Abstract
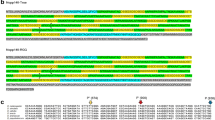
Runs of identical amino acids encoded by triplet repeats (homopolymers) are components of numerous proteins, yet their role is poorly understood. Large numbers of homopolymers are present in the Drosophila melanogaster mastermind (mam) protein surrounding several unique charged amino acid clusters. Comparison of mam sequences from D. virilis and D. melanogaster reveals a high level of amino acid conservation in the charged clusters. In contrast, significant divergence is found in repetitive regions resulting from numerous amino acid replacements and large insertions and deletions. It appears that repetitive regions are under less selective pressure than unique regions, consistent with the idea that homopolymers act as flexible spacers separating functional domains in proteins. Notwithstanding extensive length variation in intervening homopolymers, there is extreme conservation of the amino acid spacing of specific charge clusters. The results support a model where homopolymer length variability is constrained by natural selection.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Beachy P, Helfand S, Hogness D (1985) Segmental distribution of bithorax complex proteins during Drosophila development. Nature 313:545–551
Bettler D, Schmid A, Yedvobnick B (1991) Early ventral expression of the Drosophila neurogenic locus mastermind. Dev Biol 144:436–439
Beverley S, Wilson A (1984) Molecular evolution in Drosophila and higher Diptera. II. A time scale for fly evolution. J Mol Evol 21:1–13
Brendel V, Karlin S (1989) Association of charge clusters with functional domains of cellular transcription factors. Proc Natl Acad Sci USA 86:5696–5702
Chen DJ, Chan CS, Pirrotta V (1992) Conserved DNA binding and self-association domains of the Drosophila zeste protein. Mol Cel Biol 12:598–608
Chen E, Seeburg P (1985) Supercoil sequencing: a fast and simple method for sequencing plasmid DNA. DNA 4:165–170
Costa R, Peixoto A, Thackery J, Dalgleish R, Kyriacou C (1991) Length polymorphism in the threonine-glycine encoding repeat region of the period gene in Drosophila. J Mol Evol 32:238–246
Danielson M, Northrop J, Ringold G (1986) The mouse glucocorticoid receptor: Mapping of functional domains by cloning, sequencing and expression of wild-type and mutant receptor proteins. EMBO J 5:2513–2522
Dente L, Sollazzo M, Baldari C, Cesareni G, Cortese R (1985) The pEMBL family of single stranded vectors. In: Glover D (ed) DNA cloning: A practical approach, volume 1. IRL Press, Oxford, England, pp 101–108
Dover G (1986) Molecular drive in multigene families: how biological novelties arise, spread and are assimilated. Trends Genet 2:159–165
Duboule D, Haenlin M, Galliot B, Mohier E (1987) DNA sequences homologous to the Drosophila opa repeat are present in murine mRNAs that are differentially expressed in fetuses and adult. Mol Cell Biol 7:2003–2006
Farabaugh P, Schmeissner U, Hoter M, Miller J (1979) On the molecular nature of spontaneous hotspots in the lacI gene of Escherichia coli. J Mol Biol 126:847–863
Faulkner DV, Jurka J (1988) Multiple aligned sequence editor (MASE). Trends Biochem Sci 13:321–322
Finkelstein R, Smouse D, Capaci T, Spradling A, Perrimon N (1990) The orthodenticle gene encodes a novel homeodomain protein involved in the development of the Drosophila nervous system and ocellar visual structure. Genes Dev 4:516–527
Fischer J, Giniger E, Maniatis T, Ptashne M (1988) GAL4 activates transcription in Drosophila. Nature 332:853–856
Friedel C (1990) An analysis of the transcriptional activity at the neurogenic locus mastermind in Drosophila melanogaster. MS thesis, Emory University, Atlanta, GA
Fu YH, Kuhl DPA, Pizzuti A, Pieretti M, Sutcliffe J, Richards S, Verkerk AJMH, Holden JJA, Fenwick Jr RG, Warren ST, Oostra BA, Nelson DL, Caskey CT (1991) Variation of the CGG repeat at the Fragile X site results in genetic instability: resolution of the Sherman paradox. Cell 67:1047–1058
Grantham R, Gautier C, Gouy M, Jacobzone M, Mercier R (1981) Codon catalog usage is a genome strategy modulated for gene expressivity. Nucleic Acids Res 9:r43-r73
Harley H, Brook J, Rundle S, Crows S, Reardon W, Buckler A, Harper P, Houseman D, Shaw D (1992) Expansion of an unstable DNA region and phenotypic variation in myotonic dystrophy. Nature 355:545–546
Heberlein U, Rubin G (1990) Structural and functional comparisons of the Drosophila virilis and Drosophila melanogaster rough genes. Proc Natl Acad Sci USA 87:5916–5920
Jeffreys A, Royle N, Wilson V, Wong Z (1988) Spontaneous mutation rates to new length alleles at tandem repetitive hypervariable loci in human DNA. Nature 322:278–281
Jones C, Dalton M, Townley L (1991) Interspecific comparisons of the structure and regulation of the Drosophila ecdysone inducible gene E74. Genetics 127:535–543
Kassis J, Poole S, Wright D, O'Farrell P (1986) Sequence conservation in the protein coding and intron regions of the engrailed transcription unit. EMBO J 5:3583–3589
Kozak M (1989) The scanning model for translation: An update. J Cell Biol 108:262–273
LaSpada A, Wilson E, Lebahn D, Harding A, Fischback K (1991) Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352:77–79
Laughon A, Carrol S, Storter F, Riley P, Scott M (1985) Common properties of proteins encoded by the Antennapedia complex genes of Drosophila melanogaster. Cold Spring Harb Symp Quant Biol 50:253–262
Levinson G, Gutman G (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol 4:203–221
Li WH, Tanimura M (1987) The molecular clock runs more slowly in man than in apes and monkeys. Nature 326:93–96
Lyons K, Stein J, Smithies O (1988) Length polymorphism in human proline rich proteins generated by intragenic unequal crossingover. Genetics 120:267–278
Michael W, Bowtell D, Rubin G (1990) Comparison of the sevenless genes of Drosophila virilis and Drosophila melanogaster. Proc Natl Acad Sci USA 87:5351–5353
Newfeld S, Smoller D, Yedvobnick B (1991) Interspecific comparison of the unusually repetitive Drosophila locus mastermind. J Mol Evol 32:415–420
O'Neil M, Belote J (1992) Interspecific comparison of the transformer gene of Drosophila reveals an unusually high degree of evolutionary divergence. Genetics 131:113–128.
Patterson T, Dean M (1987) Preparation of high titre lambda phage lysates. Nucleic Acids Res 15:6298
Peterson M, Tanese N, Pugh B, Tjian R (1990) Functional domains and upstream activation properties of cloned human TATA binding protein. Science 248:1625–1630
Peixoto AA, Costa R, Wheeler DA, Hall JC, Kyriacou CP (1992) Evolution of the threonine-glycine repeat region of the period gene in the melanogaster species subgroup of Drosophila. J Mol Evol 35:411–419
Rothstein R, Helsm C, Rosenberg N (1987) Concerted deletions and inversions are caused by mitotic recombination between delta sequences in Saccharomyces cerevisiae. Mol Cell Biol 7:1198–1207
Smith RF, Smith TF (1990) Automatic generation of primary sequence patterns from sets of related protein sequences. Proc Natl Acad Sci USA 87:118–122
Smith RF, Smith TF (1992) Pattern Induced Multi-sequence Alignment (PIMA) algorithm employing secondary structure-dependent gap penalties for use in comparative protein modelling. Protein Engineering 5:35–111
Smoller D, Friedel C, Schmid A, Bettler D, Lam L, Yedvobnick B (1990) The Drosophila neurogenic locus mastermind encodes a nuclear protein unusually rich in amino acid homopolymers. Genes Dev 4:1688–1700
Smoller D, Petrov D, Hartl D (1991) Characterization of bacteriophage P1 library containing inserts of Drosophila DNA of 75–100 kilobase pairs. Chromosoma 100:487–494
Tachida H, Iizuka M (1992) Persistence of repeated sequences that evolve by replication slippage. Genetics 131:471–478
Tautz D (1989) Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res 17:6463–6471
Treier M, Pfeifle C, Tautz D (1989) Comparison of the gap segmentation gene hunchback between Drosophila melanogaster and Drosophila virilis reveals novel modes of evolutionary change. EMBO J 8:1517–1525
Tseng H, Green H (1988) Remodeling of the involucrin gene during primate evolution. Cell 54:411–496
Wharton K, Yedvobnick B, Finnerty V, Artavanis-Tsakonas S (1985) Opa: A novel family of transcribed repeats shared by the Notch locus and other developmentally regulated loci in D. melanogaster. Cell 40:55–62
Wheeler C, Maloney D, Fogel S, Goodenow R (1990) Microconversion between murine H-2 genes integrated into yeast. Nature 347:192–194
Wilde CD, Akam M (1987) Conserved sequence elements in the 5′ region of the Ultrabithorax transcription unit. EMBO J 5:1393–1401
Yedvobnick B, Smoller D, Young P, Mills D (1988) Molecular analysis of the neurogenic locus mastermind of Drosophila melanogaster. Genetics 118:483–497
Zhu Q, Smith T, Lathrop R, Figge J (1990) Acid helix-turn activator motif. Proteins 8:156–163
Author information
Authors and Affiliations
Additional information
Correspondence to: B. Yedvobnick
Rights and permissions
About this article
Cite this article
Newfeld, S.J., Schmid, A.T. & Yedvobnick, B. Homopolymer length variation in the Drosophila gene mastermind . J Mol Evol 37, 483–495 (1993). https://doi.org/10.1007/BF00160429
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00160429