Abstract
Background
In spite of the global economic significance of sheep production, little is known about the prevalence of various Sarcocystis spp. infecting the domestic sheep (Ovis aries) in Egypt.
Materials and methods
Muscle samples were collected from 175 sheep (> 2 years) slaughtered at El-Mahalla El-Kubra abattoir, Gharbia governorate, Egypt. Samples were initially examined by naked eye for the existence of macrosarcocysts. The microscopic sarcocysts were detected and identified using the light microscopy and the Transmission electron microscopy (TEM). Different microscopic species of ovine Sarcocystis were molecularly confirmed by PCR, sequence analyses and phylogeny.
Results
Preliminary light microscopic inspection of the muscle specimens revealed the existence of only the microscopic sarcocysts of Sarcocystis tenella and Sarcocystis arieticanis in 152 (86.8%) out of the175 examined animals. Sarcoysts of S.tenella had striated thick cyst wall that amounted from 3.5–5.5 μm in thickness whereas, S.arieticanis sarcocysts had a thin cyst wall that ranged from 1–3 μm in thickness. S.tenella sarcocysts were detected in 115 sheep (65.7%), and were more prevalent than those of S.arieticanis, observed only in 68 sheep (38.8%). No macroscopic sarcocysts were observed in any of the examined carcasses. Transmission electron microscopy (TEM) of the cyst wall of S.tenella revealed the existence of the short stubby villar protrusions (VP) with the characteristic disk-like structures at the tips of the (VP). While, TEM of S.arieticanis showed that the cyst wall had elongated tubular protrusions that measured approximately 5–7 μm in length. Each (VP) consisted of a dome-shaped base (0.3–0.9 μm in diameter), a relatively thick middle portion (0.1–0.3 μm) in width, and a thin hair-like distal portion that measured about (0.03 x 1–4.5 μm).
Conclusion
Comparative analyses of the sequences of the four genetic markers (18S rRNA, 28S rRNA, mitochondrial cox1 and ITS-1) for S.tenella and S.arieticanis isolates detected herein, revealed genetic variations of 95% and 95– 96% among the different isolates on the level of the 18S rRNA and 28S rRNA, respectively. Whereas, the cox1 and ITS-1 shared sequence identities of 76–78% and 70–73%, respectively. S.tenella was strongly related to S.capracanis infecting goats (Capra hircus). Sequence identity of 98% on the level of 18S rRNA, 28S rRNA genes was observed between the currently identified isolates of S.tenella and the formerly GenBank deposited isolates of S.capracanis. While, cox1 sequences shared identities of 92–93%. Furthermore, S.arieticanis isolates identified here were closely related to the formerly published sequences of S.hircicanis. The 18S rRNA and 28S rRNA sequences of S.arieticanis shared 98% and 94–95% identities with those of S.hircicanis, respectively. However, 87–88% homologies were observed between the cox1 sequences of S.arieticanis and S.hircicanis. Consequently, cox1 and ITS-1 gene sequences act as better genetic markers than 18S rRNA and 28S rRNA sequences for the characterization of ovine Sarcocystis spp. Maximum parsimony analyses based on the sequences of three genetic markers, (18S rRNA, 28S rRNA and mitochondrial cox1), yielded the same placement of the currently identified isolates of the two taxa (S.tenella and S.arieticanis) within a clade of Sarcocystis species with carnivorous animals as known, or assumed, final hosts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sarcocystis spp. are cyst-forming intracellular apicomplexan parasites with an obligatory two-host life cycle between carnivores as final hosts and prey animals usually (herbivores) as intermediate hosts. Sheep (Ovis aries) act as intermediate hosts for six Sarcocystis species, i.e., S. gigantea, S. medusiformis, S. tenella, S. arieticanis, S. microps and S. mihoensis, that can be morphologically characterized depending on variations in the ultrastructure of the sarcocyst wall. Sarcocysts of S. tenella and S. arieticanis are microscopic and transmitted by canine definitive hosts, whereas S. gigantea and S. medusiformis form macroscopic sarcocysts and transmitted by felids [10].
Natural Sarcocystis spp. infections in sheep have been studied in several countries throughout the world, with variable prevalence rates according to the methods used for sarcocyst detection [1, 10, 29, 31, 34, 48, 57]. However, to the best of our knowledge, infection rates of different Sarcocystis spp. in domestic sheep in Egypt are widely unknown. The ultrastructural features of the sarcocysts are the basic fundamentals for differentiation of the Sarcocystis spp. infecting the same intermediate host [2, 10, 12,13,14,15,16,17, 27, 28, 30, 32, 37]. Nonetheless, many Sarcocystis spp. isolated from diverse, but closely related intermediate hosts had morphological homologies resulting in perplexity concerning clarifying the relationships among the Sarcocystis spp. under investigation. Therefore, genetic markers, such as the small ribosomal subunit gene (18S rRNA), the large ribosomal subunit gene (28S rRNA), the internal transcribed spacer-1 region (ITS-1) and the mitochondrial cytochrome c oxidase subunit 1 gene (cox1), have been used for delineation or more comprehensive identification of novel Sarcocystis or already existent species in different hosts. Additionally, few genetic data of ovine Sarcocystis spp. are currently provided in GenBank [10, 29]. Therefore, the current study was performed for investigation of the prevalence and detailed morphology of the ovine Sarcocystis spp. in Egypt. Molecular identification of the detected sarcocysts was carried out utilizing PCR, sequencing and phylogenetic analyses of 18S rRNA, 28S rRNA, cox-1 genes and ITS-1 region.
Materials and Methods
Morphologic Examination of Sarcocysts
Muscle samples were collected from 175 slaughtered sheep (> 2 years). Animals were slaughtered at El-Mahalla El-Kubra abattoir, Gharbia governorate, Egypt (30°58′07″N 31°09′49″E), during the period extending from January 2017 till February 2018. The ages of the inspected animals were determined through teeth examination.
From each animal, fresh tissue samples from the esophagus, diaphragm, skeletal muscles, tongue and heart were grossly examined for the existence of macrosarcocysts. Approximately, 0.5 mm pieces of muscle from each collected sample were squeezed between two glass slides to inspect and preliminary identify the detected microscopic sarcocysts using stereomicroscopy.
For histopathologic investigations, the detected microscopic sarcocysts were fixed in 10% neutral buffered formalin, routinely processed, paraffin embedded, cut into 5 µm sections, and stained with H&E. Images were captured using OLYMPUS AX80 Microscope Digital Unit. Other microsarcocysts were isolated from muscular fibers using dissecting sterile fine needles then processed for transmission electron microscopy (TEM) and DNA analysis.
For TEM, sarcocysts were fixed in 2.5% glutaraldehyde and post-fixed in osmium tetroxide 1%, dehydrated in alcohols and finally embedded in epon-araldite mixture. Ultrathin sections were stained with uranyl acetate and lead citrate and then examined using a JEOL JEM-1400 TEM at 80 kV. For molecular investigations, the isolated sarcocysts were stored in sterile water at − 20 °C till processing.
Molecular Characterization, Sequencing and Phylogenic Analyses
Genomic DNA was extracted using QIAGEN DNeasy Tissue Kit® (QIAGEN®, GmbH, Hilden, Germany) from two morphologically different microsarcocysts according to the manufacturer’s instructions.
The 18S rRNA gene was amplified with the forward (S1) and reverse (B) primer pair [19, 36]. Primer sets KL1/KL3, KL4/KL5b and KL6/KL2 [39] were used for amplification of the 28S rRNA, while the cox1 gene was amplified with (SF1 forward) and (SR9 reverse) primers [22, 23], and finally, the ITS-1 region was amplified with primer pairs SU1F/5.8SR2 [25].
All PCR amplifications for the 18S rRNA, 28S rRNA genes and ITS-1 region were carried out according to the reactions steps and conditions previously described by El-Morsey et al. [14, 16]. PCR amplifications were performed utilizing Bio-Rad T100 thermal cycler (Bio-Rad Laboratories Inc., USA). PCRs for the (cox-1) gene were performed according to the amplification protocol formerly described by Gjerde [23].
PCR products were purified from gel using Qia Quick gel extraction kit® (QIAGEN®) according to the recommendations of the manufacturer. Two isolates, each from one sarcocyst, were directly sequenced utilizing ABI 3730XL automatic DNA sequencer (Applied Biosystems, USA).
The Basic Local Alignment Search Tool (BLAST) of the National Center for Biotechnology Information (NCBI) https://blast.ncbi.nlm.nih.gov/Blast.cgi was used to determine the identities of the 18S rRNA, the 28S rRNA, cox1 and ITS-1 sequences, with the previously GenBank deposited Sarcocystis spp. sequences. Prior to performing phylogenetic analyses, sequences were truncated slightly from both ends so that they begin and end with the same nucleotides. Sequences of the three target genes, i.e., 18S rRNA, the 28S rRNA and cox1 of Sarcocystis spp. infecting ruminant animals were retrieved from the GenBank database and aligned using the ClustalW program.
Phylogenetic analyses were conducted individually on nucleotide sequences of the 18S rRNA, 28S rRNA and cox1 using MEGA7 software [33]. Multiple alignments were generated with the ClustalW program within MEGA7, utilizing a gap opening penalty = 10 and a gap extension penalty of 0.2. Phylograms of the three gene sequences were constructed using the maximum parsimony (MP) analysis. The credibility of the (MP) trees was tested with the bootstrap method with 1000 replicates [18]. Bootstrap values are shown on the bifurcation points of the cladograms in Figs. 3, 4 and 5. The MP trees were obtained using the Tree–Bisection–Regrafting (TBR) algorithm [40].
The 18S rRNA gene evolutionary analysis involved 60 nucleotide sequences that belonged to 55 taxa (Fig. 3). There were a total of 2359 positions in the final dataset. The analysis was rooted on seven Eimeria spp. infecting chickens as outgroup. The seven Eimeria spp. 18S rRNA sequences included in the analysis were E. necatrix U67119, E. tenella U67121, E. praecox U67120, E. acervulina U67115, E. maxima DQ538348, E. mitis U40262 and E. brunetti U67116.
The 28S rRNA gene analysis involved 26 nucleotide sequences belonging to 18 Sarcocystis spp. infecting ruminant animals (Fig. 4). There were a total of 3587 nucleotide positions in the final dataset. One Eimeria sp. (E. acervulina; GU593707) was used as outgroup to root the 28S rRNA maximum parsimony tree.
The evolutionary history of the cox1 gene sequences of the microscopic ovine Sarcocystis isolates detected herein was deduced relying on the MP analysis (Fig. 5). The analysis included 39 nucleotide sequences belonging to 31 Sarcocystis spp. A total of 2908 nucleotide positions were existing in the final dataset. The MP tree was based on seven Eimeria spp. parasitizing chickens, as outgroup. The seven Eimeria spp. cox1 sequences integrated in the cladistic analysis were E. maxima HQ702481, E. mitis JN864949, E. brunetti HQ702480, E. acervulina HQ702479, E. praecox HQ702483, E. necatrix HQ702482 and E. tenella HQ702484.
Results
Prevalence of Ovine Sarcocystis spp. Natural Infections in Egypt
Only two morphologically distinct microscopic types of sarcocysts were detected in 152 out of 175 sheep (86.8%). Sarcocysts of S. tenella that have striated thick cyst wall and S. arieticanis with thin cyst wall were observed. Sarcocystis tenella sarcocysts were observed in 115 sheep (65.7%) and were more prevalent than those of S. arieticanis, found in 68 sheep (38.8%). The distribution of the two Sarcocystis spp. in different sheep muscles is shown in Table 1.
Light Microscopic and Ultrastructural Features of the Detected Sarcocysts
Utilizing light microscopy, the sarcocysts of S. tenella were microscopic and ranged from 350–1150 × 35–110 µm (n = 45) in size. The sarcocyst wall measured 3.5–5.5 µm in thickness and had numerous, very crowded palisade-like villar protrusions (VP) that appeared somewhat short or stubby (Fig. 1a). The interior of the sarcocyst was filled with crescent-shaped bradyzoites (BR) of variable dimensions that measured 9.5–14 × 3.5–5 µm (n = 35).
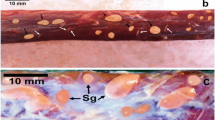
Morphologic features of S. tenella sarcocysts infecting sheep muscles from Egypt. a Light micrograph of a sarcocyst (H&E stained); showing the striated sarcocyst wall with numerous stubby palisade-like villar protrusions (VP), elongated banana-shaped bradyzoites (BR) within the cyst cavity. Scale bar = 10 µm. b Semithin section of S. tenella stained with toluidine blue 1% depicting the short stubby (VP) that are characteristic for S. tenella, ground substance (Gs), obvious septa (S), irregularly shaped lightly stained peripherally located metrocytes (Met) and elongated crescent-shaped bradyzoites (BR) Scale bar = 10 µm. c TEM micrograph depicting the sarcocyst wall with short stubby palisade-like villar protrusions (VPs). The (VPs) are characterized by their tips that have dense plaque-like structures (DP). Note the Parasitophorous vacuolar membrane (PVM), ground substance (Gs) with no obvious structures. Scale bar = 2 µm
Examination of the semithin sections of S. tenella sarcocysts revealed the existence of the distinctive short stubby upright (VP). Numerous peripherally located lightly stained metrocytes (Met) were evident under the ground substance (GS) of the cyst wall and appeared as irregularly shaped cells of different sizes. Clear septa (S) originated from the ground substance dividing the cyst into several compartments filled with myriads of banana-shaped bradyzoites (BR) (Fig. 1b).
By TEM, S. tenella cyst wall ranged from 3.5–5.5 µm in thickness and had numerous stubby and mainly upright palisade-like protrusions that amounted from 2.5–4 in length × 0.3–1.4 µm in width (n = 25). The (VP) were characterized by the existence of dense plaques (DP) at their tips; however, microtubules were missing (Fig. 1c). The ground substances (Gs) measured 0.9–1.8 µm in thickness and was situated immediately under the primary sarcocyst wall. No structures were existent within the ground substance. The ultrastructural characteristics of the cyst wall of S. tenella belonged to type 14 according to the classification of Dubey et al. [10].
By histopathological examination, S. arieticanis sarcocysts were microscopic and measured 125–985 × 30–85 µm (n = 45) in size. The cyst wall appeared smooth and thin and measured approximately from 1 to 3 µm in thickness (Fig. 2a). However, examination of the semithin sections showed that the sarcocyst wall had numerous fine hair-like protrusions that were bent nearly 90º at their middle portions then became parallel to the direction of the cyst wall surface. Prominent undulations or uneven surface were a characteristic feature of the outer surface of the cyst wall in some locations (Fig. 2b).
Morphologic characters of S. arieticanis cysts infecting sheep muscles from Egypt. a Light micrograph showing the histopathology of a sarcocyst (H&E stained); notice the smooth appearance of the thin sarcocyst wall (CW), the existence of crescent-shaped bradyzoites (BR). Scale bar = 10 µm. b Semithin section of S. arieticanis; note the hair-like villar protrusions (VP) with the prominent undulations or uneven surface of the cyst wall in some locations, ground substance (GS), very evident septa (S), and banana-like bradyzoites (BR). Scale bar = 10 µm. c, d TEM micrographs depicting the ultrastructure of the cyst wall of S. arieticanis that is characterized by the elongated tubular protrusions (VP) that consists of dome-shaped bases, relatively thick middle portions and hair-like distal ends. d Some locations of the cyst wall, the (VPs) appeared as non-branched bone-like structures. In addition, the parasitophorous vacuolar membrane (PVM) appeared highly convoluted or corrugated in the dome-shaped base of the villi forming electron dense indentations. Prominent septa (S) originated from the ground substance (Gs) dividing the cyst into different chambers containing lightly stained metrocytes (Met) and Bradyzoites (BR). Scale bars = 1 µm and 500 nm for (c) and (d), respectively
Ultrastructurally, the cyst wall of S. arieticanis had elongated tubular (VP) that measured approximately 5–7 µm in length. Each (VP) consisted of a dome-shaped base (0.3–0.9 µm in diameter), a relatively thick middle portion (0.1–0.3 µm) and a thin hair-like distal portion that measured about (0.03 × 1–4.5 µm). Sometimes, the hair-like distal regions of the (VP) were highly convoluted forming conical tufts that were deeply embedded inside the cytoplasm of the host cells. In addition, the parasitophorous vacuolar membrane (PVM) appeared highly corrugated in the dome-shaped base of the villi forming electron dense indentations. Just above the dome-shaped base, the protrusions turned 90° to the cyst wall so that the middle and distal portions became nearly parallel to the surface of the sarcocyst. The ground substance (Gs) ranged from 0.9 to 1.5 µm in thickness and was located immediately under the primary sarcocyst wall. In some locations of the cyst wall, the (VP) appeared as non-branched bone-like structures. Obvious septa (S) were found dividing the sarcocysts of S. arieticanis into chambers containing ovoid or irregularly shaped metrocytes (Met) of various diameters and bradyzoites (BR) that measured 8.5–12 × 2–3.5 µm (n = 55) in size (Fig 2c, d). Features of the sarcocyst wall of S. arieticanis described herein belonged to cyst wall type 7a [10].
DNA and Comparative Analyses of the Sequences
Characterization of the 18S rRNA Sequences
The two 18S rRNA nucleotide sequences of S. tenella were 1828 bp in length and were completely identical. Hence, only one sequence (MH413034) of isolate (1 ST) was submitted to GenBank. Each sequence was obtained from an individual sarcocyst. Running the obtained sequence on the BLAST of the GenBank revealed 99% similarity with those of S. tenella (KC209734, KC209737 and MF039329) reported from sheep and the sequences (KP263752 to KP263759) isolated from the chamois (Rupicapra rupicapra). Homology of 98% was observed with the sequences of S. capracanis (L76472, KU820982 and KU820983) reported from goats (Capra hircus).
Sarcocystis arieticanis 18S rRNA sequences (MH413035; 1SA and MH413036; 2SA) were 1840 bp in length. Each sequence was obtained from an individual sarcocyst of S. arieticanis. The differences between the two isolates comprised two nucleotide substitutions. A similarity of 96% was shared between the new S. arieticanis 18S RNA sequences and that of S. tenella (MF039329) infecting sheep from China, whereas the most homologous sequences (sharing 99% identities), were those of S. arieticanis under accessions MF039330 and MF039331. Sarcocystis arieticanis (L24382) and S. hircicanis (KU820984; KU820985) showed 98% identity with the isolates of S. arieticanis identified in the current study.
The phylogenetic tree of the partial 18S rRNA sequences (Fig. 3), showed that the isolate of S. tenella obtained in the current study (MH413034; isolate 1 ST), was placed within a well-supported clade formed by S. tenella (MF039329 and KP263759). Meanwhile, the three isolates were strongly related together with S. capracanis (L76472), S. heydorni (KX057997), S. alces (KF831274) and S. gracilis (KY019031) forming a major group. On the other hand, sequences of the two isolates of S. arieticanis (MH413035; 1SA and MH413036; 2SA) were situated inside another robustly supported clade that comprised isolates of S. arieticanis (MF039331, L24382) and S. hircicanis (KU820985).
Phylogram of the maximum parsimony analysis of the 18S rRNA sequences of the Sarcocystis spp. infecting ruminant animals depicting the robust association of the currently identified isolates of S. tenella and S. arieticanis from domestic sheep with the formerly detected isolates of the same species. Bootstrap values are the percentages located at the bifurcation points of the tree. The newly identified Sarcocystis isolates are marked by black circles (MH413034 for S. tenella isolate 1ST, while MH413035 and MH413036 for S. arieticanis isolates 1SA, 2SA, respectively). The evolutionary analysis was conducted in MEGA7 using 7 Eimeria spp. of chickens as outgroup
Characterization of the 28S rRNA Sequences
The 28S rRNA nucleotide sequences of S. tenella were deposited in GenBank under accessions (MH413037 for isolate 2aST and MH413038 for isolate 2bST). Each sequence was derived from an individual sarcocyst of S. tenella. Both sequences were 3465 bp in length. Variations between the two isolates involved five nucleotide substitutions.
The most homologous sequences deposited in GenBank were those of S. tenella (AF076899 reported from Australia and MF039325; MF039326 isolated from China) with 99% identity, followed by the sequences of S. capracanis (AF012885, KU820978 and KU820979) (sharing 98% identity), S. arieticanis (MF039327; MF039328) (96% identity), S. cruzi (AF076903) and S. arieticanis (AF076904) (95% identity) and S. hircicanis (KU820980 and KU820981) (94% identity).
The 28S rRNA sequences of the two S. arieticanis isolates (1aSA and 1bSA), each obtained from an individual sarcocyst, were accepted in GenBank under accession numbers (MH413039 and MH413040), respectively. Isolate (1aSA) was 3492 bp long, whereas isolate (1bSA) was 3515 bp in length. The differences between the sequences of the two isolates comprised 35 nucleotide substitutions.
The similarity of S. arieticanis 28S rRNA sequences obtained herein was 97% with S. tenella isolate (MF039326) and 96% with S. tenella isolates (AF076899) and (MF039325). The highly similar sequences in GenBank were those of S. arieticanis (MF039327) (sharing 99% identity), followed by S. arieticanis (MF039328) (showing 98% identity), whereas 97% homology was observed with S. arieticanis (AF076904). Homology of 96% was observed with S. capracanis (AF012885, KU820978; KU820979), while S. cruzi (AF076903) and S. hircicanis isolates (KU820980; KU820981) shared 94–95% identities.
As a result, 28S rRNA sequence identities of 99% were shared between the currently identified isolates of S. tenella and the previously GenBank deposited sequences of the same species. In the same way, a homology of 97%–99% was observed between S. arieticanis isolates detected herein, and those published on GenBank and belonging to the same taxon.
Collectively, sequence similarities ranging from 94 to 99% on the level of the 28S rRNA gene were shared between the presently identified isolates of both S. tenella and S. arieticanis and the previously GenBank deposited sequences of S. tenella, S. arieticanis, S. hircicanis, S. capracanis and S. cruzi. Furthermore, sequence identities on the level of 28S rRNA gene varying from 95 to 96% were observed among the currently detected isolates of S. tenella and S. arieticanis. The nucleotide differences between the newly identified isolates of both species involved approximately 140–175 nucleotide substitutions.
The phylogram of the 28S rRNA sequences (Fig. 4) placed the two recently detected isolates of S. tenella (MH413037; 2aST and MH413038; 2bST) inside a highly supported group including the previously GenBank deposited isolates (MF039325, MF039326, AF076899) of S. tenella and S. capracanis (KU820978). Furthermore, S. arieticanis isolates (1aSA MH413039 and 1bSA MH413040) were located within a clade containing the formerly GenBank released sequences of S. arieticanis (MF039327, MF039328 and AF076904).
A phylogram showing the evolutionary history of members of the Sarcocystidae that was inferred using the maximum parsimony method. Bootstrap values are shown on the nodes of the cladogram. The recently identified isolates of S. tenella and S. arieticanis were marked by black circles. 28S rRNA sequences of S. tenella were deposited in GenBank with accessions (MH413037; 2aST and MH413038; 2bST) while S. arieticanis sequences had accession numbers (MH413039; 1aSA and MH413040; 1bSA). The phylogenic analysis was conducted using the program MEGA7. The tree was rooted on E. acervulina (GU593707) as outgroup
Characterization of the cox1 Sequences
The (cox1) nucleotide sequences (MH413045 and MH413046), each from a single sarcocyst of S. tenella, were 1035 bp long and shared 99% similarity. The variation between them comprised eight nucleotide substitutions.
The two isolates of S. tenella, MH413045 for isolate (1cST) and MH413046 for isolate (2cST), shared the highest homologies with those of S. tenella MF039322 (99%) and MF039323 (98%) infecting domestic sheep from China. They shared identities of 97% with those of S. tenella (KC209725–KC209729; KC209731; KC209732) isolated from sheep, S. tenella (KP263744, KP263746, KP263747, KP263748, KP263749) reported from chamois (Rupicapra rupicapra tatrica). Identities of 96% were observed with isolates of S. tenella from sheep (KC209723; KC209730; KC209724) and chamois (Rupicapra rupicapra tatrica) (KP263750; KP263745; KP263751). On the other hand, S. capracanis (KU820974; KU820977) shared similarities of 92–93% with the current isolates, whereas 90% homology was found between the isolates reported in the present study and those of S. heydorni (KX057994; KX057995).
The (cox1) nucleotide sequences (MH413047; isolate 1CSA and MH413048; isolate 2CSA), each from a single sarcocyst of S. arieticanis, were 1040 bp long, and showed 99% homology. The variations between them comprised 11 nucleotide substitutions.
Sarcocystis arieticanis cox1 sequence of domestic sheep under accession MF039324 shared the highest identity (95%) with the present isolates (MH413047; isolate 1CSA and MH413048; isolate 2CSA).
Similarities of 87–88% were observed between the isolates of S. arieticanis reported in the current study and those of S. hircicanis (KU820975 and KU820976), while 80–82% homologies were observed with the isolates of S. grueneri (KC209615–KC209624) infecting the reindeer. Sequences of cox-1 of S. capreolicanis (KY018938–KY018944) infecting the roe deer (Capreolus capreolus) shared 79–80% similarities.
Sarcocystis cruzi sequences (KC209599; KT901095; KT901079; KT901090; LC171859) shared 79% identities with the isolates of S. arieticanis reported herein, while isolates (KT901084; LC171862; LC171861) that belong to the same species showed 80% homology. Finally, 78% similarity was observed with S. cruzi under accessions (KC209597; KT901094; KT901088; KT901085; KT901083; KT901078).
The cox1 nucleotide sequence homologies between the isolates of S. arieticanis and S. tenella detected herein, varied from 76 to 78%. The nucleotide differences included approximately 230–247 nucleotide substitutions.
The cladistic analysis depending on the cox1 sequences (Fig. 5), placed the two new isolates of S. tenella (1cST; MH413045 and 2cST; MH413046) in a well-supported clade comprising the sequences of S. tenella (MF039322, MF039323, KC209727 and KP263749), whereas the S. arieticanis isolates (1CSA; MH413047 and 2CSA; MH413048) were grouped with S. arieticanis (MF039324).
A cladogram showing the evolutionary history of the Sarcocystis spp. of the Sarcocystidae family that was inferred using the maximum parsimony method. Bootstrap values are shown on the bifurcation points of the cladogram. The newly detected isolates of the cox1 sequences of (S. tenella and S. arieticanis) with accession numbers (MH413045; MH413046 and MH413047; MH413048), respectively, were marked by black circles. The phylogenic tree was constructed using MEGA7 software. Seven Eimeria spp. cox-1 sequences were used as outgroup
Characterization of the ITS-1 Sequences
Sarcocystis tenella ITS-1 nucleotide sequences were deposited in GenBank under accession numbers (MH413041) for isolate (1iST) and (MH413042) for isolate (2iST). Each isolate was derived from a single sarcocyst. The first isolate (1iST) was 791 bp, while the second (2iST) was 795 bp in length. The identity between the two isolates was 99%. The differences between both isolates comprised 10 nucleotide substitutions. When the ITS-1 sequences S. tenella and S. arieticanis were compared with sequences previously released on GenBank, a similarity of 96% was observed between the isolates identified herein and S. tenella (MF039318) infecting domestic sheep from China, whereas isolate (MF039319) of S. tenella showed 93% homology.
The ITS-1 sequences MH413043 (1iSA) and MH413044 (2iSA), each from one sarcocyst of S. arieticanis, were 789 bp and 785 bp long, respectively. The similarity between them was 98%, and the variations involved 15 nucleotide substitutions. A homology of 97% was observed between the current isolates and S. arieticanis (MF039320), while isolate (MF039321) of S. arieticanis shared 94% similarity.
The identities between ITS-1 nucleotide sequences reported herein for S. tenella and S. arieticanis were ranging from 70 to 73%. The nucleotide variations involved approximately 115–251 nucleotide substitutions.
Discussion
Sarcocystis species are highly prevalent apicomplexan parasites in domestic animals and some of them may have significant economic impacts particularly, when causing clinical and subclinical disease [10].
No macroscopic Sarcocystis spp. were detected in the examined sheep carcasses in the current study. Only two distinct microscopic Sarcocystis species were found in 152 out of 175 sheep (86.8%). Sarcocystis tenella sarcocysts were detected in 115 sheep (65.7%) and were more prevalent than those of S. arieticanis, observed in 68 sheep (38.8%).
To our knowledge, Sarcocystis species infecting sheep were not thoroughly investigated from Egypt in the previous studies. Only a single study, Mahran [35], who detected both macroscopic and microscopic sarcocysts in 229 (41.26%) out of 555 sheep slaughtered at Shalatin abattoir, Red Sea governorate. Depending only on the macroscopic and histologic examination of the collected muscle samples, the microscopic S. tenella sarcocysts were detected in 81.1% of the investigated sheep. Additionally, macroscopic sarcocysts were found in (9.9%) of the examined carcasses.
Prevalences of ovine Sarcocystis spp. as low as 9% to approximately 100% have been observed in several former investigations [10, 29]. Variations in the prevalence of Sarcocystis spp. infecting sheep depend on many factors like, cessation of life cycle, management conditions, existence of stray dogs and cats in close proximity to sheep and habits of final and intermediate hosts [10].
Sarcocystis tenella cyst wall morphologic characters observed in the current study were consistent with the ultrastructural features of the cyst wall type-14 according to the classification of Dubey et al. [10]. Similar cyst wall features have been detected in Sarcocystis spp. infecting other ruminant hosts, as those of S. tenella in domestic sheep [1, 29, 55], sarcocysts of S. pseudois from the Himalayan blue sheep (Pseudois nayaur) reported by Odening [42], sarcocysts of S. gazellae infecting the springbok (Antidorcas marsupialis) that were recorded by Odening et al. [43], S. cf. capracanis isolated from the blackbuck (Antilope cervicapra) [53], Sarcocystis tenella-like sarcocysts detected in wild sheep (Ovis musimon) by Nigro et al. [41], a Sarcocystis sp. parasitizing the chamois [44], S. capracanis-like cysts from the alpine ibex (Capra ibex) that were observed by Cornaglia et al. [6], S. capracanis observed in goats from Japan [52], S. capracanis sarcocysts described in goats (Capra hircus) from Philippines [4] and S. capracanis sarcocysts infecting domestic goats in Egypt [38].
Sarcocystis capracanis of the domestic goats (Capra hircus) is a sister species of S. tenella, and the two species have the same morphologic features under light microscope. Nonetheless, there are some distinctive variations in the sarcocyst wall ultrastructure, i.e., the existence of electron dense disk-like structures or condensations in the tips of the (VPs) of S. tenella, whereas vesicles were observed at the bases of S. capracanis (VPs).
The morphology of S. arieticanis observed in the current study belonged to the cyst wall (type-7a) according to the classification of Dubey et al. [10]. The ultrastructural features described herein, are homologous to some extent with those of S. arieticanis identified from sheep slaughtered in USA [9], S. arieticanis infecting sheep from China [29], S. hircicanis sarcocysts infecting domesticated goats from China [31], and sarcocysts of S. hircicanis from the Japanese goats that were reported by Saito et al. [51].
Sarcocystis spp. having the same ultrastructural characteristics have been reported from other domestic and wild hosts, like; S. arieticanis parasitizing the domestic sheep [46], S. cruzi described by Claveria et al. [5] from cattle, sarcocysts of S. rangi infecting reindeer identified by Gjerde [21], a Sarcocystis species infecting the springbok reported by Odening et al. [43], sarcocysts of S. levinei infecting the water buffalo (Bubalus bubalis) reported by Claveria and Cruz [3], a Sarcocystis sp. parasitizing the Mongolian gazelle (Procapra gutturosa) recorded by Odening et al. [45], S. cf. cruzi reported from the Nilgai (Boselaphus tragocamelus) by Stolte et al. [53], S. arieticanis-like sarcocysts detected from the Alpine ibex by Cornaglia et al. [6], Sarcocystis sp. number II found in the raccoon (Procyon lotor) [54], and Sarcocystis species number 1 isolated from the European hare (Lepus europaeus) reported by Odening et al. [47].
The two distinct sarcocyst wall morphologic types (14 and 7a) detected in the current study for (S. tenella and S. arieticanis), respectively, have been also identified in various but closely related intermediate hosts; nonetheless, the relationships among these Sarcocystis species were not clear. Additionally, previous investigations using only the 18S rRNA gene sequences for the characterization of Sarcocystis spp. infections in several ruminant intermediate hosts have led to controversy concerning the exact identity of some taxa of Sarcocystis a result of the great identities (> 99%) between the sequences of the 18S rRNA genes. For example, detection of the bradyzoites of S. capracanis in the cerebrospinal fluids from sheep [11, 20], S. cruzi infections in either water buffalo or cattle [56, 58] and several Sarcocystis spp. infections detected in cervids such as (reindeer and red deer, moose and red deer) [7, 8], or (moose and roe deer) Gjerde [24]. Therefore, there was a necessity to differentiate or re-evaluate descriptions of existing or recently discovered taxa of Sarcocystis in different hosts, using different genetic markers for delineating their phylogenetic relationships.
In the present study, the sequences of (18S rRNA, 28S rRNA, cox1 genes and the ITS-1 region) for the microsarcocysts of S. tenella and S. arieticanis in Egyptian sheep were characterized. When comparing these sequences with the previously released sequences in GenBank, sequences of 18S rRNA and 28S rRNA for S. tenella shared similarity percentages of 98% with those of S. capracanis, whereas 92–93% homologies were observed on the level of cox1 sequences. On the other hand, S. hircicanis showed homologies of 98%, 94–95%, and 87–88% with the currently identified isolates of S. arieticanis on the levels of 18S rRNA, 28S rRNA and cox1 genes, respectively.
High identities among the sequences of 18S rRNA genes were observed herein and in former studies [1, 11, 20, 29,30,31,32, 49, 50, 56, 58] as a result of the slowly evolving character of such target gene [22]. Moreover, sequence analysis utilizing one gene marker may be inadequate as some Sarcocystis species have more intra-species sequences variations in a particular region as elucidated recently [26]. Consequently, the (cox1) gene sequences appeared to act as better genetic markers for Sarcocystis spp. than 18S rRNA and 28S rRNA for discrimination of the sarcocysts of S. tenella from that of S. capracanis, and S. arieticanis from those of S. hircicanis.
The comparison of the four sequences, i: e (18S rRNA, 28S rRNA, cox1, and ITS-1) for the isolates of S. tenella and S. arieticanis detected in the current investigation, showed similarities of 95%, 95–96%, 76–78%, and 70–73%, respectively. Accordingly, the cox1 gene and the ITS-1 sequences could be more precise than the 18S rRNA and 28S rRNA genes for distinguishing the closely related Sarcocystis spp. infecting the same intermediate host because of their high divergence.
The cladistic analyses based on the sequences of the three genetic markers (18S rRNA, 28S rRNA and cox1) grouped the newly detected isolates of S. tenella and S. arieticanis inside a clade consisting of Sarcocystis spp. having carnivores as the known, or assumed, final hosts.
In conclusion, the current investigation is the first to demonstrate the presence of the microscopic Sarcocystis spp., i.e., (S. tenella and S. arieticanis), in domestic sheep slaughtered for human consumption in Egypt depending on comprehensive ultrastructural and molecular identification of the isolated sarcocysts.
References
Bittencourt MV, Meneses IDS, Ribeiro-Andrade M, de Jesus RF, de Araújo FR, Gondim LFP (2016) Sarcocystis spp. in sheep and goats: frequency of infection and species identification by morphological, ultrastructural, and molecular tests in Bahia, Brazil. Parasitol Res 115:1683–1689
Choi TI, Hong EJ, Ryu SY, Sim C, Chae JS, Kim HC, Park J, Choi KS, Yu DH, Yoo JG, Park BK (2018) Detection and identification of Sarcocystis cruzi (Protozoa: Apicomplexa) by molecular and ultrastructural studies in naturally infected Korean Cattle (Bos taurus coreanae) from Daejeon, Korea. Korean J Parasitol 56:121–127
Claveria FG, Cruz MJ (2000) Sarcocystis levinei infection in Philippine water buffaloes (Bubalus bubalis). Parasitol Int 48:243–247. https://doi.org/10.1016/S1383-5769(99)00025-2
Claveria FG, San Pedro-Lim MR, Tan JE, Flores-Cruz MJ (2004) Sarcocystis capracanis infection in Philippine domestic goats (Capra hircus): ultrastructural studies. Philipp J Sci 133:33–37
Claveria FG, San-Pedro LR, Cruz-Flores MJ, Nagasawa H, Suzuki N, De La Pena C (2001) Ultrastructural studies of Sarcocystis cruzi (Hasselmann, 1926) Wenyon, 1926 infection in cattle (Bos taurus): Philippine cases. Parasite 8:251–254
Cornaglia E, Giaccherino AR, Peracino V (1998) Ultrastructural morphology of sarcosporidiosis in alpine ibex (Capra ibex). Vet Parasitol 75:21–32. https://doi.org/10.1016/S0304-4017(97)00185-4
Dahlgren SS, Gjerde B (2010) Molecular characterization of five Sarcocystis species in red deer (Cervus elaphus), including Sarcocystis hjorti n. sp., reveals that these species are not intermediate host specific. Parasitology 137:815–840
Dahlgren SS, Gjerde B (2010) The red fox (Vulpes vulpes) and the arctic fox (Vulpes lagopus) are definitive hosts of Sarcocystis alces and Sarcocystis hjorti from moose (Alces alces). Parasitology 137:1547–1557
Dubey JP, Lindsay DS, Speer CA, Fayer R, Livingston CW (1988) Sarcocystis arieticanis and other Sarcocystis Species in sheep in the United States. J Parasitol 74(6):1033–1038
Dubey JP, Calero-Bernal R, Rosenthal BM, Speer CA, Fayer R (2016) Sarcocystosis of animals and humans, 2nd edn. CRC Press, Boca Raton
Dubey JP, Rosenthal BM (2013) Sarcocystis capracanis-associated encephalitis in sheep. Vet Parasitol 197:407–408. https://doi.org/10.1016/j.vetpar.2013.04.027
El-Morsey A (2010) Studies on Sarcocystis species infecting water buffaloes in Egypt. M.D. thesis Parasitology Dept. Fac. of .Vet. Med. KafrElsheikh University, Egypt, pp 1–94
El-Morsey A (2016) Studies on Sarcocystis species infecting wild and migratory birds. Ph.D. thesis Parasitology Dept. Fac. of .Vet. Med. KafrElsheikh University, Egypt, pp 1–115
El-Morsey A, El-Seify M, Desouky AR, Abdel-Aziz MM, El-Dakhly KM, Kasem S, Abdo W, Haridy M, Sakai H, Yanai T (2015) Morphologic and molecular characteristics of Sarcocystis atraii n. sp. (Apicomplexa: Sarcocystidae) infecting the common coot (Fulica atra) from Egypt. Acta Parasitol 60:691–699. https://doi.org/10.1515/ap-2015-0098
El-Morsey A, El-Seify M, Desouky AY, Abdel-Aziz MM, Sakai H, Yanai T (2014) Morphologic identification of a new Sarcocystis sp. in the common moorhen (Gallinula chloropus) (Aves: Gruiformes: Rallidae) from Brolos Lake, Egypt. Parasitol Res 113:391–397. https://doi.org/10.1007/s00436-013-3667-x
El-Morsey A, EL-Seify M, Desouky AY, Abdel-Aziz MM, Sakai H, Yanai T (2015) Sarcocystis chloropusae (protozoa: Sarcocystidae) n. sp. from the common moorhen (Gallinula chloropus) from Egypt. Parasitology 142:1063–1065. https://doi.org/10.1017/s0031182015000293
El-Seify M, El-Morsey A, Hilali M, Zayed A, El-Dakhly K, Haridy M, Sakai H, Yanai T (2014) Molecular characterization of Sarcocystis fusiformis and Sarcocystis buffalonis infecting water buffaloes (Bubalus bubalis) from Egypt. Am J Anim Vet Sci 9:95–104. https://doi.org/10.3844/ajavssp.2014.95.104
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fischer S, Odening K (1998) Characterization of bovine Sarcocystis species by analysis of their 18S ribosomal DNA sequences. J Parasitol 84:50–54. https://doi.org/10.2307/3284529
Formisano P, Aldridge B, Alony Y, Beekhuis L, Davies E, Del Pozo J, Dunn K, English K, Morrison L, Sargison N, Seguino A, Summers BA, Wilson D, Milne E, Beard PM (2013) Identification of Sarcocystis capracanis in cerebrospinal fluid from sheep with neurological disease. Vet Parasitol 193:252–255. https://doi.org/10.1016/j.vetpar.2012.12.016
Gjerde B (1985) Ultrastructure of the cysts of Sarcocystis rangi from skeletal muscle of reindeer (Rangifer tarandus tarandus). Rangifer 5:43–52
Gjerde B (2013) Phylogenetic relationships among Sarcocystis species in cervids, cattle and sheep inferred from the mitochondrial cytochrome c oxidase subunit I gene. Int J Parasitol 43:579–591. https://doi.org/10.1016/j.ijpara.2013.02.004
Gjerde B (2014) Sarcocystis species in red deer revisited: with a redescription of two known species as Sarcocystis elongata n. sp. and Sarcocystis truncata n. sp. based on mitochondrial cox1 sequences. Parasitology 141:441–452. https://doi.org/10.1017/S0031182013001819
Gjerde B (2012) Morphological and molecular characterization and phylogenetic placement of Sarcocystis capreolicanis and Sarcocystis silva n. sp. from roe deer (Capreolus capreolus) in Norway. Parasitol Res 110:1225–1237. https://doi.org/10.1007/s00436-011-2619-6
Gjerde B (2014) Molecular characterization of Sarcocystis rileyi from a common eider (Somateria mollissima) in Norway. Parasitol Res 113:3501–3509
Gjerde B, Josefsen TD (2015) Molecular characterisation of Sarcocystis lutrae n. sp. and Toxoplasma gondii from the musculature of two Eurasian otters (Lutra lutra) in Norway. Parasitol Res 114:873–886. https://doi.org/10.1007/s00436-014-4251-8
Gual I, Bartley PM, Katzer F, Innes EA, Canton GJ, Moore DP (2017) Molecular confirmation of Sarcocystis gigantea in a naturally infected sheep in Argentina: a case report. Vet Parasitol 248:25–27
Hilali M, El-Seify M, Zayed A, El-Morsey A, Dubey JP (2011) Sarcocystis dubeyi (Huong and Uggla 1999) infection in water buffaloes (Bubalus bubalis) from Egypt. J Parasitol 97:527–528. https://doi.org/10.1645/GE-2656.1
Hu JJ, Huang S, Wen T, Esch GW, Liang Y, Li HL (2017) Sarcocystis spp. in domestic sheep in Kunming City, China: prevalence, morphology, and molecular characteristics. Parasite 24:30. https://doi.org/10.1051/parasite/2017025
Hu JJ, Huang S, Wen T, Esch GW, Liang Y, Li HL (2017) Morphology, molecular characteristics, and demonstration of a definitive host for Sarcocystis rommeli from cattle (Bos taurus) in China. J Parasitol 103:471–476
Hu JJ, Liu TT, Liu Q, Esch GW, Chen JQ, Huang S, Wen T (2016) Prevalence, morphology, and molecular characteristics of Sarcocystis spp. in domestic goats (Capra hircus) from Kunming, China. Parasitol Res 115:3973–3981
Hu JJ, Wen T, Chen XW, Lu TT, Esch GW, Huang S (2016) Prevalence, morphology, and molecular characterization of Sarcocystis heydorni Sarcocysts from cattle (Bos Taurus) in China. J Parasitol 102:545–548
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Maca O (2018) Molecular identification of Sarcocystis lutrae in the European otter (Lutra lutra) and the European badger (Meles meles) from the Czech Republic. Parasitol Res 117(3):943–945
Mahran OM (2009) Sarcocystis infection in sheep and goats slaughtered in Shalatin abattoir, Red Sea Governorate, Egypt. Assiut Vet Med J 55(121):1–15
Medlin L, Elwood HJ, Stickel S, Sogin ML (1988) The characterization of enzymatically eukaryotic 16S-like rRNA-coding regions. Gene 71:491–499. https://doi.org/10.1016/0378-1119(88)90066-2
Morsy K, Abdel-Ghaffar F, Dajem SB, Abdel-Gaber R, El Gazar F (2018) First molecular characterization and morphological aspects of Sarcocystis fusiformis infecting water buffalo (Bubalus bubalis) in Egypt. Acta Parasitol 63(2):333–345
Morsy K, Saleh A, Al-Ghamdi A, Abdel-Ghaffar F, Al-Rasheid K, Bashtar AR, Al Quraishy S, Mehlhorn H (2011) Prevalence pattern and biology of Sarcocystis capracanis infection in the Egyptian goats: a light and ultrastructural study. Vet Parasitol 181:75–82. https://doi.org/10.1016/j.vetpar.2011.05.010
Mugridge NB, Morrison DA, Heckeroth AR, Johnson AM, Tenter AM (1999) Phylogenetic analysis based on full-length large subunit ribosomal RNA gene sequence comparison reveals that Neospora caninum is more closely related to Hammondia heydorni than to Toxoplasma gondii. Int J Parasitol 29:1545–1556
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nigro M, Mancianti F, Rossetti P, Poli A (1991) Ultrastructure of the cyst and life cycle of Sarcocystis sp. from wild sheep (Ovis musimon). J Wildl Dis 27:217–224
Odening K (1998) The present state of species-systematics in Sarcocystis Lankester, 1882 (Protista, Sporozoa, Coccidia). Syst Parasitol 41:209–233. https://doi.org/10.1023/A:1006090232343
Odening K, Rudolph M, Quandt S, Bengis RG, Bockhardt I, Viertel D (1998) Sarcocystis spp. in antelopes from southern Africa. Acta Protozool 37:149–158
Odening K, Stolte M, Bockhardt I (1996) On the diagnostics of Sarcocystis in chamois (Rupicapra rupicapra). Appl Parasitol 37:153–160
Odening K, Stolte M, Lux E, Bockhardt I (1996) The Mongolian gazelle (Procapra gutturosa, Bovidae) as an intermediate host of three Sarcocystis species in Mongolia. Appl Parasitol 37:54–65
Odening K, Stolte M, Walter G, Bockhardt I (1995) Cyst wall ultrastructure of two Sarcocystis spp. from European mouflon (Ovis ammon musimon) in Germany compared with domestic sheep. J Wildl Dis 31:550–554. https://doi.org/10.7589/0090-3558-31.4.550
Odening K, Wesemeier HH, Pinkowski M, Walter G, Sedlaczek J, Bockhardt I (1994) European hare and European rabbit (Lagomorpha) as intermediate hosts of Sarcocystis species (Sporozoa) in central Europe. Acta Protozool 33:177–189
Phythian CJ, Jackson B, Bell R, Citer L, Barwell R, Windsor PA (2018) Abattoir surveillance of Sarcocystis spp., Cysticercosis ovis and Echinococcus granulosus in Tasmanian slaughter sheep, 2007–2013. Aust Vet J 96:62–68. https://doi.org/10.1111/avj.12670
Prakas P, Butkauskas D, Rudaitytė E, Kutkienė L, Sruoga A, Pūraitė I (2016) Morphological and molecular characterization of Sarcocystis taeniata and Sarcocystis pilosa n. sp. from the sika deer (Cervus nippon) in Lithuania. Parasitol Res 115:3021–3032. https://doi.org/10.1007/s00436-016-5057-7
Prakas P, Rudaitytė E, Butkauskas D, Kutkienė L (2017) Sarcocystis entzerothi n. sp. from the European roe deer (Capreolus capreolus). Parasitol Res 116:271–279. https://doi.org/10.1007/s00436-016-5288-7
Saito M, Shibata Y, Itagaki H (1995) Sarcocystis capracanis and S. hircicanis from goats in Japan. Jpn J Parasitol 44:391–395
Saito M, Shibata Y, Kobayashi T, Kobayashi M, Kubo M, Itagaki H (1996) Ultrastructure of the cyst wall of Sarcocystis species with canine final host in Japan. J Vet Med Sci 58:861–867. https://doi.org/10.1292/jvms.58.861
Stolte M, Odening K, Bockhardt I (1996) Antelopes (Bovidae) kept in European zoological gardens as intermediate hosts of Sarcocystis species. Parassitologia 38:565–570
Stolte M, Odening K, Walter G, Bockhardt I (1996) The raccoon as intermediate host of three Sarcocystis species in Europe. J Helminthol Soc Wash 63:145–149
Vlemmas I, Kanakoudis G, Tsangaris T, Theodorides I, Kaldrymidou E (1989) Ultrastructure of Sarcocystis tenella (Sarcocystis ovicanis). Vet Parasitol 33:207–217. https://doi.org/10.1016/0304-4017(89)90130-1
Xiang Z, He Y, Zhao H, Rosenthal BM, Dunams DB, Li X, Zuo Y, Feng G, Cui L, Yang Z (2011) Sarcocystis cruzi: comparative studies confirm natural infections of buffaloes. Exp Parasitol 127:460–466. https://doi.org/10.1016/j.exppara.2010.10.012
Yang Y, Dong H, Su R, Wang Y, Wang R, Jiang Y, Tong Z (2018) High prevalence of Sarcocystis spp. infections in cattle (Bos taurus) from central China. Parasitol Int 67:800–804. https://doi.org/10.1016/j.parint.2018.08.006
Yang ZQ, Zuo YX, Yao YG, Chen XW, Yang GC, Zhang YP (2001) Analysis of the 18S rRNA genes of Sarcocystis species suggests that the morphologically similar organisms from cattle and water buffalo should be considered the same species. Mol Biochem Parasitol 115:283–288. https://doi.org/10.1016/S0166-6851(01)00283-3
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest associated with this manuscript.
Ethical Approval
Parasite collection from the examined animals was carried out according to the regulatory laws and ethical considerations regarding experimental ethics of animal use and collecting permits.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
El-Morsey, A., Abdo, W., Sultan, K. et al. Ultrastructural and Molecular Identification of the sarcocysts of Sarcocystis tenella and Sarcocystis arieticanis Infecting Domestic Sheep (Ovis aries) from Egypt. Acta Parasit. 64, 501–513 (2019). https://doi.org/10.2478/s11686-019-00070-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11686-019-00070-8