Abstract
Since its emergence in China in 2019, the coronavirus has exerted its impact on every nation. Throughout this period, epidemiological models have been employed to anticipate the disease’s patterns. These models rely on the random diffusion of the virus, overlooking the inherent social interactions within communities. A pioneering approach involves incorporating the concept of networks to account for these human interactions. The present study presents visual depictions of numerical simulations utilizing the Susceptible–Infected–Recovered model on the Barabási–Albert network. This enables an analysis of coronavirus dynamics in Pakistan while considering diverse mitigation strategies. The findings underscore that effective infection management involves preventing its spread into central hubs-nodes with extensive connections-and significantly curtailing social interactions within society.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
In December 2019, the outbreak of the highly contagious COVID-19 disease in Wuhan, China, as documented in [1], sent shockwaves worldwide due to its rapid and life-threatening transmission. Recognizing the gravity of the situation, the World Health Organization (WHO) officially declared it a pandemic in March 2020, as cited in [2]. Subsequently, numerous nations enacted comprehensive strategies aimed at containing and suppressing the virus, with remarkable success. These governments relied on invaluable insights derived from a range of mathematical epidemiological models, including those highlighted in [3,4,5,6], to gauge the severity of the disease and gauge the effectiveness of measures such as lockdowns and social distancing.
In 1927, Kermack and McKendrick laid the foundation for the study of infectious diseases by introducing the concept of a compartmental model. Their pioneering work involved categorizing the entire population into three distinct groups: those susceptible to infection, those currently infected, and those who had recovered. This groundbreaking concept, now recognized as the Susceptible–Infected–Recovered (SIR) model [7], has since become fundamental in epidemiology. Subsequently, an adapted version, known as the Susceptible–Exposed–Infectious–Recovered (SEIR) model, emerged to account for individuals who could carry the virus without yet being infectious, classifying them as “exposed” [8,9,10].
As time has progressed, epidemiologists have continued to refine these models by incorporating various factors. These refinements include considerations for age [11], the impact of mitigation strategies [12], and population density [13], all of which have contributed to enhancing the accuracy and reliability of predictive outcomes in the field of infectious disease modeling.
In the realm of epidemiology, there has been a growing recognition of the limitations of classical models. One prominent issue identified in 2020 by Manzo was the failure of these models to consider the interconnectedness of various interventions [14]. To address this shortcoming, he proposed the integration of social networks as a promising solution.
Social networks, often depicted as connected graphs, provide a powerful framework for understanding the dynamics of infectious diseases within a society. In this conceptualization, individuals are represented as nodes, and their interactions are depicted as edges. Nodes that share edges are considered neighbors, and the degree of a node reflects the number of its neighbors. Particularly influential nodes, known as hubs, exhibit an exceptionally high number of connections. A network is classified as scale-free when the distribution of node degrees follows a power law, suggesting that the number of nodes is related to degrees raised to a specific power.
To delve deeper into the significance of networks in epidemiology, you can explore resources such as [15,16,17,18,19]. For insights into the relationship between networks and the dynamics of the coronavirus, you may find [20] informative. Additionally, to gain a more comprehensive understanding of how coronavirus dynamics are influenced by isolation measures, consider examining [21].
For a broader exploration of social networks and their applications, you can refer to resources like [22, 23].
Although the network approach has proven to be a valuable tool for making predictions, it has certain limitations when it comes to fully representing the complexities of infectious diseases. In the real-world scenario of a disease outbreak within a population, it is unlikely that every individual will become infected. However, network models tend to assume a scenario where all individuals eventually succumb to the infection.
2 Methodology
Our methodology involves the following steps: We will utilize the COVID-19 data from Pakistan and conduct simulations of the SIR model using the Barabási–Albert network. This approach aims to elucidate the disease’s dynamics within this context.
2.1 The coronavirus data
Based on data retrieved from the COVID-19 Health Advisory Platform, a resource established by the Ministry of Health Services Regulations and Coordination of Pakistan, the period spanning from June 16, 2022, to July 15, 2022, witnessed an average of approximately 1.5376 million confirmed COVID-19 cases in Pakistan. During this same timeframe, the corresponding average for individuals who had successfully recovered from the disease reached approximately 1.5 million. These statistics are derived from a total population estimate of 235.8 million. To put these numbers into perspective, the confirmed cases represent around 0.65% of the overall population, while the recovered cases account for approximately 0.63%.
For a more visual representation of this data, please consult Fig. 1. If you wish to access the data source directly, it is available at https://covid.gov.pk/stats/pakistan.
2.2 The SIR model
In the year 1927, Kermack and McKendrick introduced a groundbreaking concept in the study of infectious diseases by pioneering what is now known as a compartmental model. Their innovative framework suggested that the entire population, denoted as N for a given society, could be categorized into three distinct groups: Those who are susceptible (S), those who are currently infected (I), and those who have successfully recovered (R). This influential model is commonly referred to as the Susceptible–Infected–Recovered (SIR) model, or more simply, the SIR model.
The SIR model operates on the foundational principles that a susceptible individual represents someone who is in good health but remains vulnerable to infection. In contrast, an infected individual is actively carrying the infectious agent and has the potential to transmit it to susceptible individuals. Once an individual recovers from the infection, they not only cease to carry the disease but also acquire immunity against subsequent infections of the same kind. In practical terms, this model partitions the entire population (N) into the susceptible (S), infected (I), and recovered (R) categories, as visually depicted in Fig. 2.
It is important to note that, within the framework of the SIR model, the infectious disease exclusively targets healthy individuals (those in the susceptible category). Additionally, as individuals recover and gain immunity, the sum of individuals in the susceptible, infected, and recovered categories remains constant and equal to the total population (N).
The original SIR model in the form of system of differential equations is shown in Fig. 3. Here, \(\beta\) and \(\gamma\) represent rate of infection and rate of recovery, respectively.
3 Results and discussion
In this segment, our objective is to comprehend the impact of the coronavirus on the population of Pakistan while implementing distinct mitigation strategies. To achieve this, we envision the entire population as an interconnected network of individuals. Given the substantial resemblance between this network and the Barabási–Albert network, we will conduct simulations of the SIR model on this framework. These simulations will encompass all the parameters inherent to the SIR model.
To expedite the simulation process, we’ve downscaled the population of Pakistan by a factor of 99.99, The simulation algorithm operates in the following manner: An initial network is generated with an average connectivity of \(k=3\). To start, 496 nodes out of the total 550 are designated as susceptible (S), while 4 nodes are randomly assigned to the infected state. The infection probability is established at \(p=0.1\), denoting the likelihood that a susceptible individual becomes infected upon contact with an infected person. Consequently, the infection rate is calculated as \(\beta = k \times p = 0.3\). The recovery period for an infected individual is set to \(\frac{1}{\gamma } = 6\) days, aligning with the consistent duration observed for recovery from the coronavirus. To obtain a reliable trend for disease spread, 30 samples are taken. A visual representation of the network, depicted in Fig. 4, showcases a subset of 50 individuals, among whom 4 are initially marked as infected.
Upon constructing the Barabási–Albert (BA) network, certain nodes acquire an exceptionally high number of connections, earning them the designation of hubs. Notably, we discern a significant alteration in the disease dynamics based on the positioning of the initial infected individuals: whether they are located exclusively within hubs, solely outside hubs, or a combination of both hubs and non-hub nodes.
3.1 Hub mitigation
We proceed to generate the Barabási–Albert (BA) network, encompassing a total of \(N=550\) nodes to symbolize the complete population of Pakistan. This network is constructed with an average connectivity of \(k=3\), an infection probability of \(p=0.1\), and an initial infected count of \(I=4\). As previously noted, this network inherently comprises nodes characterized by significantly elevated degrees, commonly referred to as hubs. The presence of hubs within such networks is a natural occurrence. To illustrate, in an organizational context, a public relations officer could serve as a hub due to their heightened interactions with visitors compared to other members of the organization.
It becomes apparent that when all four initially infected individuals originate from hubs, the infection spreads at an accelerated pace. The outcomes of our simulations, depicted in Fig. 5a, highlight that the infection reaches its zenith within merely 11 days. Furthermore, nearly the entire populace becomes infected, signifying a critical predicament for the society.
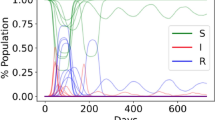
Simulated results of 30 samples of the SIR model on the Barabási–Albert network with \(N=550\) nodes and \(I=4\) infected nodes. The probability of getting infection per contact of a healthy person from an infected person is \(p=0.1\). Figures a and b represent respectively the simulations on the BA network before and after mitigation
How can this issue be addressed? By targeting the hubs. To address this, we adopt the strategy of disrupting hubs. This involves randomly removing edges within the hubs in a manner that no hub retains more than six connections. This approach, known as hub mitigation, is designed to curtail the propagation of infectious diseases. The application of this mitigation technique yields notable improvements in the situation. In Fig. 5b, you can observe that the pace of infection transmission decelerates. The infection now requires 17 days, as opposed to the initial 11 days, to reach its peak. Furthermore, the infection no longer engulfs the entire population, signifying a substantial enhancement.
3.2 Random mitigation
The second scenario involves the initial four infected individuals being positioned anywhere within the network. Given that we are conducting 30 simultaneous samples, the probability of an infected person being located both inside and outside the hubs is approximately equal. Under this circumstance, the infection rate is slightly lower compared to the hub-centric scenario. In this case, the infection reaches its peak within 12 days and subsequently diminishes by day 19. Figure 6 visually illustrates this trend, demonstrating that the disease impacts all individuals within the social network before being eradicated from the community.
To manage the disease, we employ a strategy termed random mitigation, involving the random removal of connections until no node retains a degree exceeding six. The outcomes are promising, as the infection rate diminishes, the peak of infections lowers, and a smaller proportion of the population becomes infected (as demonstrated in Fig. 6).
3.3 Global mitigation
Presently, we are introducing an additional method of mitigation. Initially, we require individuals who are infected to remain in areas outside of central hubs. Subsequently, we implement a random removal of connections, aiming to limit the degree of each node to six or lower. This approach exerts a significant level of control over the spread of the disease. As a result of these measures, the infection rate reaches its minimum point, and the disease’s impact is greatly diminished. It is noteworthy that the disease scarcely affects even fifty percent of the total population, as illustrated in Fig. 7. This underscores the effectiveness of this technique as a means to manage various infectious diseases, particularly the coronavirus.
4 Conclusion
This paper delves into our simulation of the SIR model on the Barab’asi-Albert network, with the primary aim of analyzing the COVID-19 dynamics in Pakistan under various mitigation strategies. Our focus extended to three distinct approaches: random, hub-based, and global mitigations. Each of these strategies exhibited notable shifts in infection and recovery rates. Particularly striking was the rapid spread of infection upon its entry into hubs.
Our findings strongly advocate for a two-pronged approach to disease control: preventing the disease’s ingress into hubs while simultaneously minimizing interaction frequencies. This twofold strategy demonstrated the most effective means of disease management.
Furthermore, our study underscores the potential for enhanced efficacy in our conclusions by factoring in the influences of vaccination and age on the dynamics. Incorporating these elements could provide a more comprehensive understanding of COVID-19 dynamics and subsequently refine our mitigation recommendations.
Data Availability
The official source for information on the coronavirus in Pakistan can be found at https://covid.gov.pk/stats/pakistan, which is hosted by the COVID-19 Health Advisory Platform administered by the Ministry of Health Services Regulations and Coordination of Pakistan.
References
J.T. Wu, K. Leung, M. Bushman, N. Kishore, R. Niehus, P.M. de Salazar, B.J. Cowling, M. Lipsitch, G.M. Leung, Estimating clinical severity of COVID-19 from the transmission dynamics in Wuhan, China. Nat. Med. 26, 506–510 (2020)
H. Harapan, N. Itoh, A. Yufika, W. Winardi, S. Keam, H. Te, D. Megawati, Z. Hayati, A.L. Wagner, M. Mudatsir, Coronavirus disease 2019 (COVID-19): a literature review. J. Infect. Public Health 13, 667–673 (2020)
R.M. Anderson, H. Heesterbeek, D. Klinkenberg, T.D. Hollingsworth, How will country-based mitigation measures influence the course of the COVID-19 epidemic? The Lancet 395, 931–934 (2020)
N. M. Ferguson, D. Laydon, G. Nedjati-Gilani, N. Imai, K. Ainslie, M. Baguelin, S. Bhatia, A. Boonyasiri, Z. Cucunubá, G. Cuomo-Dannenburg, others, Impact of non-pharmaceutical interventions (NPIs) to reduce COVID-19 mortality and healthcare demand (2020)
J. Hellewell, S. Abbott, A. Gimma, N.I. Bosse, C.I. Jarvis, T.W. Russell, J.D. Munday, A.J. Kucharski, W.J. Edmunds, F. Sun, S. Flasche, Feasibility of controlling COVID-19 outbreaks by isolation of cases and contacts. The Lancet Glob. Health 8, e488–e496 (2020)
S. Zhao, H. Chen, Modeling the epidemic dynamics and control of COVID-19 outbreak in China. Quant. Biol. 8, 11–19 (2020)
W.O. Kermack, A.G. McKendrick, A contribution to the mathematical theory of epidemics, Proceedings of the royal society of London. Series A Contain. Papers of a Math. Phys. Character 115, 700–721 (1927)
X. He, E.H. Lau, P. Wu, X. Deng, J. Wang, X. Hao, Y.C. Lau, J.Y. Wong, Y. Guan, X. Tan, X. Mo, Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat. Med. 26, 672–675 (2020)
A. Rǎdulescu, C. Williams, K. Cavanagh, Management strategies in a SEIR-type model of COVID 19 community spread. Sci. Rep. 10, 1–16 (2020)
J.T. Wu, K. Leung, G.M. Leung, Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study. The Lancet 395, 689–697 (2020)
S.L. Chang, N. Harding, C. Zachreson, O.M. Cliff, M. Prokopenko, Modelling transmission and control of the COVID-19 pandemic in Australia. Nat. Commun. 11, 1–13 (2020)
H.A. Herrmann, J.-M. Schwartz, Why COVID-19 models should incorporate the network of social interactions. Phys. Biol. 17, 065008 (2020)
X. Luo, S. Feng, J. Yang, X.-L. Peng, X. Cao, J. Zhang, M. Yao, H. Zhu, M. Y. Li, H. Wang, others, Analysis of potential risk of COVID-19 infections in China based on a pairwise epidemic model (2020)
G. Manzo, Complex social networks are missing in the dominant COVID-19 epidemic models. Sociologica 14, 31–49 (2020)
Y. Okabe, A. Shudo, Spread of variants of epidemic disease based on the microscopic numerical simulations on networks, Scientific reports 12 (2022) 1–9. Publisher: Nature Publishing Group
Z. Dezső, A.-L. Barabási, Halting viruses in scale-free networks. Phys. Rev. E 65, 055103 (2002). (Publisher: APS)
M.J. Keeling, K.T. Eames, Networks and epidemic models. J. R. Soc. Interface 2, 295–307 (2005)
M.E. Newman, Spread of epidemic disease on networks. Phys. Rev. E 66, 016128 (2002)
R. Pastor-Satorras, A. Vespignani, Epidemic spreading in scale-free networks. Phys. Rev. Lett. 86, 3200 (2001)
M. Rafiq, A.R. Nizami, D. Baleanu, N. Ahmad, Numerical simulations on scale-free and random networks for the spread of COVID-19 in Pakistan. Alex. Eng. J. 62, 75–83 (2023). (Publisher: Elsevier)
K. Choi, H. Choi, B. Kahng, Covid-19 epidemic under the K-quarantine model: Network approach, arXiv preprint arXiv:2010.07157 (2020)
G. Caldarelli, M. Catanzaro, Networks: A Very Short Introduction, vol. 335 (Oxford University Press, 2012)
Y. Okabe, A. Shudo, Microscopic numerical simulations of epidemic models on networks. Mathematics 9, 932 (2021)
Acknowledgements
The authors would like to acknowledge Deanship of Scientific Research, Taif University for funding this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nizami, A.R., Rafiq, M., Inc, M. et al. Numerical simulations with mitigation strategies on Barabási–Albert network for the spread of coronavirus in Pakistan. Eur. Phys. J. Plus 138, 972 (2023). https://doi.org/10.1140/epjp/s13360-023-04628-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjp/s13360-023-04628-4