Abstract
LINC00467 is an example of long intergenic non-coding RNAs whose roles in human disorders are being identified. This gene coding LINC00467 is located on chromosome 1: 211,382,736 − 211,435,570 forward strand. This lncRNA has been firstly recognized through a microarray-based lncRNA profiling as an N-Myc target in neuroblastoma cells. Further studies have shown up-regulation of LINC00467 in different cancer including those originated from brain, gastrointestinal tract, lung and breast. It acts as a molecular sponge for miR-339, miR-138-5p, miR-107, miR-133b, miR-451a, miR-485-5p, miR-7-5p, miR-485-5p, miR-339-3p, miR-200a, miR-1285-3p, miR-299-5p, miR-509-3p, miR-18a-5p, miR-9-5p and miR-20b-5p. LINC00467 can regulate activity of NF-κB, STAT1, Wnt/b-catenin, Akt and ERK1/2 signaling pathways. Accumulating evidence indicates oncogenic role of LINC00467. The current review article aims at providing an overview of LINC00467 in the carcinogenesis.
Similar content being viewed by others
Introduction
Long non-coding RNAs (lncRNAs) are a group of transcripts having sizes larger than 200 nt. They are regarded as important epigenetic regulators that control epigenetic mechanisms principally in the nucleus, modulating transcription of genes through changing histone or DNA methylation and acetylation marks [1]. The majority of identified lncRNAs are transcribed by RNA polymerase II, thus having similar structures with mRNAs. While sharing many features with mRNAs, these widely expressed transcripts have distinct roles from mRNAs. Notably, function of lncRNAs is related with their particular subcellular localization [2]. In addition to modulation of chromatin function, lncRNAs can influence establishment and functions of nuclear bodies, change mRNAs stability and their translation and affect activity of signaling pathways [2].
GENECODE catalog of lncRNAs have classified these transcript into distinct categories of long intergenic non-coding (linc)-RNAs, antisense transcripts, intronic, and non-overlapping antisense transcripts [3]. LINC00467 is an example of the first group of lncRNAs whose roles in human disorders are being identified. This transcript is encoded by a gene located on chromosome 1: 211,382,736 − 211,435,570 forward strand. This gene has 28 transcripts with sizes ranging from 3536 bp (LINC00467-201) to 469 bp (LINC00467-204).
This lncRNA has been firstly recognized as an N-Myc target in neuroblastoma cells through a microarray-based transcriptome profiling [4]. Further studies have indicated abnormal expression of LINC00467 in a wide variety of cancer cell lines and clinical samples. Moreover, several studies have assessed functional roles of LINC00467 in xenograft models of cancers. The current review article aims at providing an overview of LINC00467 in the carcinogenesis through summarization of three mentioned lines of evidence.
Cell line studies
Sponging effects of LINC00467
Expression of LINC00467 has been shown to be elevated in acute myeloid leukemia (AML) cell lines. LINC00467 silencing has inhibited the malignant features of these cells. Notably, expression of miR-339 has been up-regulated after LINC00467 silencing. Moreover, expression of miR-339 target gene SKI has been decreased following this intervention. Since miR-339 silencing can chiefly eliminate the impact of LINC00467 silencing in AML cell lines, miR-339/SKI axis has been proposed as the molecular axis mediating the effects of LINC00467 [5].
In breast cancer cells, LINC00467 silencing has impeded proliferation, migratory potential, invasive features and epithelial-to-mesenchymal transition (EMT), while its up-regulation has led to opposite impacts. LINC00467 could down-regulate miR-138-5p through functioning as a molecular sponge for this miRNA. Moreover, LINC00467 could enhance expression of LIN28B through directly interacting with it [6]. Another in silico study in breast cancer has shown possible role of LINC00467 in the regulation of peroxisomal lipid metabolism and immune response through targeting miRNAs [7].
In cervical cancer cells, expression assays have detected high expression of LINC00467 and KIF23, and down-regulation of miR-107. LINC00467 has been shown to be mainly localized in the cytoplasm, where it acts as a molecular sponge for miR-107. LINC00467 silencing or miR-107 overexpression has blocked proliferation and decreased migration, invasion, and EMT [8].
In squamous cell carcinoma cells, LINC00467 can also enhance EMT through influencing activity of miR-299‐5p/USP48 axis [9].
Moreover, LINC00467 can influence response of hepatocellular cancer cells to Axitinib via acting as a molecular sponge for miR-509-3p and enhancing expression of PDGFRA [10]. miR-18a‐5p/NEDD9 [11] and miR-9-5p/PPARA [12] molecular axes are other routes of participation of LINC00467 in the pathoetiology of hepatocellular carcinoma as revealed through in vitro assays.
In osteosarcoma cells, LINC00467 has been shown to sponge miR‑217 and increase expression of KPNA4 [13] which facilitates progression of this type of cancer. Moreover, the sponging effect of LINC00467 on this miRNA leads to up-regulation of HMGA1 which enhances growth and metastatic abilities of these cells [14].
LINC00467 has also been shown to increase proliferation of lung adenocarcinoma cells through influencing miR-20b-5p/CCND1 activity [15]. Moreover, LINC00467 increases stemness of lung cancer cells through sequestering miR-4779 and miR‐7978 [16].
Association of LINC00467 with transcription factors
Experiments in bladder cancer cells have shown the role of LINC00467 in enhancement of proliferation and invasive properties of these cells. Mechanistically, LINC00467 directly binds to NF-kb-p65 transcript, enhances its stability and promotes its nuclear translocation for further activation of the NF-κB signaling [17].
SiRNA-mediated LINC00467 silencing has suppressed proliferation, invasiveness and metastatic potential of colorectal cancer cells. Mechanistically, LINC00467 could affect expression of Cyclins D1 and A1, CDK2, CDK4, Twist1 and E‑cadherin [18].
LINC00467 can also promote invasive properties and block apoptosis of squamous cell carcinoma cells through sponging miR-1285-3p and enhancing expression of TFAP2A [19]. In hepatocellular carcinoma cells, LINC00467 has been shown to bind with IGF2BP3 and stabilize TRAF5, thus promoting proliferation and metastatic abilities of these cells [20].
Upstream regulators of LINC00467
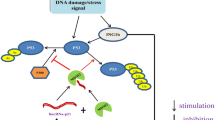
Expression of LINC00467 has been shown to be suppressed by N-Myc. In fact, N-Myc directly binds to the promoter of LINC00467 gene, decreasing its promoter activity. N-Myc has also inhibited expression of the down-stream gene of LINC00467, i.e. RD3 via directly binding to its promoter (Fig. 1). SiRNA-mediated silencing of LINC00467 has led to up-regulation of the tumor suppressor gene DKK1. This intervention has also decreased viability of neuroblastoma cells and increased their apoptosis. Notably, co-transfection of LINC00467 siRNA and DKK1 siRNA has blocked the effect of LINC00467 silencing [4].
Table 1 shows function of LINC00467 in cell lines derived from different types of cancers.
Mouse studies
Up-regulation of LINC00467 has enhanced breast cancer growth, whereas its silencing has inhibited lung metastases in vivo [6]. Furthermore, LINC00467 knock down or miR-107 over-expression has suppressed tumorigenic ability of cervical cancer cell in xenograft models [8]. Similar studies in AML, bladder cancer, colorectal cancer, esophageal carcinoma, glioma, hepatocellular carcinoma, lung cancer and prostate cancer have consistently confirmed oncogenic effects of LINC00467 (Table 2).
Clinical studies
Assessment of expression data from a GEO dataset and the TCGA database has revealed up-regulation of LINC00467 in bladder cancer samples and negative correlation between its expression and patients’ prognosis [17]. Expression assays in patients with breast cancer has also verified over-expression of LINC00467 in cancerous tissues compared with nearby normal samples. Moreover, up-regulation of LINC00467 has been associated with poor overall survival (OS) [6]. Another study has indicated association between LINC00467 over-expression and tumor metastases and poor prognosis. Genomic and epigenetic analyses have shown the impact of copy number amplification, chromatin configuration, and methylation status of DNA on expression of this lncRNA. Copy number amplification and up-regulation of LINC00467 has been associated with the lower levels CD8 + and CD4 + T cells infiltrations [7]. LINC00467 level has also been reported to be elevated in colorectal cancer tissues compared with normal colon mucosal counterparts. In silico analyses available datasets have confirmed correlation between over-expression of LINC00467 and poor OS and recurrent-free survival rate [18]. The association between over-expression of LINC00467 and poor clinical outcome has been verified in different cancers, including bladder cancer, breast cancer, colorectal cancer, glioma, lung cancer, osteosarcoma and testicular germ cell tumor (Table 3).
Discussion
Numerous studies have indicated up-regulation of LINC00467 in different types of cancers. Mechanistically, this lncRNA can be up-regulated through DNA demethylation and copy number variations.
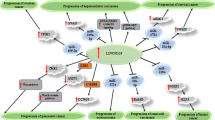
The sponging effect of LINC00467 on miRNAs has been well assessed in different cancer cell lines. Through this mechanistical route, LINC00467 can affect activity of miR-339/SKI, miR-107/KIF23, miR-133b/FTL, miR-485-5p/DPAGT1, miR-7-5p/EGFR, miR-339-3p/IP6K2, miR-200a/E2F3, miR-1285-3p/TFAP2A, miR-299-5p/USP48, miR-509-3p/PDGFRA, miR-18a-5p/NEDD9, miR-9-5p/PPARA, miR-20b-5p/CCND1, miR-125a-3p/SIRT6, miR-217/KPNA4, miR-217/HMGA1 and miR-494-3p/STAT3 axes. Moreover, LINC00467 can influence activity of NF-κB, STAT1, Wnt/b-catenin, Akt and ERK1/2 signaling pathways. Most notably, LINC00467 has been shown to increase EMT in breast, cervical, colorectal, head and neck and prostate cancer as well as osteosarcoma. Thus, strategies to decrease expression of LINC00467 are expected to affect tumor invasion and metastasis.
LINC00467 has a possible role in the tumor microenvironment and immune evasion. Copy number variations within LINC00467 have been associated expression levels of this lncRNA, immune infiltration in lung adenocarcinoma and poor clinical outcome [38]. Moreover, LINC00467 expression in breast cancer has been associated with immune infiltration [7].
Up-regulation of LINC00467 has been associated with poor prognosis of patients with bladder cancer, breast cancer, colorectal cancer, glioma, lung cancer, osteosarcoma and testicular germ cell tumor. Thus, LINC00467 is a putative prognostic marker in cancers. However, the potential of this lncRNA as a diagnostic marker has not well studied. Future studies should focus on this aspect. Expression assays of LINC00467 particularly in biofluids such as serum and urine would pave the way for establishment of non-invasive methods for cancer diagnosis.
Identification of additional miRNA targets of LINC00467 is expected to clarify the molecular mechanisms and signaling pathways being affected by this lncRNA. This would help in design of novel and efficient targeted therapies for cancer. Based on the critical roles of LINC00467 in the regulation of cell apoptosis, it is expected that modification of its expression affects response of cancer cells to anti-cancer modalities. This function of LINC00467 has been verified in hepatocellular carcinoma cells where its silencing has enhanced sensitivity to Axitinib ([10].
Finally, the presence of single nucleotide polymorphisms within LINC00467 would affect expression or function of this lncRNA. Therefore, genotyping of these variants would help in recognition of risk factors for different types of cancer.
Conclusions and future prospects
LINC00467 is regarded as an oncogenic lncRNA in humans. Thus, strategies to down-regulate its expression are theoretically effective in reduction of tumor burden. The most challenging issue in this regard is establishment of effective ways to convey LINC00467-targetted therapies in a specific way to cancer cells and avoid off-target effects.
Data Availability
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
References
Zhang X, Wang W, Zhu W, Dong J, Cheng Y, Yin Z, et al. Mechanisms and Functions of Long Non-Coding RNAs at Multiple Regulatory Levels. Int J Mol Sci. 2019;20(22):5573. PubMed PMID: 31717266. eng.
Statello L, Guo C-J, Chen L-L, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22(2):96–118.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22(9):1775–89.
Atmadibrata B, Liu PY, Sokolowski N, Zhang L, Wong M, Tee AE, et al. The novel long noncoding RNA linc00467 promotes cell survival but is down-regulated by N-Myc. PLoS ONE. 2014;9(2):e88112.
Lu J, Wu X, Wang L, Li T, Sun L. Long noncoding RNA LINC00467 facilitates the progression of acute myeloid leukemia by targeting the miR-339/SKI pathway. Leuk Lymphoma. 2021;62(2):428–37.
Zhang Y, Sun Y, Ding L, Shi W, Ding K, Zhu Y. Long non-coding RNA LINC00467 correlates to poor prognosis and aggressiveness of breast cancer. Frontiers in oncology. 2021;11.
Bo H, Zhang W, Zhong X, Chen J, Liu Y, Cheong K-L, et al. LINC00467, Driven by Copy Number Amplification and DNA Demethylation, Is Associated with Oxidative Lipid Metabolism and Immune Infiltration in Breast Cancer. Oxidative medicine and cellular longevity. 2021;2021.
Li G-C, Xin L, Wang Y-S, Chen Y. Long intervening noncoding 00467 RNA contributes to tumorigenesis by acting as a competing endogenous RNA against miR-107 in cervical cancer cells. Am J Pathol. 2019;189(11):2293–310.
Chen Y, Ding Y. LINC00467 enhances head and neck squamous cell carcinoma progression and the epithelial–mesenchymal transition process via miR-299‐5p/ubiquitin specific protease‐48 axis. J Gene Med. 2020;22(7):e3184.
Li W, He Y, Chen W, Man W, Fu Q, Tan H, et al. Knockdown of LINC00467 contributed to Axitinib sensitivity in hepatocellular carcinoma through miR-509-3p/PDGFRA axis. Gene Ther. 2021;28(10):634–45.
Zheng Y, Nie P, Xu S. Long noncoding RNA linc00467 plays an oncogenic role in hepatocellular carcinoma by regulating the miR-18a‐5p/NEDD9 axis. J Cell Biochem. 2020;121(5–6):3135–44.
Cai K, Li T, Guo L, Guo H, Zhu W, Yan L, et al. Long non-coding RNA LINC00467 regulates hepatocellular carcinoma progression by modulating miR-9-5p/PPARA expression. Open biology. 2019;9(9):190074.
Yan J, Fang T, Zhang M, Zhou Q. LINC00467 facilitates osteosarcoma progression by sponging miR–217 to regulate KPNA4 expression. Int J Mol Med. 2021;47(3):1-.
Ma H, Wang J, Shi J, Zhang W, Zhou D. LncRNA LINC00467 contributes to osteosarcoma growth and metastasis through regulating HMGA1 by directly targeting miR-217. Eur Rev Med Pharmacol Sci. 2020;24(11):5933–45.
Ding H, Luo Y, Hu K, Liu P, Xiong M. Linc00467 promotes lung adenocarcinoma proliferation via sponging miR-20b-5p to activate CCND1 expression. OncoTargets and therapy. 2019;12:6733.
Chang Y, Yang L. LINC00467 promotes cell proliferation and stemness in lung adenocarcinoma by sponging miR-4779 and miR‐7978. J Cell Biochem. 2020;121(8–9):3691–9.
Xiao J, Gong L, Xiao M, He D, Xiang L, Wang Z, et al. LINC00467 Promotes Tumor Progression via Regulation of the NF-kb Signal Axis in Bladder Cancer. Front Oncol. 2021;11:1958.
He X, Li S, Yu B, Kuang G, Wu Y, Zhang M, et al. Up-regulation of LINC00467 promotes the tumourigenesis in colorectal cancer. J Cancer. 2019;10(25):6405.
Liang Y, Cheng G, Huang D, Yuan F. Linc00467 promotes invasion and inhibits apoptosis of head and neck squamous cell carcinoma by regulating miR-1285-3p/TFAP2A. Am J Translational Res. 2021;13(6):6248.
Jiang W, Cheng X, Wang T, Song X, Zheng Y, Wang L. LINC00467 promotes cell proliferation and metastasis by binding with IGF2BP3 to enhance the mRNA stability of TRAF5 in hepatocellular carcinoma. J Gene Med. 2020;22(3):e3134.
Han Y, Cai Y, Lai X, Wang Z, Wei S, Tan K, et al. lncRNA RMRP Prevents Mitochondrial Dysfunction and Cardiomyocyte Apoptosis via the miR-1-5p/hsp70 Axis in LPS-Induced Sepsis Mice. Inflammation. 2020:1–14.
Bai Y, Wu H, Han B, Xu K, Liu Y, Liu Y, et al. Long intergenic non–protein coding RNA–467 targets microRNA–451a in human colorectal cancer. Oncol Lett. 2020;20(5):1-.
Liu Z, Yang S, Chen X, Dong S, Zhou S, Xu S. LncRNA LINC00467 acted as an oncogene in esophageal squamous cell carcinoma by accelerating cell proliferation and preventing cell apoptosis via the miR-485‐5p/DPAGT1 axis. J Gastroenterol Hepatol. 2021;36(3):721–30.
Deng L-H, Zhao H, Bai L-P, Xie J, Liu K, Yan F. Linc00467 promotion of gastric cancer development by directly regulating miR-7-5p expression and downstream epidermal growth factor receptor. Bioengineered. 2021;12(2):9484–95.
Xu L, Liu C, Ye Z, Wu C, Ding Y, Huang J. Overexpressed LINC00467 promotes the viability and proliferation yet inhibits apoptosis of gastric cancer cells via raising ITGB3 level. Tissue Cell. 2021;73:101644.
Jiang X, Liu Y. LINC00467 promotes proliferation and invasion in glioma via interacting with miRNA-485-5p. Eur Rev Med Pharmacol Sci. 2020;24:766–72.
Zhang Y, Zhang Y, Wang S, Cao B, Hu D, Jia J, et al. LINC00467 facilitates the proliferation, migration and invasion of glioma via promoting the expression of inositol hexakisphosphate kinase 2 by binding to miR-339-3p. Bioengineered. 2022;13(2):3370–82.
Zhang Y, Jiang X, Wu Z, Hu D, Jia J, Guo J, et al. Long noncoding RNA LINC00467 promotes glioma progression through inhibiting P53 expression via binding to DNMT1. J Cancer. 2020;11(10):2935.
Gao S, Duan H, An D, Yi X, Li J, Liao C. Knockdown of long non-coding RNA LINC00467 inhibits glioma cell progression via modulation of E2F3 targeted by miR-200a. Cell Cycle. 2020;19(16):2040–53.
Liang R, Tang Y. LINC00467 knockdown repressed cell proliferation but stimulated cell apoptosis in glioblastoma via miR-339-3p/IP6K2 axis. Cancer Biomarkers. 2020;28(2):169–80.
Wang H, Guo Q, Nampoukime KPB, Yang P, Ma K. Long non-coding RNA LINC00467 drives hepatocellular carcinoma progression via inhibiting NR4A3. J Cell Mol Med. 2020;24(7):3822–36.
Yang J, Liu Y, Mai X, Lu S, Jin L, Tai X. STAT1-induced upregulation of LINC00467 promotes the proliferation migration of lung adenocarcinoma cells by epigenetically silencing DKK1 to activate Wnt/β-catenin signaling pathway. Biochem Biophys Res Commun. 2019;514(1):118–26.
Wang X, Liu H, Shen K, Pan X, Wei Y, Lv T, et al. Long intergenic non–coding RNA 00467 promotes lung adenocarcinoma proliferation, migration and invasion by binding with EZH2 and repressing HTRA3 expression. Mol Med Rep. 2019;20(1):640–54.
Zhu Y, Li J, Bo H, He D, Xiao M, Xiang L, et al. LINC00467 is up-regulated by TDG-mediated acetylation in non-small cell lung cancer and promotes tumor progression. Oncogene. 2020;39(38):6071–84.
Xue F, Yang C, Yun K, Jiang C, Cai R, Liang M, et al. Reduced LINC00467 elevates microRNA-125a-3p to suppress cisplatin resistance in non-small cell lung cancer through inhibiting sirtuin 6 and inactivating the ERK1/2 signaling pathway. Cell Biology and Toxicology. 2021:1–17.
Jiang H, Deng W, Zhu K, Zeng Z, Hu B, Zhou Z, et al. LINC00467 promotes prostate cancer progression via M2 macrophage polarization and the miR-494-3p/STAT3 axis. Front Oncol. 2021;11:1688.
Bo H, Zhu F, Liu Z, Deng Q, Liu G, Li R, et al. Integrated analysis of high-throughput sequencing data reveals the key role of LINC00467 in the invasion and metastasis of testicular germ cell tumors. Cell Death Discovery. 2021;7(1):1–12.
Wang W, Bo H, Liang Y, Li G. LINC00467 Is Upregulated by DNA Copy Number Amplification and Hypomethylation and Shows ceRNA Potential in Lung Adenocarcinoma. Frontiers in endocrinology. 2021;12.
Acknowledgements
This study was financially supported by Grant from Medical School of Shahid Beheshti University of Medical Sciences.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
SGF wrote the manuscript and revised it. MT supervised and designed the study. TK, MH and BMH collected the data and designed the figure and tables. All authors read and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participant
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent forms were obtained from all study participants. The study protocol was approved by the ethical committee of Shahid Beheshti University of Medical Sciences. All methods were performed in accordance with the relevant guidelines and regulations.
Consent of publication
Not applicable.
Competing interests
The authors declare they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Ghafouri-Fard, S., Khoshbakht, T., Hussen, B.M. et al. A review on the role of LINC00467 in the carcinogenesis. Cancer Cell Int 22, 319 (2022). https://doi.org/10.1186/s12935-022-02724-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12935-022-02724-6