Summary
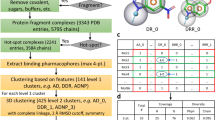
A new method is presented for computer-aided ligand design by combinatorial selection of fragments that bind favorably to a macromolecular target of known three-dimensional structure. Firstly, the multiple-copy simultaneous-search procedure (MCSS) is used to exhaustively search for optimal positions and orientations of functional groups on the surface of the macromolecule (enzyme or receptor fragment). The MCSS minima are then sorted according to an approximated binding free energy, whose solvation component is expressed as a sum of separate electrostatic and nonpolar contributions. The electrostatic solvation energy is calculated by the numerical solution of the linearized Poisson-Boltzmann equation, while the nonpolar contribution to the binding free energy is assumed to be proportional to the loss in solvent-accessible surface area. The program developed for computational combinatorial ligand design (CCLD) allows the fast and automatic generation of a multitude of highly diverse compounds, by connecting in a combinatorial fashion the functional groups in their minimized positions. The fragments are linked as two atoms may be either fused, or connected by a covalent bond or a small linker unit. To avoid the combinatorial explosion problem, pruning of the growing ligand is performed according to the average value of the approximated binding free energy of its fragments. The method is illustrated here by constructing candidate ligands for the active site of human α-thrombin. The MCSS minima with favorable binding free energy reproduce the interaction patterns of known inhibitors. Starting from these fragments, CCLD generates a set of compounds that are closely related to high-affinity thrombin inhibitors. In addition, putative ligands with novel binding motifs are suggested. Probable implications of the MCSS-CCLD approach for the evolving scenario of drug discovery are discussed.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Greer J., Erickson J.W., Baldwin J.J. and Varney M.D., J. Med. Chem., 37 (1994) 1035.
Lam P.Y.S., Jadhav P.K., Eyermann C.J., Hodge C.N., Ru Y., Bacheler L.T., Meek J.L., Otto M.J., Rayner M.M., Wong Y.N., Chang C.H., Weber P.C., Jackson D.A., Sharpe T.R. and Erickson-Viitanen S.K., Science, 264 (1994) 380.
Hilpert K., Ackermann J., Banner D.W., Gast A., Gubernator K., Hadvary P., Labler L., Müller K., Schmid G., Tschopp T. and Van de Waterbeemd H., J. Med. Chem., 37 (1994) 3889.
Karplus M. and Petsko G.A., Nature, 347 (1990) 631.
Van Gunsteren W.F. and Berendsen H.J.C., Angew. Chem. Int. Ed. Engl., 29 (1990) 992.
Honig B. and Nicholls A., Science, 268 (1995) 1144.
Appelt K., Perspect. Drug Discov. Design, 1 (1993) 23.
Clore G.M. and Gronenborn A.M., Science, 252 (1991) 1390.
Fesik S.W., J. Med. Chem., 34 (1991) 2937.
Greer J., Proteins Struct. Funct. Genet., 7 (1990) 317.
Havel T.F., J. Mol. Simul., 10 (1993) 175.
Šali A. and Blundell T.L., J. Mol. Biol., 234 (1993) 779.
Caflisch A. and Karplus M., Perspect. Drug Discov. Design, 3 (1995) 51.
Miranker A. and Karplus M., Proteins Struct. Funct. Genet., 11 (1991) 29.
Stubbs M.T. and Bode W., Perspect. Drug Discov. Design, 1 (1993) 431.
Noble M.E.M., Verlinde C.L.M.J., Groendijk H., Kalk K.H., Wierenga R.K. and Hol W.G.J., J. Med. Chem., 34 (1991) 2709.
Caflisch A., Miranker A. and Karplus M., J. Med. Chem., 36 (1993) 2142.
Eisen M.B., Wiley D.C., Karplus M. and Hubbard R.E., Proteins Struct. Funct. Genet., 19 (1994) 199.
Sitkoff D., Sharp K.A. and Honig B., J. Phys. Chem., 98 (1994) 1978.
Warwicker J. and Watson H.C., J. Mol. Biol., 157 (1982) 671.
Gilson M.K. and Honig B.H., Proteins Struct. Funct. Genet., 4 (1988) 7.
Hermann R.B., J. Phys. Chem., 76 (1972) 2754.
Lee B. and Richards F.M., J. Mol. Biol., 55 (1971) 379.
Tapparelli C., Metternich R., Ehrhardt C. and Cook N.S., Trends Pharmacol. Sci., 14 (1993) 366.
Bode W., Mayr I., Baumann U., Huber R., Stone S.R. and Hofsteenge J., EMBO J., 8 (1989) 3467.
Banner D.W. and Hadvary P., J. Biol. Chem., 266 (1991) 20085.
Lyle T.A., Perspect. Drug Discov. Design, 1 (1993) 453.
Grootenhuis P.D.J. and Karplus M., J. Comput.-Aided Mol. Design, 10 (1996) 1.
Rotstein S.H. and Murcko M.A., J. Med. Chem., 36 (1993) 1700.
Bohacek R.S. and McMartin C., J. Am. Chem. Soc., 116 (1994) 5560.
Kuntz I.D., Science, 257 (1992) 1078.
Kettner C. and Shaw E., Thromb. Res., 14 (1979) 969.
Brünger A. and Karplus M., Proteins Struct. Funct. Genet., 4 (1988) 148.
Brooks B.R., Bruccoleri R.E., Olafson B.D., States D.J., Swaminathan S. and Karplus M., J. Comput. Chem., 4 (1983) 187.
Elber R. and Karplus M., J. Am. Chem. Soc., 112 (1990) 9161.
Dirac P.A.M., Proc. Cambridge Phil. Soc., 26 (1930) 376.
Hestenes M.R. and Stiefel E., J. Res. N.B.S., 49 (1952) 409.
Bashford D. and Karplus M., Biochemistry, 29 (1990) 10219.
Davis M.E., Madura J.D., Luty B.A. and McCammon J.A., Comput. Phys. Commun., 62 (1991) 187.
Press W.H., Teukolsky S.A., Vetterling W.T. and Flannery B.P., Numerical Recipes in Fortran, Cambridge University Press, Cambridge, U.K., 1992.
Davis M.E. and McCammon J.A., J. Comput. Chem., 10 (1989) 386.
Davis M.E. and McCammon J.A., J. Comput. Chem., 11 (1990) 401.
Davis M.E. and McCammon J.A., J. Comput. Chem., 12 (1991) 909.
Lim C., Bashford D. and Karplus M., J. Phys. Chem., 95 (1991) 5610.
Edmonds D.T., Rogers N.K. and Sternberg M.J.E., Mol. Phys., 52 (1984) 1487.
Mohan V., Davis M.E., McCammon J.A. and Pettitt B.M., J. Phys. Chem., 96 (1992) 6428.
Gilson M.K., Sharp K.A. and Honig B.H., J. Comput. Chem., 9 (1988) 327.
Luty B.A., Davis M.E. and McCammon J.A., J. Comput. Chem., 13 (1992) 768.
Still W.C., Tempczyk A., Hawley R.C. and Hendrickson T., J. Am. Chem. Soc., 112 (1990) 6127.
Chothia C., Nature, 248 (1974) 338.
Cabani S., Gianni P., Mollica V. and Lepori L., J. Solution Chem., 10 (1981) 563.
Maryanoff B.E., Qiu X., Padmanabhan K.P., Tulinsky A., AlmondJr. H.R., Andrade-Gordon P., Greco M.N., Kauffman J.A., Nicolaou K.C., Liu A., Brungs P. and Fusetani N., Proc. Natl. Acad. Sci. USA, 90 (1993) 8048.
Weber P.C., Lee S.L., Lewandowski F.A., Schadt M.C., Chang C.H. and Kettner C.A., Biochemistry, 34 (1995) 3750.
Tabernero L., Chang C.Y., Ohringer S., Lau W.F., Iwanowicz E.J., Han W.C., Wang T.C., Seiler S.M., Roberts D.G.M. and Sack J.S., J. Mol. Biol., 246 (1995) 14.
Gubernator K., Broger C., Bur D., Doran D.M., Gerber P.R., Müller K. and Schaumann T.M., In Hermann E.C. and Franke R. (Eds.) Computer-Aided Drug Design in Industrial Research, Springer, Berlin, Germany, 1995, pp. 61–77.
Gerber P.R. and Müller K., J. Comput.-Aided Mol. Design, 9 (1995) 251.
Obst U., Gramlich V., Diederich F., Weber L. and Banner D.W., Angew. Chem., 107 (1995) 1874.
Rydel T.J., Tulinsky A., Bode W. and Huber R., J. Mol. Biol., 221 (1991) 583.
Müller K., In Schwartz T.W., Hjorth S.A. and Sandholm Kastrup J., (Eds.) Structure and Function of 7TM Receptors, Munksgaard, Copenhagen, Denmark, 1996, pp. 414–421.
Gallop M.A., Barrett R.W., Dower W.J., Fodor S.P.A. and Gordon E.M., J. Med. Chem., 37 (1994) 1233.
Gordon E.M., Barrett R.W., Dower W.J., Fodor S.P.A. and Gallop M.A., J. Med. Chem., 37 (1994) 1385.
Weber L., Wallbaum S., Broger C. and Gubernator K., Angew. Chem. Int. Ed. Engl., 34 (1995) 2280.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Caflisch, A. Computational combinatorial ligand design: Application to human α-thrombin. J Computer-Aided Mol Des 10, 372–396 (1996). https://doi.org/10.1007/BF00124471
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00124471