chloroplasts. J Bacteriol 170:3584–3592
Goldman N (1993) Statistical tests of models of DNA substitution. J Mol Evol 36:182–198
Grace SC (1990) Phylogenetic distribution of superoxid dismutase supports an endosymbiotic origin for chloroplasts and mitochondria. Life Sci 47:1875–1886
Gutell RB, Fox GE (1988) A compilation of large subunit RNA sequences presented in a structural format. Nucleic Acids Res 16:r175-r269
Gutell RB, Weiser B, Woese CR, Noller HF (1985) Comparative anatomy of 16S-like ribosomal RNA. Prog Nucleic Acid Res Mol Biol 32:155–216
Heath I (1980) Variant mitosis in lower eukaryotes, indication of the evolution of mitosis. Int Rev Cytol 64:1–80
Hendriks L, De Baere R, Van de Peer Y, Neefs J, Goris A, De Wachter R (1991) The evolutionary position of the rhodophyte Porphyra umbilicalis and the basidiomycete Leucosporidium scottii among other eukaryotes as deduced from complete sequences of small ribosomal subunit RNA. J Mol Evol 32:167–177
Herzog M, von Boletsky S, Soyer MO (1984) Ultrastructural and biochemical nuclear aspects of eukaryote classification: independent evolution of the dinoflagellates as a sister group of the actual eukaryotes? Orig Life 13:205–312
Higgins DG, Sharp PM (1988) CLUSTAL: a package for performing multiple sequence alignments on a microcomputer. Gene 73:237–244
Hillis DM, Huelsenbeck JP (1992) Signal, noise and reliability in molecular phylogenetic analysis. J Hered 83:189–195
Hori H, Osawa S (1987) Origin and evolution of organisms as deduced from 5S ribosomal RNA sequences. Mol Biol Evol 4:445–472
Janssen I, Jakowitsch J, Michalowski CB, Bohnert HJ, Löffelhardt W (1989) Evolutionary relationship of psbA genes from cyanobacteria, cyanelles and plastids. Curr Genet 15:335–340
Johnson PW, Sieburth J (1982) In situ morphology and occurrence of eukaryotic phototrophs of bacterial size in the picoplankton of estuarian and oceanic waters. J Phycol 18:318–327
Kimura M (1980) A simple model for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kraus M, Götz M, Löffelhardt W (1990) The cyanelle str operon from Cyanophora paradoxa: sequence analysis and phylogenetic implications. Plant Mol Biol 15:561–573
Lenaers G, Scholin Ch, Bhaud Y, Saint-Hilaire D, Herzog M (1991) A molecular model of Dinoflagellate protist (Pyrrhophyta) inferred from the sequence of 24S rRNA divergent domains D1 and D8. J Mol Evol 32:53–63
Lenaers G, Maroteaux L, Michot B, Herzog M (1989) Dinoflagellates in evolution. A molecular phylogenetic analysis of large subunit ribosomal RNA. J Mol Evol 29:40–51
Macaya G, Thiery JP, Bernardi G (1976) An approach to the organization of eukaryotic genomes at a macromolecular level. J Mol Biol 108:237–254
Manhart JR, Kelly K, Dudock BS, Palmer JD (1989) Unusual characteristics of Codium fragile chloroplast DNA revealed by physical and gene mapping. Mol Gen Genet 216:417–421
Margulis L (1981) Symbiosis in cell evolution. WH Freeman and Co., San Francisco, CA, pp 233–283
Markowicz Y, Loiseaux-de Goer S (1991) Plastid genomes of the Rhodophyta and Chromophyta constitute a distinct lineage which differs from that of the Chlorophyra and have a composite phylogenetic origin, perhaps like that of the Euglenophyta. Curr Genet 20:427–430
Michot B, Qu LH, Bachellerie JP (1990) Evolution of large-subunit rRNA structure. The diversification of divergent D3 domain among major phylogenete groups. Eur J Biochem 188:219–229
Michot B, Bachellerie JP (1987) Comparisons of large subunit rRNA reveal some eukaryote-specific elements of secondary structure. Biochimie 69:11–23
Morden W, Delwichen CF, Kuhsel M, Palmer JD (1992) Gene phylogenies and the endosymbiotic origin of plastids. Biosystems 28: 75–90
Mortimer RK, Contopoulou CR, King JS (1992) Genetic and physical maps of S. cerevisiae, edition 11. Yeast 8:817–902
Palmer JD (1985) Comparative organization of chloroplast genomes. Annu Rev Genet 19:325–354
Penny D, Hendy MD, Steel MA (1992) Progress with methods for constructing evolutionary trees. Trends Ecol Evol 7:73–79
Rausch H, Larsen N, Schmitt R (1989) Phylogenetic relationship of the green alga Volvox carteri deduced from small-subunit ribosomal RNA comparisons. J Mol Evol 29:255–265
Rigby PW, Dieckmann M, Rhodes C, Berg P (1977) Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA-Polymerase I. J Mol Biol 113:237–251
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Sargent M, Zahn RK, Walters B, Gupta R, Kain B (1988) Nucleotide sequence of the 18S rRNA from the microalga Nanochlorum eukaryotum. Nucleic Acids Res 16:41–56
Schwartz RM, Dayhoff MO (1978) Origin of prokaryotes, eukaryotes, mitochondria, and chloroplasts. Science 199:395–403
Shivji MS, Li N, Cattolico RA (1992) Structure and organization of rhodophyte and chromophyte plastid genomes: implications for the ancestry of plastids. Mol Gen Genet 232:65–73
Szybalski W, Szybalski EH (1971) Equilibrium density gradient centrifugation. In: Cantoni GL, Davis DR (eds) Procedures in nucleic acid research, vol II. Harper and Row, New York, pp 333–343
Thinh LV, Griffiths DJ (1985) A small marine Chlorella from the waters of a coral reef. Bot Mar 28:41–46
Thorne JL, Kishino H, Felsenstein J (1991) An evolutionary model for maximum likelihood alignment of DNA sequences. J Mol Evol 33:114–124
Turner MF, Gowen RJ (1984) Some aspects of the nutrition and taxonomy of fourteen small green and yellow-green algae. Bot Mar 27:249
Turner S, Burger-Wiersma T, Giovannoni SJ, Mur LR, Pace NR (1989) The relationship of a prochlorophyte Prochlorothrix hollandica to green chloroplasts. Nature 337:380–385
Valentin K, Zetsche K (1990) Rubisco genes indicate a close phylogenetic relation between the plastids of Chromophyta and Rhodophyta. Plant Mol Biol 15:575–584
Van de Peer Y, Neefs JM, De Wachter R (1990) Small ribosomal subunit RNA sequences, evolutionary relationship among different life forms, and mitochondrial origins. J Mol Evol 30:463–476
Wetmur JG, Davidson N (1968) Kinetics of renaturation of DNA. J Mol Biol 31:349–370
Wilhelm C, Eisenbeis G, Wild A, Zahn RK (1982) Nanochlorum eukaryotum: a very reduced coccoid species of marine chlorophyceae. Z Naturforsch 37C:107–114
Woese CR (1987) Bacterial evolution. Microbiol Rev 51:221–271
Yamada T, Sakaguchi K (1981) Protoplast induction in Chlorella species. Agric Biol Chem 45:1905–1909
Zahn RK (1984) A green alga with minimal eukaryofc features: Nanochlorum eukaryotum. Orig Life 13:289–303
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary informations. Nucleic Acids Res 9:133–148
Author information
Authors and Affiliations
Additional information
Correspondence to: D. Weinblum
Rights and permissions
About this article
Cite this article
Schreiner, M., Geisert, M., Oed, M. et al. Phylogenetic relationship of the green alga Nanochlorum eukaryotum deduced from its chloroplast rRNA sequences. J Mol Evol 40, 428–442 (1995). https://doi.org/10.1007/BF00164029
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00164029