Abstract
Background
The loss of PBRM1 expression (as identified by immunohistochemistry) is associated with a high risk of postoperative recurrence for patients with clear cell renal cell carcinoma (ccRCC). The authors developed a scoring system to predict recurrence based on clinicopathologic factors incorporating PBRM1 expression.
Methods
This study retrospectively reviewed 479 ccRCC patients who underwent radical surgery between 2006 and 2017. The study extracted a subset of 389 non-metastatic ccRCC patients for whom relevant clinicopathologic factors were available. The primary end point was recurrence-free survival (RFS). The Kaplan–Meier method and the Cox proportional hazards model were used for statistical analysis. Leibovich score, SSIGN score, and University of California, Los Angeles (UCLA) Integrated Staging System were included as conventional prediction models.
Results
Of the 389 patients, 53 (13.6%) experienced recurrence during a median period of 61 months. Multivariable analyses showed that that the independent factors for RFS were ≥ pT3 (hazard ratio [HR] 3.64; P < 0.001), sarcomatoid or rhabdoid component (HR 3.29; P = 0.005), PBRM1 negativity (HR 3.39; P = 0.001), and necrosis (HR 3.60; P < 0.001). A scoring system calculated with these factors, named the SSPN (stage, sarcomatoid, PBRM1 expression, and necrosis) score, showed significant differences in RFS among the following four groups; low-risk group (0 factors), intermediate-risk group (1 factor), high-risk group (2 to 3 factors), and very high-risk group (4 factors) (P < 0.001). The authors’ model also showed a greater predictive accuracy for 5-year RFS than the conventional models (0.841 vs 0.747–0.792).
Conclusions
The SSPN score, which integrates clinicopathologic findings and PBRM1 expression, can accurately predict postoperative recurrence for patients with non-metastatic ccRCC after radical surgery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Renal cell carcinoma (RCC) is newly diagnosed for an estimated 403,000 people and causes approximately 175,000 deaths annually.1 Clear cell renal cell carcinoma (ccRCC), the predominant histologic subtype of kidney cancer, accounts for approximately 75% of RCCs.2
Surgical resection, such as radical or partial nephrectomy, is the definitive treatment for localized ccRCC. However, 30% of patients who undergo radical surgery experience recurrence.3 Although several models for predicting ccRCC recurrence and mortality using clinicopathologic findings have been reported,4,5,6 a prognostic model that integrates molecular factors is needed to improve predictive accuracy.
Recently, several groups have investigated the molecular pathogenesis of ccRCC using whole-genome and exome sequencing. In ccRCC, PBRM1, located on chromosome 3p21, is a commonly mutated gene (~ 40% to 50%).
Detection of PBRM1 mutation on the basis of immunohistochemistry (IHC)-based assays has proved to be highly reliable.7 Some studies have shown that loss of PBRM1 expression detected by IHC is associated with a high risk of recurrence for ccRCC patients after surgery.8,9 To date, no prognostic model has incorporated PBRM1 expression for recurrence after radical surgery in ccRCC. Therefore, this study verified whether PBRM1 expression detected by IHC could be a significant biomarker for predicting recurrence in patients with non-metastatic ccRCC. Furthermore, we developed a novel scoring system combining standard clinicopathologic factors and PBRM1 expression to accurately predict recurrence of non-metastatic ccRCC. Finally, we validated the predictive accuracy of our scoring system compared with existing models.4,5,6
Materials and methods
Patent Selection
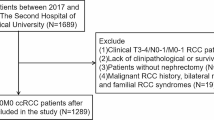
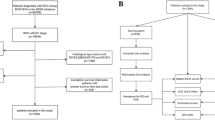
The medical records of 476 patients who underwent radical or partial nephrectomy for ccRCC at Kansai Medical University Hospital between January 2006 and December 2017 were retrospectively reviewed. Of these 476 patients, 87 were excluded from this study for the following reasons: synchronous or metachronous bilateral tumors, presurgical or adjuvant treatment with tyrosine kinase inhibitors, death due to operation-related complications, presence of metastasis, and insufficient pathologic materials (Fig. S1). Ultimately, 389 non-metastatic ccRCC (cT1-4N0-2M0) patients were analyzed in the current study, which was approved by the institutional review board of KMU Hospital (approval no. 2018109).
Clinical Data Collection
The following clinical data were obtained from the patients’ medical records: age at the time of surgery, sex (male/female), body mass index, and prognostic data such as recurrence and cancer-specific mortality. Recurrence was considered to have occurred if imaging showed new local or distant lesions. Lesions adjacent to the resection site were considered to indicate local recurrence, and those in distant organs were considered to indicate distant recurrence.
All the patients underwent preoperative imaging, including computed tomography (CT) or magnetic resonance imaging (MRI), for evaluation of the clinical stage and the presence of local or distant metastasis. All the patients were postoperatively followed up for more than 1 year, with the initial visit after 3 to 6 months, followed by semi-annual or annual visits. Laboratory testing and CT imaging were performed at each visit.
Pathologic Examination
All pathologic specimens were rereviewed and reclassified by a single urologic pathologist (C.O.) who was blinded to the clinical outcome, as previously described.10 Histologic features including subtype (clear cell/non-clear cell), the World Health Organization/International Society of Urological Pathology (WHO/ISUP) grade, the sarcomatoid or rhabdoid component (SC/RC), coagulative tumor necrosis, and lymphovascular invasion (LVI) were assessed according to the ISUP guidelines and the WHO classification.11 The pT stage was classified according to the UICC/AJCC 8th edition tumor-node-metastasis (TNM) staging system.12
Tissue Microarray (TMA) Construction and IHC of PBRM1
Construction of TMA from 2-mm cores of a formalin-fixed, paraffin-embedded tissue block was performed. Two representative locations (the highest and the most common grade areas) were identified in the hematoxylin and eosin-stained slides, and two tumor cores were sampled from each case.
All retrieved tissues for the TMA were reviewed to confirm whether the intended areas were accurately included. Immunohistochemistry staining was performed on 4-µm-thick sections sliced from TMA blocks using the Ventana Discovery Ultra Autostainer (Roche Diagnostics, Indianapolis, IN, USA). A primary antibody against PBRM1 (rabbit polyclonal, dilution 1:200; Atlas Antibodies AB, Bromma, Sweden) was used according to the manufacturer’s protocols. Antibody reactivity was visualized with OptiView DAB IHC detection and the OptiView amplification kit (Ventana Medical System, Tucson, AZ, USA).
The PBRM1 nuclear staining of the tumor cells was evaluated as positive (diffuse and strong [Fig. 1A and B]; focal weak [Fig. 1C and D]) or negative (lack of staining [Fig. 1E and F]), referring to the internal positive controls (PBRM1-positive for lymphocytes, stromal fibroblasts, and/or endothelial cells) as previously described.9 Two TMA cores were evaluated for each case. Tumor heterogeneity was considered negative if a tumor showed different staining patterns between two cores (positive/negative)0.9 In cases with only one TMA core available (n = 64), the result was used and considered positive or negative based on the presence or absence of PBRM1 expression. All IHC slides were reviewed independently by two authors (C.O., H.O.) (kappa statistic, 0.877), with the final determination made by consensus.
Representative immunohistochemical expression of PBRM1. A,B Tumor cells expressing diffuse and strong nuclear staining of PBRM1: positive. C,D Tumor cells showing focally strong or weak nuclear staining of PBRM1: positive. E,F Tumor cells showing lack of nuclear staining of PBRM1 with accurate internal control: negative. Scale bars indicate (A,C,E) 500 µm and (B,D,F) 20 µm. Original magnification × 2 and × 400, respectively
Statistical Analysis
The primary outcome measure was recurrence-free survival (RFS), defined as the time from surgery to initial recurrence shown on imaging. The secondary outcome measure was cancer-specific survival (CSS). All continuous data are shown as median values and interquartile ranges (IQRs). Survival analysis was assessed using the Kaplan–Meier method with the log-rank test and the Cox proportional hazards model. The associations among clinicopathologic factors were analyzed using the logistic regression model. Backward step-down selection was used to create most of the informative scoring system for predicting RFS with the fewest variables (reduced models). Multiple comparisons were performed with Bonferroni correction. The predictive accuracies of the models were analyzed using the area under the receiver operating characteristic curve. Hazard ratios (HRs) estimated from the Cox analyses were reported as relative risk with corresponding 95% confidence intervals (CIs). All statistical analyses were performed using EZR version 1.29 (Saitama Medical Center, Jichi, Japan)13 A two-sided p value lower than 0.05 was considered statistically significant.
Results
Patient Characteristics
The clinicopathologic findings are shown in Table 1. Of the 389 patients, 53 (13.6%) experienced recurrence, and 14 (3.6%) died of ccRCC during a median follow-up period of 61 months (IQR, 33.3–93.8 months). The median time to recurrence after surgery was 31.5 months (IQR, 13.1–49.3 months). Immunohistochemistry showed 161 patients (41.4%) to be PBRM1-negative. Positive surgical margins were not observed in any pathologic specimen.
Association Between PBRM1 Expression and Clinicopathologic Characteristics
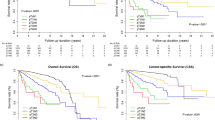
The Kaplan–Meier survival analysis showed that the 5-year RFS rate was significantly worse for the PBRM1-negative patients than for the PBRM1-positive patients (74.3% vs 96.1%; P < 0.001; Fig. 2). Similarly, the 5-year CSS rate was significantly worse for the PBRM1-negative patients than for the PBRM1-positive patients (93.9% vs 100%; P < 0.001; Fig. S2).
The correlations of standard pathologic factors with PBRM1 expression were assessed using logistic regression analysis (Table S1). The findings showed that PBRM1 negativity was strongly associated with WHO/ISUP grade (odds ratio [OR], 1.91), LVI (OR, 4.09), and pT stage (OR, 3.10) (all P < 0.05).
Uni- and Multivariable Models Predicting Recurrence
The associations between clinicopathologic factors and recurrence after nephrectomy are shown in Table 2. In the univariable analysis, pT stage, WHO/ISUP grade, SC/RC, LVI, necrosis, and PBRM1 negativity were found to be significantly associated with recurrence (all P < 0.001). These variables then were incorporated into the multivariable analysis, which identified pT stage (HR 3.64; P < 0.001), SC/RC (HR 3.29; P = 0.005), necrosis (HR 3.60; P < 0.001), and PBRM1 negativity (HR 3.39; P = 0.001) as independent prognostic factors for RFS, with a c-index of 0.843.
Development of a Novel Scoring System for Predicting Recurrence
A reduced model incorporating four readily available factors was used to create a scoring system. Because all the HRs were relatively similar, the score was assigned as follows: pT stage (< pT3, 0; ≥ pT3, 1), SC/RC (absent, 0; present, 1), PBRM1 (positive, 0; negative, 1), and necrosis (absent, 0; present, 1). The probability of RFS clearly stratified by the accumulated score is shown in Fig. 3A (P < 0.001). The scoring system is named the SSPN score to emphasize the features on which it is based (stage, sarcomatoid, PBRM1 expression, and necrosis).
A Recurrence-free survival (RFS) for all patients stratified by the accumulated score with pT stage (< pT3, 0; ≥ pT3, 1), SC/RC (absent, 0; present, 1), necrosis (absent, 0; present, 1), and PBRM1 (positive, 0; negative, 1). B The novel scoring system integrating the low-risk group (0 factors), the intermediate-risk group (1 factor), the high-risk group (2–3 factors), and the very high-risk group (4 factors). Bonferroni correction was used for multiple comparisons of the RFS among the risk groups. SC/RC, World Health Organization/International Society of Urological Pathology; CI, confidence interval; HR, hazard ratio; NE, not estimated
Because RFS did not differ significantly between groups of two and three factors (P = 0.338), the two groups were combined, resulting in a scoring system constructed with four risk groups: a low-risk group (0 factors), an intermediate-risk group (1 factor), a high-risk group (2–3 factors), and a very high-risk group (4 factors), with respective 5-year RFS rates of 97.4%, 90.8%, 65.5%, and 0.0% (P < 0.001; Fig. 3B). The high-risk group had 70 PBRM1-negative patients (93.3%), and the very high-risk group had 11 PBRM1-negative patients (100%). The 5-year CSS rates based on our scoring system were 100% in both the low- and intermediate-risk groups, 91.2% in the high-risk group, and 67.3% in the very high-risk group (P < 0.001; Fig. S3).
Comparison of Accuracy Between Our Scoring System and Conventional Risk Models
Finally, we compared the predictive accuracy of 5-year RFS between our model and conventional risk models based on the Leibovich score,4 tumor stage, size, grade and necrosis (SSIGN) score,5 and the University of California Los Angeles Integrated Staging System.6 To assess these models, data on their performance status, tumor size, and Fuhrman grade were additionally collected from patients’ medical records (Table S2). All the Kaplan–Meier survival curves based on the conventional risk models showed clearly discriminated RFS rates (Fig. S4 A–C).
Regarding the predictive ability of 5-year RFS, our model showed a higher area under the receiver operating characteristic curve than conventional risk models (0.841 vs 0.747–0.792) in our cohort (Fig. 4). < F4>
Discussion
The current study demonstrated that PBRM1 negativity detected by IHC was a significant factor in predicting the risk of postoperative recurrence for patients with non-metastatic ccRCC. Additionally, PBRM1 expression was correlated with pathologic features such as the WHO/ISUP grade, LVI, and pT stage. The SSPN score, a novel scoring system integrating four readily available factors, had the best accuracy for predicting postoperative recurrence compared with existing risk models.4,5,–6 Thus, our model can provide prognostic information for patient counseling and improve clinical decision-making regarding candidates for adjuvant therapy or clinical trials with novel agents after radical surgery for ccRCC. The acronym SSPN means “scoring system incorporating PBRM1 to predict recurrence after nephrectomy,” which is the concept of this study.
Technological advances in genome sequencing have enabled identification of novel driver genes mutated in renal cancer, such as PBRM1 (~ 50%), BAP1 (~ 15%), and SETD2 (~ 15%) 14 Specifically, PBRM1 encodes BRG1-associated factor 180 (BAF180), a subunit of the SWI/SNF chromatin remodeling complex that affects DNA transcription by altering chromatin structure.15 Mutations of PBRM1 and BAP1, also located on chromosome 3p21, are largely mutually exclusive in ccRCC and define biologically distinct ccRCC subtypes.7 Cancer genome atlas research has demonstrated that the inactivation of PBRM1 comprises a second major mutation in ccRCC tumorigenesis, followed by VHL.16 In addition, siRNA-mediated knockdown of wild-type PBRM1 significantly increases proliferation in several ccRCC cell lines.17 Therefore, PBRM1 is considered a key driver gene of ccRCC, making it a strong prognostic factor and a novel therapeutic target.
A significant correlation between PBRM1 gene mutation and PBRM1 protein deficiency detected by IHC has been reported in several previous studies.7,18 Our PBRM1-negative patients accounted for 41.4% of the entire cohort, which corresponds to the previously reported frequency of PBRM1 mutation in ccRCC,7,14,16 Regarding the association of pathologic features with PBRM1 expression, Pawłowski et al.18 and Bihr et al.19 found that advanced pT stage, nuclear grade, and LVI were associated with the PBRM1-negative status in univariable analyses. Our study confirmed that this is true even in multivariable analysis. Notably, we found that LVI, which may be a precursor of micrometastasis, was a particularly independent factor for the presence of PBRM1 in primary ccRCC. Thus, PBRM1-negative ccRCC has a greater potential to infiltrate the lymphovascular space, which supports the evidence that PBRM1 deficiency is related to tumor metastasis.
Some reports have shown that PBRM1-deficient tumors are more likely to recur after a radical surgery.8,9 Da Costa et al.8 analyzed 112 ccRCC patients and found that the PBRM1-negative status was significantly associated with a poor RFS rate compared with PBRM1-positive status (66.7% vs. 87.3%, P = 0.048). However, it did not remain as an independent factor for RFS (P = 0.575), ly because of small cohorts. Joseph et al.9 found that the PBRM1-negative status was an independent factor for metastasis-free survival (HR 1.50; P = 0.0025) when they adjusted for age and SSIGN score in 1330 ccRCC patients.
The current study also found that PBRM1 expression significantly discriminated RFS rates (Fig. 1) and was an independent factor in the multivariable analysis (HR 3.39; P = 0.001). Combined with standard pathologic factors, PBRM1 expression improved the model’s c-index, from 0.819 to 0.843. Accordingly, we have successfully established a novel risk stratification model for predicting recurrence after surgery, and it is superior to existing typical models4,5,–6 in terms of discrimination ability.
Recent studies have focused on gene mutations as biomarkers for predicting response to systemic RCC therapy. Two previous studies that investigated whether PBRM1 alteration is a marker of response to anti-PD-1 treatment suggested that patients with ccRCC harboring PBRM1-LOF had significantly better survival than those with intact PBRM1. 20,21 However, because gene mutations can be used as a marker only to some extent, complementing it with IHC assay is recommended to offer a stronger biomarker for more effective medical treatment in current clinical practice.22 According to our preliminary data investigating the efficacy of salvage treatment with nivolumab (n = 9), the PBRM1-negative patients had a better response than the PBRM1-positive patients (Fig. S5A). Moreover, the maximum change from baseline in tumor burden was − 16.3% for the PBRM1-negative patients and + 55.9% for the PBRM1-positive patients (P = 0.095; Fig. S5B). Although tumor heterogeneity between the primary and metastatic sites has been reported,23 these results suggest that the PBRM1-negative status may be a surrogate marker reflecting better efficacy of ICI treatment in preventing recurrence. Further investigations with a larger cohort are required to support our findings.
Systemic adjuvant therapy is considered to improve the outcome for RCC patients at high risk of recurrence. Four clinical trials investigating the role of tyrosine kinase inhibitors in the adjuvant setting resulted in unfavorable clinical outcomes, except the Sunitinib Treatment of Renal Adjuvant Cancer (S-TRAC) trial.24,25,26,–27 Adjuvant ICI treatment is therefore expected to develop as a new therapeutic strategy. Four clinical trials have already been conducted.28 No clinical risk predictive model is currently intended to select patients who are candidates for ICI adjuvant therapy. Interestingly, our scoring system may be suitable for such selection because in the current study, 94.2% (81/86) of the patients in the high- and very high-risk groups had PBRM1-negative ccRCC, indicating that they possibly benefit from ICIs, according to previous studies.20,21 Although external validation of this model is required, our model may provide clinicians with useful information regarding patients’ prognoses and candidates for ICI adjuvant therapy to improve oncologic outcomes.
Our results should be interpreted with caution because of several limitations. First, this study was a retrospective observational investigation. Second, the surgeries involved different procedures and were performed by different surgeons, which could have influenced the oncologic outcomes. Third, our results should be externally validated with other cohorts (e.g., at other institutions where the procedure is performed and with other races of patients). Fourth, PBRM1 expression was evaluated using only TMA and not the whole section, which may have caused unidentified bias. Finally, we did not evaluate genetic correlation with PBRM1 expression detected by IHC because such investigations already exist.7,14,17 Despite these limitations, we believe our results add new evidence to the management for ccRCC in this era of ICI treatment.
Conclusion
We present a new scoring system to predict recurrence after radical surgery using four significant factors based on standard pathologic findings and PBRM1 expression detected by IHC. Our model may therefore improve the prediction of oncologic outcomes for patients with ccRCC and may subsequently facilitate shared clinical decision-making (regarding candidates for adjuvant therapy) for patients with non-metastatic ccRCC after radical surgery.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. https://doi.org/10.3322/caac.21492.
Shuch B, Amin A, Armstrong AJ, et al. Understanding pathologic variants of renal cell carcinoma: distilling therapeutic opportunities from biologic complexity. Eur Urol. 2015; 67: 85–97. https://doi.org/10.1016/j.eururo.2014.04.029.
Eggener SE, Yossepowitch O, Pettus JA, Snyder ME, Motzer RH, Russo P. Renal cell carcinoma recurrence after nephrectomy for localized disease: predicting survival from time of recurrence. J Clin Oncol. 2006;24:3101–6. https://doi.org/10.1200/jco.2005.04.8280.
Leibovich BC, Blute ML, Cheville JC, et al. Prediction of progression after radical nephrectomy for patients with clear cell renal cell carcinoma: a stratification tool for prospective clinical trials. Cancer. 2003;97:1663–71. https://doi.org/10.1002/cncr.11234.
Frank I, Blute ML, Cheville JC, Lohse CM, Weaver AL, Zincke H. An outcome prediction model for patients with clear cell renal cell carcinoma treated with radical nephrectomy based on tumor stage, size, grade, and necrosis: the SSIGN score. J Urol. 2002;168:2395–400. https://doi.org/10.1097/01.ju.0000035885.91935.d5.
Zisman A, Pantuck AJ, Dorey F, et al. Improved prognostication of renal cell carcinoma using an integrated staging system. J Clin Oncol. 2001;19:1649–57. https://doi.org/10.1200/jco.2001.19.6.1649.
Peña-Llopis S, Vega-Rubín-de-Celis S, Liao A, et al. BAP1 loss defines a new class of renal cell carcinoma. Nat Genet. 2012;44:751–9. https://doi.org/10.1038/ng.2323.
Da Costa WH, Rezende M, Carneiro FC, et al. Polybromo‐1 (PBRM 1), a SWI/SNF complex subunit is a prognostic marker in clear cell renal cell carcinoma. BJU Int. 2014;113:E157–63. https://doi.org/10.1111/bju.12426.
Joseph RW, Kapur P, Serie DJ, et al. Clear cell renal cell carcinoma subtypes identified by BAP1 and PBRM1 expression. J Urol. 2016;195:180–7. https://doi.org/10.1016/j.juro.2015.07.113.
Yoshida T, Ohe C, Tsuzuki T, et al. Clinical impact of segmental renal vein invasion on recurrence in patients with clinical T1 renal cell carcinoma undergoing partial nephrectomy. Int J Clin Oncol. 2020;25:464–71. https://doi.org/10.1007/s10147-019-01543-6.
Moch H, Humphrey PA, Ulbright TM, et al. WHO Classification of Tumours of the Urinary System and Male Genital Organs. 4th ed. IARC, Lyon, 2016.
Brierley JD, Gospodarowics MK, Wittekind C. Union for International Cancer Control. TNM Classification of Malignant Tumours. 8th ed. Wiley, New York, 2017.
Kanda Y. Investigation of the freely available easy-to-use software ‘EZR’for medical statistics. Bone Marrow Transplant. 2013;48:452–8.
Brugarolas J. PBRM1 and BAP1 as novel targets for renal cell carcinoma. Cancer J. 2013; 19: 324–32. https://doi.org/10.1097/ppo.0b013e3182a102d1.
Reisman D, Glaros S, Thompson EA. The SWI/SNF complex and cancer. Oncogene. 2009;28:1653–68. https://doi.org/10.1038/onc.2009.4.
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature. 2013;499:43–9. https://doi.org/10.1038/nature12222.
Varela I, Tarpey P, Raine K, et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature. 2011;469:539–42. https://doi.org/10.1038/nature09639.
Pawłowski R, Mühl SM, Sulser T, Krek W, Moch H, Schraml P. Loss of PBRM1 expression is associated with renal cell carcinoma progression. Int J Cancer. 2013;132:E11–7. https://doi.org/10.1002/ijc.27822.
Bihr S, Ohashi R, Moore AL, et al. Expression and mutation patterns of PBRM1, BAP1, and SETD2 mirror specific evolutionary subtypes in clear cell renal cell carcinoma. Neoplasia. 2019;21:247–56. https://doi.org/10.1016/j.neo.2018.12.006.
Miao D, Margolis CA, Gao W, et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science. 2018;359:801–6. https://doi.org/10.1126/science.aan5951.
Braun DA, Ishii Y, Walsh AM, et al. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 2019;5:1631–3. https://doi.org/10.1001/jamaoncol.2019.3158.
Lu S, Stein JE, Rimm DL, et al. Comparison of biomarker modalities for predicting response to PD-1/PD-L1 checkpoint blockade: a systematic review and meta-analysis. JAMA Oncol. 2019;5:1195–204. https://doi.org/10.1001/jamaoncol.2019.1549.
Serie DJ, Joseph RW, Cheville JC, et al. Clear cell type A and B molecular subtypes in metastatic clear cell renal cell carcinoma: tumor heterogeneity and aggressiveness. Eur Urol. 2017;71:979–85. https://doi.org/10.1016/j.eururo.2016.11.018.
Ravaud A, Motzer RJ, Pandha HS, et al. Adjuvant sunitinib in high-risk renal-cell carcinoma after nephrectomy. N Engl J Med. 2016;375:2246–54. https://doi.org/10.1056/nejmoa1611406.
Haas NB, Manola J, Uzzo RG, et al. Adjuvant sunitinib or sorafenib for high-risk, non-metastatic renal-cell carcinoma (ECOG-ACRIN E2805): a double-blind, placebo-controlled, randomised, phase 3 trial. Lancet. 2016;387:2008–16. https://doi.org/10.1016/s0140-6736(16)00559-6.
Motzer RJ, Haas NB, Donskov F, et al. Randomized phase III trial of adjuvant pazopanib versus placebo after nephrectomy in patients with localized or locally advanced renal cell carcinoma. J Clin Oncol. 2017;35:3916–23. https://doi.org/10.1200/jco.2017.73.5324.
Gross-Goupil M, Kwon TG, Eto M, et al. Axitinib versus placebo as an adjuvant treatment of renal cell carcinoma: results from the phase III, randomized ATLAS trial. Ann Oncol. 2018;29:2371–8. https://doi.org/10.1093/annonc/mdy454.
Gul A, Rini BI. Adjuvant therapy in renal cell carcinoma. Cancer. 2019;125:2935–44. https://doi.org/10.1002/cncr.32144.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
Chisato Ohe received research funding from Chugai Pharmaceutical Co. Ltd. for work outside the subject matter discussed in the manuscript. The remaining authors have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ohsugi, H., Yoshida, T., Ohe, C. et al. The SSPN Score, a Novel Scoring System Incorporating PBRM1 Expression, Predicts Postoperative Recurrence for Patients with Non-metastatic Clear Cell Renal Cell Carcinoma. Ann Surg Oncol 28, 2359–2366 (2021). https://doi.org/10.1245/s10434-020-09075-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-020-09075-4