Abstract
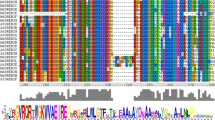
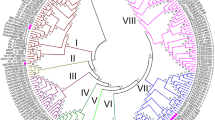
The WRKY family, a large family of transcription factors (TFs) found in higher plants, plays central roles in many aspects of biological processes and adaption to environment. However, little information is available on this family in apple (Malus domestica). In the present study, a total of 119 candidate WRKY genes in apple genome were identified and classified into three main groups (Group I–III) based on the structure of the conserved domains. Each group or subgroup showed similar exon–intron structures and motif compositions. The evolution analysis showed that 44 MdWRKY genes clustered into 20 intensive regions (<100 kb) and 78 MdWRKY formed 85 pairs of collinear relationships, suggesting that both tandem and segmental duplications played an important role in the evolution and diversification of the WRKY gene family in apple. Furthermore, the expression of the MdWRKY genes in apple leaves in response to biotic stress (Alternaria alternate) and hormone treatments [salicylic acid (SA), methyl jasmonate (MeJA) and ethephon] was examined by using RNA-seq and qRT-PCR. The results showed that 63 MdWRKY genes had differential expression in their transcript abundance in response to Alternaria alternata apple pathotype infection. Moreover, most pathogen responsive MdWRKY genes were also changed significantly when apple leaves were treated by SA, MeJA or ethephon plant growth regulations, suggesting an interaction between SA, JA and ethylene (Eth) hormone signaling under biotic stress. This work may provide the basis for future studies of the genetic modification of WRKY genes for pathogen resistance in apple.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Abe K, Iwanami H, Kotoda N, Moriya S, Takahashi Sumiyoshi S (2010) Evaluation of apple genotypes and Malus species for resistance to Alternaria blotch caused by Alternaria alternata apple pathotype using detached-leaf method. Plant Breeding 129:208–218
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7:986–995
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS. (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Bari R, Jones JDG (2009) Role of plant hormones in plant defence responses. Plant Mol Biol 69:473–488
Bulajic A, Filajdic N, Babovic M, Sutton TB (1996) First report of Alternaria mali on apples in Yugoslavia. Plant Dis 80:709
Cai B, Yang X, Tuskan GA, Cheng ZM (2011) MicroSyn: a user friendly tool for detection of microsynteny in a gene family. BMC Bioinformatics 12:79
Cai C, Niu E, Du H, Zhao L, Feng Y, Guo W (2014) Genome-wide analysis of the WRKY transcription factor gene family in Gossypium raimondii and the expression of orthologs in cultivated tetraploid cotton. The Crop Journal 2:87–101
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10
Cao Z, Zhang S, Wang R, Zhang R, Hao Y (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdoMYB121 gene confering abiotic stress tolerance in plants. PLoS ONE 8:e69955
Chang S, Jeff P, John C (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Chen C, Chen Z (2002) Potentiation of developmentally regulated plant defense response by AtWRKY18, a pathogen-induced Arabidopsis transcription factor. Plant Physiol 129:706–716
Chen C, Chen Z (2000) Isolation and characterization of two pathogenand salicylic acid-induced genes encoding WRKY DNA-binding proteins from tobacco. Plant Mol Biol 42:387–396
Chen M, Tan Q, Sun M, Li D, Fu X, Chen X, Xiao W, Li L, Gao D (2016) Genome-wide identification of WRKY family genes in peach and analysis of WRKY expression during bud dormancy. Mol Genet Genomics 291(3):1319–1332
Chen X, Liu J, Lin G, Wang A, Wang Z, Lu G (2013) Overexpression of AtWRKY28 and AtWRKY75 in Arabidopsis enhances resistance to oxalic acid and Sclerotinia sclerotiorum. Plant Cell Rep 32:1589–1599
Choi C, Park YH, Kwon SI, Yun C, Ahn I, Park SR, Hwang D (2014) Identification of AtWRKY75 as a transcriptional regulator in the defense response to Pcc through the screening of Arabidopsis activation-tagged lines. Plant Biotechnol Rep 8:183–192
Ciolkowski I, Wanke D, Birkenbihl RP, Somssich IE (2008) Studies on DNA-binding selectivity of WRKY transcription factors lend structural clues into WRKY-domain function. Plant Mol Biol 68:81–92
Dang F, Wang Y, She J, Lei Y, Liu Z, Eulgem T, Lai Y, Lin J, Yu L, Lei D, Others (2014) Overexpression of CaWRKY27, a subgroup IIe WRKY transcription factor of Capsicum annuum, positively regulates tobacco resistance to Ralstonia solanacearum infection. Physiol Plantarum 150:397–411
Dang F, Wang Y, Yu L, Eulgem T, Lai Y, Liu Z, Wang X, Qiu A, Zhang T, Lin J, Chen Y, Guan D, Cai H, Mou S, He S (2013) CaWRKY40, a WRKY protein of pepper, plays an important role in the regulation of tolerance to heat stress and resistance to Ralstonia solanacearum infection. Plant, Cell Environ 36:757–774
De Grassi A, Lanave C, Saccone C (2008) Genome duplication and gene-family evolution: the case of three OXPHOS gene families. Gene 421:1–6
Delaney TP, Uknes S, Vernooij B, Friedrich L, Weymann K, Negrotto D, Gaffney T, Gut-Rella M, Kessmann H, Ward E, Ryals J (1994) A central role of salicylic acid in plant disease resistance. Science 266:1247–1250
Dong J, Chen C, Chen Z (2003) Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol Biol 51:21–37
Duan GF, Li L J, Liu QL (2014). A wrky transcription factor from malus domestica, negatively regulates dehydration stress in transgenic arabidopsis. Acta Physiol Plant, 36(2):541–548
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:113
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206
Eulgem T, Rushton PJ, Schmelzer E, Hahlbrock K, Somssich IE (1999) Early nuclear events in plant defence signalling: rapid gene activation by WRKY transcription factors. EMBO J 18:4689–4699
Eulgem T, Somssich IE (2007) Networks of WRKY transcription factors in defense signaling. Curr Opin Plant Biol 10:366–371
Fan H, Wang F, Gao H, Wang L, Xu J, Zhao Z (2011) Pathogeninduced MdWRKY1 in ‘Qinguan’ apple enhances disease resistance. J Plant Biol 54:150–158
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi- Shinozaki K, Shinozaki K (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9:436–442
Guo C, Guo R, Xu X, Gao M, Li X, Song J, Zheng Y, Wang X (2014) Evolution and expression analysis of the grape (Vitis vinifera L.) WRKY gene family. J Exp Bot 65:1513–1528
He H, Dong Q, Shao Y, Jiang H, Zhu S, Cheng B, Xiang Y (2012) Genome-wide survey and characterization of the WRKY gene family in Populus trichocarpa. Plant Cell Rep 31:1199–1217
Huang X, Li K, Xu X, Yao Z, Jin C, Zhang S (2015) Genome-wide analysis of WRKY transcription factors in white pear (Pyrus bretschneideri) reveals evolution and patterns under drought stress. BMC Genomics 16
Ishiguro S, Nakamura K (1994) Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 5'-upstream regions of genes coding for sporamin and β-amylase from sweet potato. Mol General Genet 244:563–571
Jiang Y, Duan Y, Yin J, Ye S, Zhu J, Zhang F, Lu W, Fan D, Luo K (2014) Genome-wide identification and characterization of the Populus WRKY transcription factor family and analysis of their expression in response to biotic and abiotic stresses. J Exp Bot 65:6629–6644
Journot-Catalino N, Somssich IE, Roby D, Kroj T (2006) The transcription factors WRKY11 and WRKY17 act as negative regulators of basal resistance in Arabidopsis thaliana. Plant Cell 18:3289–3302
Li J (2004) The WRKY70 transcription factor: a node of convergence for jasmonate-mediated and salicylate-mediated signals in plant defense. Plant Cell 16:319–331
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967
Li S, Fu Q, Chen L, Huang W, Yu D (2011) Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta 233:1237–1252
Ling J, Jiang W, Zhang Y, Yu H, Mao Z, Gu X, Huang S, Xie B (2011) Genome-wide analysis of WRKY gene family in Cucumis sativus. BMC Genomics 12:471
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCt Method. Methods 25:402–408
Mangelsen E, Kilian J, Berendzen KW, Kolukisaoglu UH, Harter K, Jansson C, Wanke D (2008) Phylogenetic and comparative gene expression analysis of barley (Hordeum vulgare) WRKY transcription factor family reveals putatively retained functions between monocots and dicots. BMC Genomics 9:194
Meng D, Li Y, Bai Y, Li M, Cheng L (2016) Genome-wide identification and characterization of WRKY transcriptional factor family in apple and analysis of their responses to waterlogging and drought stress. Plant Physiol Bioch 103:71–83
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNASeq. Nat Methods 5:621–628
Murray SL, Ingle RA, Petersen LN, Denby KJ (2007) Basal resistance against Pseudomonas syringae in Arabidopsis involves WRKY53 and a protein with homology to a nematode resistance protein. Mol Plant Microbe Interact 20:1431–1438
Pavy N, Boyle B, Nelson C, Paule C, Giguere I, Caron S, Parsons LS, Dallaire N, Bedon F, Berube H, Cooke J, Mackay J (2008) Identification of conserved core xylem gene sets: conifer cDNA microarray development, transcript profiling and computational analyses. New Phytol 180:766–786
Qin Y, Tian Y, Han L, Yang X (2013) Constitutive expression of a salinity-induced wheat WRKY transcription factor enhances salinity and ionic stress tolerance in transgenic Arabidopsis thaliana. Bioch Biophy Res Commun 441:476–481
Rairdan G, Moffett P (2007) Brothers in arms? Common and contrasting themes in pathogen perception by plant NB-LRR and animal NACHT-LRR proteins. Microbes Infect 9:677–686
Ramamoorthy R, Jiang SY, Kumar N, Venkatesh PN, Ramachandran S (2008) A comprehensive transcriptional profiling of the WRKY gene family in rice under various abiotic and phytohormone treatments. Plant Cell Physiol 49:865–879
Ramiro D, Jalloul A, Petitot A, Grossi De Sá MF, Maluf MP, Fernandez D (2010) Identification of coffee WRKY transcription factor genes and expression profiling in resistance responses to pathogens. Tree Genet Genomes 6:767–781
Rinerson CI, Rabara RC, Tripathi P, Shen QJ, Rushton PJ (2015) The evolution of WRKY transcription factors. BMC Plant Biol 15:66
Robatzek S, Somssich IE (2001) A new member of the Arabidopsis WRKY transcription factor family, AtWRKY6, is associated with both senescence-and defence-related processes. Plant J 28:123–133
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends in Plant Sci 15:247–258
Ryu HS, Han M, Lee SK, Cho JI, Ryoo N, Heu S, Lee YH, Bhoo SH, Wang GL, Hahn TR, Jeon JS (2006) A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defense response. Plant Cell Rep 25:836–847
Saito A, Nakazawa N, Suzuki M (2001) Selection of mutants resistant to Alternaria blotch from in vitro-cultured apple shoots irradiated with X-and γ-rays. J Plant Physiol 158:391–400
Su H, Zhang S, Yuan X, Chen C, Wang X, Hao Y (2013) Genomewide analysis and identification of stress-responsive genes of the NAM—ATAF1, 2—CUC2 transcription factor family in apple. Plant Physiol Bioch 71:11–21
Sun C, Palmqvist S, Olsson H, Bor E N M, Ahlandsberg S, Jansson C (2003) A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell 15:2076–2092
Tang Y, Kuang J, Wang F, Chen L, Hong K, Xiao Y, Xie H, Lu W, Chen J (2013) Molecular characterization of PR and WRKY genes during SA-and MeJA-induced resistance against Colletotrichum musae in banana fruit. Postharvest Biol Technol 79:62–68
Tiwari SB, Hagen G, Guilfoyle TJ (2004) Aux/IAA proteins contain a potent transcriptional repression domain. Plant Cell 16:533–543
Tripathi P, Rabara RC, Rushton PJ (2014) A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 239:255–266
Ülker B, Mukhtar MS, Somssich IE (2007) The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta 226:125–137
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D, Salvi S, Pindo M, Baldi P, Castelletti S, Cavaiuolo M, Coppola G, Costa F, Cova V, Dal Ri A, Goremykin V, Komjanc M, Longhi S, Magnago P, Malacarne G, Malnoy M, Micheletti D, Moretto M, Perazzolli M, Si-Ammour A, Vezzulli S, Zini E, Eldredge G, Fitzgerald LM, Gutin N, Lanchbury J, Macalma T, Mitchell JT, Reid J, Wardell B, Kodira C, Chen Z, Desany B, Niazi F, Palmer M, Koepke T, Jiwan D, Schaeffer S, Krishnan V, Wu C, Chu VT, King ST, Vick J, Tao Q, Mraz A, Stormo A, Stormo K, Bogden R, Ederle D, Stella A, Vecchietti A, Kater MM, Masiero S, Lasserre P, Lespinasse Y, Allan AC, Bus V, Chagné D, Crowhurst RN, Gleave AP, Lavezzo E, Fawcett JA, Proost S, Rouzé P, Sterck L, Toppo S, Lazzari B, Hellens RP, Durel C, Gutin A, Bumgarner RE, Gardiner SE, Skolnick M, Egholm M, Van de Peer Y, Salamini F, Viola R (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42:833–839
Vision TJ, Brown DG, Tanksley SD (2000) The origins of genomic duplications in Arabidopsis. Science 290:2114–2117
Wang D, Amornsiripanitch N, Dong X (2006) A genomic approach to identify regulatory nodes in the transcriptional network of systemic acquired resistance in plants. PLoS Pathog 2:e123
Wang L, Zhu W, Fang L, Sun X, Su L, Liang Z, Wang N, Londo JP, Li S, Xin H (2014) Genome-wide identification of WRKY family genes and their response to cold stress in Vitis vinifera. BMC Plant Biol 14:103
Wang M, Vannozzi A, Wang G, Liang Y, Tornielli GB, Zenoni S, Cavallini E, Pezzotti M, Cheng ZM (2014) Genome and transcriptome analysis of the grapevine (Vitis vinifera L.) WRKY gene family. Hort Res 1:14016
Wang X, Zhang S, Su L, Liu X, Hao Y (2013) A genome-wide analysis of the LBD (LATERAL ORGAN BOUNDARIES domain) gene family in Malus domestica with a functional characterization of MdLBD11. PloS ONE 8:e57044
Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X, Lee TH, Jin H, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49
Wei W, Hu Y, Han YT, Zhang K, Zhao FL, Feng JY (2016) The WRKY transcription factors in the diploid woodland strawberry Fragaria vesca: Identification and expression analysis under biotic and abiotic stresses. Plant Physiol Biochem 105:129–144
Wu KL, Guo ZJ, Wang HH, Li J (2005) The WRKY family of transcription factors in rice and Arabidopsis and their origins. DNA Res 12:9–26
Yao Q, Xia E, Liu F, Gao L (2015) Genome-wide identification and comparative expression analysis reveal a rapid expansion and functional divergence of duplicated genes in the WRKY gene family of cabbage, Brassica oleracea var. capitata. Gene 557:35–42
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18:292–298
Zhang S, Xu R, Luo X, Jiang Z, Shu H (2013) Genome-wide identification and expression analysis of MAPK and MAPKK gene family in Malus domestica. Gene 531:377–387
Zhang Y, Lubberstedt T, Xu M (2013) The genetic and molecular basis of plant resistance to pathogens. J Genet Genomics 40:23–35
Zhang Y, Wang L (2005) The WRKY transcription factor superfamily: its origin in eukaryotes and expansion in plants. BMC Evol Biol 5:1
Zheng Z, Mosher SL, Fan B, Klessig DF, Chen Z (2007) Functional analysis of Arabidopsis WRKY25 transcription factor in plant defense against Pseudomonas syringae. BMC Plant Biol 7:2
Zheng Z, Qamar SA, Chen Z, Mengiste T (2006) Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J 48:592–605
Zhou Q, Tian A, Zou H, Xie Z, Lei G, Huang J, Wang C, Wang H, Zhang J, Chen S (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol 6:486–503
Zou C, Jiang W, Yu D (2010) Male gametophyte-specific WRKY34 transcription factor mediates cold sensitivity of mature pollen in Arabidopsis. J Exp Bot 61:3901–3914
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lui, S., Luo, C., Zhu, L. et al. Identification and expression analysis of WRKY transcription factor genes in response to fungal pathogen and hormone treatments in apple (Malus domestica). J. Plant Biol. 60, 215–230 (2017). https://doi.org/10.1007/s12374-016-0577-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-016-0577-3