Abstract
Objective: The “fall armyworm” also known as Spodoptera frugiperda, is a maize pest that is native to North America. It invaded Africa in 2016 and caused serious economic damage, forcing the continent’s nations to take swift action to fend off this new invading pest. It is urgently necessary to produce powerful insecticides for the efficient management of this insect pest since this parasite is a novel invading species and its resistance to insecticides has to be investigated. Methods: The synthesized thiazole-owing hydrazone derivatives are the subject of the first study on the harmful effects they cause, followed by structural relationships. Following the stated protocols, four molecules of thiazole-owing hydrazone derivatives were synthesized as pure active ingredients, and their efficacy as potential insecticides against S. frugiperda was tested. Results and Discussion: The toxicity data exhibited that compound (V) with LC50 of 13.66 and 106.25 ppm was more toxic against the 2nd and 4th larvae instars, respectively, than other synthesized compounds; the other screened compounds showed weak to strong toxicological activity against S. frugiperda. A molecular docking investigation of four synthetic compounds (III–VI) was performed against acetylcholinesterase (AChE). The molecular docking results revealed that compound (V) has the more negative docking score and best binding (docking score = –9.01 kcal/mol), followed by compounds (III), (IV), and (VI) with docking scores of –8.76, –8.53, and –7.41 kcal/mol, respectively. The protein-ligand docking postures revealed that these substances exhibited a strong affinity for the target enzyme’s active site (Protein Data Bank ID: 2ACE). This work suggests that these substances may have insecticidal and AChE inhibitory properties, and it may be possible to further explore them in the process of creating pesticides that target AChE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
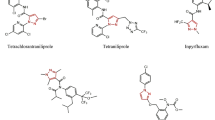
Agrochemicals, which include fungicides, insecticides, and herbicides, raise the quality and quantity of agricultural products. Modern agriculture has come to recognize innovative agrochemicals as a pillar supporting strong crop production [1]. In this respect, many potent tools have been developed since the discovery of agrochemicals [2–4]. Among these instances, the generation of organosulfur agrochemicals plays a growing role. Organosulfur agrochemicals have advanced significantly and found widespread use in contemporary crop protection over the past few decades [5]. Additionally, the types of sulfur-containing compounds found in these agrochemicals can be broadly categorized as thioethers, thiophosphates, sulfonamides, sulfones, sulfoxides, thioureas, and sulfur heterocycles (including thiazolidine, thiophene, thiazole, and benzothiazole) [5]. Thiazole is a crucial heterocyclic component widely applied in commercial pesticides, such as insecticides (e.g., thiamethoxam and clothianidin, nicotinic acetylcholine receptor competitive regulators), nematicides (e.g., fluensulfone, metabolic inhibitors of energy storage processes), and fungicides (e.g., thifluzamide, thiabendazole, and ethaboxam) (Fig. 1) [6–8]. Thiazole derivatives have found applications in different fields, such as medicinal chemistry [9, 10], agrochemicals [11], and biologically active natural products [12], and materials science [13], especially in liquid crystals, molecular switches, sensors, or sunscreens in the cosmetic industry. Hydrazone has garnered greater interest as a characteristic active component and is seen as crucial for pesticide discovery [14]. It has received significant attention as an achievable herbicide, insecticide, antibacterial, antifungal, and antiviral agent [15].
The autumn armyworm (FAW), also known as Spodoptera frugiperda (J.E. Smith), is a destructive insect pest that has developed into a significant global threat to agricultural output [16]. The fall armyworm causes damage to different crops [17], including maize, rice, and beans. Sorghum, sugarcane, beetroot, cotton, groundnuts, soybeans, alfalfa, onions, pasture grasses, millet, tomato, and potato S. frugiperda grazing on the host plant’s reproductive and vegetative components results in decreased yields. Early larvae feed by first feeding near the ground and later by chewing through leaves from the outside in. The density of larval populations is halved as a result of cannibalism [18]. Maize, also grown commercially in Egypt, is known as the “Queen of Cereals” and is one of the most significant grains [19, 20]. In light of the aforementioned facts, the current work intends to create certain thiazole derivatives and estimated as insecticides agent against 2nd and 4th instar larvae of S. frugiperda.
RESULTS AND DISCUSSION
Chemistry
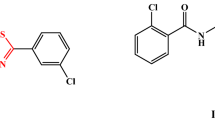
The target compounds are N-(4-phenyl-thiazol-2-yl)N-[4-(piperidin-1-yl)benzylidenyl]hydrazine (III), N-(4methyl-thiazol-2-yl)-N-[4-(piperidin-1-yl)benzylidenyl]hydrazine (IV), 2-[N-(4-piperidin-1-yl)benzylidenyl]hydrazino]-4,5-dihydrothiazol-4-one (V), and 2-[N-(4(piperidin-1-yl)benzylidenyl)hydrazino]naphtho[2,3-d]thiazol-4,9-dione (VI) has been generated using the procedures we previously provided [21]. The synthetic route of the target compounds is summarized in (Scheme 1). The 4-(piperidin-1-yl)benzylidenyl-thiosemicarbazide (II) was generated once 4-piperidinobenzaldehyde (I) was subjected to treatment with thiosemicarbazide in refluxing ethanol. The corresponding N-(4-phenyl-thiazol2-yl)-N-(4-piperidin-1-yl-benzylidene) hydrazine (III) was obtained by reacting compound (II) with phenacyl bromide in ethanol in the presence of fused sodium acetate at ambient temperature. Similarly, compound (II) was cycloalkylated with chloroacetone, ethyl chloroacetate in refluxing ethanol, and sodium acetate to yield the thiazole derivatives (IV) and (V), respectively. The cyclocondensation of compound (II) with 2,3-dichloro naphthoquinone in N,N-dimethylformamide (DMF) in the presence of anhydrous potassium carbonate at reflux temperature created the naphtho[2,3-d]thiazole derivative (VI).
All the target insecticidal compounds have been screened for insecticidal bioefficacy, as explained below.
Toxicological Activity Test for 2nd Instar Larvae of S. frugiperda after 72 h of Treatment
The insecticidal test efficacy of (III–VI) under laboratory conditions against 2nd instar larvae of S. frugiperda is shown in Table 1. The LC50 values were 48.09, 84.12, 13.66, and 62.44 ppm for (III–VI), respectively. In addition, each had slope values of 0.915, 1.35, 0.911, and 1.345, demonstrating the homogeneity of the larvae, whose toxic ratios were 28.40, 16.20, 100, and 21.87. Results exhibited that target compound (V), was more toxic (LC50 = 13.66 ppm) than the other synthesized compounds (Fig. 2).
Toxicological Activity Test for 4th Instar Larvae of S. frugiperda After 72 h of Treatment
The results of compounds (III–VI) which were tested as insecticidal agents against 4th instar larvae of S. frugiperda, are shown in Table 1. After 72 h of treatment, the bioefficacy results of synthesized compounds exhibit high to low toxicological activity because some of them cause a higher kill percentage against insect larvae than others with LC50 values ranging from 106.25 to 158.01 ppm, whereas the LC50 values of compounds (III–VI) were 126.32, 158.01, 106.25, and 137.36 ppm, respectively. In addition, each had slope values of 1.178, 1.129, 1.195, and 1.143, demonstrating the homogeneity of the larvae. From these results, the toxicity of compound (V) against 4th instar larvae of S. frugiperda was the most efficient, with an LC50 value of 106.25 ppm (Fig. 3).
Molecular Docking
Molecular docking is a method that predicts the preferred orientation and binding affinity of one molecule (a ligand) to another molecule (a receptor) when they form a stable complex [22, 23]. Molecular docking is important for understanding the molecular interactions that underlie biological processes such as signal transduction, enzyme catalysis, and drug action [24]. Because molecular docking could be used to find potential drug candidates that bind to specific target proteins, it is also frequently utilized in structure-based drug design [25]. First, the re-docking and superimposition methods were used to validate the docking operation [26].
The 2ACE’s natural ligand was taken out and docked back into the active site. Re-docking was done to evaluate the efficiency of the docking process. The re-docking operation was carried out utilizing the same approaches as the compounds under investigation. The co-crystallized ligand’s binding pattern was successfully recreated in the re-docking validation stage, proving the utilized docking protocol was appropriate for the intended docking investigation. The superimposition between the re-docked ligand and the native co-crystallized one with small RMSD of 1.012 Å was shown in Fig. S1 (supplementary information). The docking scores of the investigated compounds (III–VI) against the Acetylcholinesterase (AChE) enzyme (PDB ID: 2ACE) are shown in Table 1. The docking scores for the compounds ranged from –9.01 kcal/mol for compound (V) to –7.41 kcal/mol for compound (VI). The compound (V) was observed to have the most negative docking score (S) and best binding (S = –9.01 kcal/mol), followed by compound (III) (S = –8.76 kcal/mol), compound (IV) (S = –8.53 kcal/mol), and finally compound (VI) with lowest score (S = –7.41 kcal/mol). As shown in Fig. 4 depict the binding location of the investigated compounds in the active site of 2ACE interaction 3D and 2D, while, Table 2 lists the docking data. The analyses of the molecular contacts, the compound (V); two H-acceptor bonds are formed at distances of 3.14 Å between O21 with HIS440, Table 2. Additionally, two pi-H bonds are formed between the C4 with TRP279 at a distance of 3.88, and 4.38 Å, Table 2. On the other hand, two H-acceptors are formed between N2 with HIS440, at a distance of 3.47, in the case of compound (III). Moreover, three pi-pi interactions were formed between 6-ring with TRP279 at distance of 3.60, and 3.54 Å, Table 2.
Structure-Action Relationship
As a continuation of this work, the relationship between structure and action was introduced here in accordance with the toxicity value in Table 1. All target chemical compounds get their basic structure from the thiazole-owing hydrazone, which may be divided into three major chemical components that are crucial to the toxicological effectiveness of all derivatives: piperidine ring, benzylidene, and thiazole. The piperidine ring and benzylidenyl part are present in all chemical compounds (III–VI). The only difference between the 4 insecticidal agents is the thiazole ring, which is attached to different substituents like phenyl, methyl, and naphthyl. Toxicity results indicate that compound (V), which consists of thiazol-4(5H)-one as a substituent, is the most effective insecticidal agent against both the 2nd and 4th larval instars of S. frugiperda. So, the thiazol-4(5H)-one ring is more efficient than other substitutions on compounds (III), (IV), and (VI), responsible for more toxicological activity, which showed a low LC50 value. The order of the insecticidal activity for the synthesized thiazoleowing hydrazone derivatives with the toxicity index was, (V) > (III) > (VI) > (IV). Hydrazone derivatives with the toxicity index was (V) > (III) > (VI) > (IV). This work confirms that organic and bioactive compounds are very important substances due to their different uses as reported before [49–60].
EXPERIMENTAL
Materials and methods. Chemicals. The target compounds N-(4-phenyl-thiazol-2-yl)- N′-[4-(piperidin-1-yl)benzylidenyl]hydrazine (III), N-(4-methyl-thiazol-2yl)- N′-[4-(piperidin-1-yl)benzylidenyl] hydrazine (IV), 2-[N-(4-piperidin-1-yl)benzyl-idenyl]hydrazino]-4,5dihydro-thiazol-4-one (V), and 2-[N-(4-(piperidin-1-yl)benzylidenyl)hydrazino]naphtho[2,3-d] thiazol-4,9-dione (VI) have been tested as insecticidal against the 2nd and 4th larvae instar of S. frugiperda under laboratory conditions.
Formation of thiazole derivatives. General synthetic procedure for N-(4-phenyl-thiazol-2-yl)-N-[4-(piperidin1-yl)benzylid enyl]hydrazine (III), N-(4-methyl-thiazol2-yl)-N′-[4-(piperidin-1-yl-benzylidenyl]-hydrazine (IV) and 2-[N-(4-piperidin-1-yl)benzylidenyl]hydrazino]4,5-dihydro-thiazol-4-one (V) 2-[N-(4-(piperidin-1-yl)benzylidenyl hydrazino]naphtho[2,3-d]thiazol-4,9-dione (VI). A mixture of (II) (0.01 mol), α-halocarbonyl compound (0.01 mol), and sodium acetate (1 g) in ethanol (30 mL) was heated under reflux for 1 h. After cooling, the resulting solid product was collected by filtration, washed with water, and the crude product recrystallized from ethanol.
Insect rearing. The S. frugiperda autumn armyworms were cultured in a lab at the Plant Protection Research Institute Agriculture Research Center. The incubation period was kept constant at 25 ± 1°C, 60 ± 5% (RH), and a 14 : 10 h light-dark cycle. The larvae were raised on fresh castor leaves, and they were raised separately to prevent cannibalism in tiny cups (7 cm in diameter by 3.5 cm in height) filled with sawdust to cut down on moisture.
Laboratory bioassay. Using the leaf dip bioassay techniques, the insecticidal activity of the pesticides mentioned was assessed [27–33]. The target compounds’ findings from laboratory testing are presented here in order to determine the concentrations needed to kill 50% (LC50) of the second- and fourth-instar larvae of S. frugiperda insects. There are five different pesticide concentrations in this study [34–42]. Nearly identical-sized 10 sec instar larvae and 5 fourth instar larvae insects (three replicates) were placed in castor bean leaf discs (9 cm in diameter), which were submerged in the concentration under test for 10 sec, dried, and then provided to the second and fourth instar larvae, respectively [43–52]. Five glass jars containing the larvae were used. The mortality percentage was recovered after 72 h for all insecticides. Mortality was redressed by Abbott’s formula [53]. The measurements of the mortality relapse line were dissected by probit analysis [54]. The harmful index was determined by sun equations [55].
Molecular docking. Molecular docking analyses of the compounds were carried out with the help of the MOE (Molecular Operating Environment) [56]. The structures of the compounds (III–VI) and the standard native ligand 9-(3-Iodobenzylamino)-1,2,3,4tetrahydroacridine were optimized to have the lowest energy levels feasible using the MMFF94x force field. The atomic coordinates of the crystal structures of the target enzyme, acetylcholine esterase (AChE) with the PDB ID of 2ACE, were downloaded from the protein databank (https://www.rcsb.org/structure/2ACE). Before docking or doing any analysis, the target structures had polar hydrogen atoms added to them, and any accessible water molecules, native ligands, and undesirable chains were eliminated [57]. With regard to the other parameters, the default values were implemented [58]. Re-docking and the superimposition approach were used to validate the docking operation. Removed from the 2ACE and re-docked into the active site was the typical ligand 9-(3-Iodobenzylamino)-1,2,3,4-tetrahydroacridine [59, 60].
CONCLUSIONS
Thiazole-owing hydrazone derivatives were chemically prepared. The toxicity of these synthesized compounds was estimated as insecticides against 2nd and 4th instar larvae of S. frugiperda. Toxicity experiments indicate that compound (V) had the best insecticidal activity against 2nd instar and 4th instar larvae of S. frugiperda compared to the other synthesized compounds with LC50 of 13.66 and 106.25 ppm, respectively. Followed by compound (III), who’s LC50 was 48.09 and 126.32 ppm against the 2nd and 4th larval instars, respectively. The insecticidal activity of compound (V) may be due to the presence of a thiazolidone ring as a substituent attached to the basic structure. These results are hopeful and valuable for additional work on the improvement of new and other potent pesticides. The study tested four synthetic compounds (III–VI) for their ability to bind and inhibit Acetylcholinesterase (AChE), an enzyme involved in insecticide action. The results showed that compound (V) had the strongest binding and inhibition of AChE, followed by compounds (III), (IV), and (VI), according to docking score (S). The compounds interacted with the active site of the enzyme, suggesting that they could be potential candidates for developing new insecticides that target AChE.
DATA AVAILABILITY
The data that support the findings of this study are available from the corresponding author upon reasonable request.
REFERENCES
Marchand, P.A., Environ. Sci. Pollut. Res., 2019,vol. 26, pp. 17996–18000. https://doi.org/10.1007/s11356-022-24057-7
Jeanmart, S., Edmunds, A.J.F., Lamberth, C., Pouliot, M., and Morris, J.A., Bioorg. Med. Chem., 2021,vol. 39, Article ID: 116162. https://doi.org/10.1016/j.bmc.2021.116162
Jeanmart, S., Edmunds, A.J.F., Lamberth, C., and Pouliot, M., Bioorg. Med. Chem., 2016, vol. 24,pp. 317–341. https://doi.org/10.1016/j.bmc.2015.12.014
Jeschke, P., Pest. Manag. Sci., 2016, vol. 72, pp. 210–225. https://doi.org/10.1002/ps.4170
Yu, J. and Jiang, X., Adv. Agrochem., 2023, vol. 2,pp. 3–14. https://doi.org/10.1016/j.aac.2022.12.003
Petrou, A., Fesatidou, M., and Geronikaki, A.,Molecules, 2021, vol. 26, Article ID: 3166. https://doi.org/10.3390/molecules26113166
Yadav, C.K., Nandeshwarappa, B.P., and Pasha, K.M.M.,Chimica Techno Acta, 2023, vol. 10, Article ID:202310110. https://doi.org/10.15826/chimtech.2023.10.1.10
Colorado-Peralta, R., Olivares-Romero, J.L., RoseteLuna, S., García-Barradas, O., Reyes-Márquez, V., Hernández-Romero, D., and Morales-Morales, D.,Inorganics, 2023, vol. 11, Article ID: 185. https://doi.org/10.3390/inorganics11050185
Subramaniyan, Raja, Subash, R., Chandra, Bose, R.,and Arunkumar, A., Mater. Sci.: Mater. Electron,2021, vol. 32, pp. 2987–2998. https://doi.org/10.1007/s10854-020-05050-7
Niu, Z., Wang, Y., Zhang, S., Li Y., Chen, X.,Wang, S.,and Liu, H., J. Eur. Med. Chem., 2023, vol. 250,Article ID: 115172. https://doi.org/10.1016/j.ejmech.2023.115172
Maienfisch, P. and Edmunds, A.J.F., Adv. Heterocyclic.Chem., 2017, vol. 212, pp. 35–88. https://doi.org/10.1016/bs.aihch.2016.04.010
Ayati, A., Emami, S., Asadipour, A., Shafiee, A.,and Foroumadi, A., Eur. J. Med. Chem., 2015, vol. 97,pp. 699–718. https://doi.org/10.1016/j.ejmech.2015.04.015
Mori, A., Sekiguchi, A., Masui, K., Shimada, T.,Horie, M., Osakada, K., Kawamoto, M., and Ikeda, T.,J. Am. Chem. Soc., 2003, vol. 125, pp. 1700–1701. https://doi.org/10.1021/ja0289189
Wang, Y., Guo, SH., Yu, L., Zhang, W., Wang, Z.,Chi, Y.R., and Wu, J., Chinese Chem. Lett., 2024,vol. 35, Article ID: 108207. https://doi.org/10.1016/j.cclet.2023.108207
Betancourth, J.G., Castaño, J.A., Visbal, R., andChaur, M.N., Eur. J. Org. Chem., 2022, vol. 28, Article ID:e202200228. https://doi.org/10.1002/ejoc.202200836
Alkorbi, F., Abdelaziz, M.A., Mazi, W., Omer, N.,Jam, R., Gad, M.A., Omran, A.O., and Ali, M.A.,Bull. Chem. Soc. Ethiop., 2024, vol. 38, pp. 765–774. https://doi.org/10.4314/bcse.v38i3.17
Meagher, R.L. and Nagoshi, R.N., Ecol. Entomol.,2004, vol. 29, pp. 614–620. https://doi.org/10.1111/j.0307-6946.2004.00629.x
Jasinski, J.P., Akkurt, M., Mohamed, Sh.K., Gad, M.A.,and Albayati, M.R., Acta Cryst., 2015, vol. 71, pp. 56–57. https://doi.org/10.1107/S2056989014027133
Bakry, M.M.S. and Abdel-Baky, N.F., Braz. J.Biol., 2023, vol. 83, Article ID: e271354. https://doi.org/10.1590/1519-6984.271354
Groot, A.T., Marr, M., Heckel, D.G., and Schöfl, G., Ecol. Entomol., 2010, vol. 35, pp. 105–118. https://doi.org/10.1111/j.1365-2311.2009.01138.x
El-Gaby, M.S.A., J. Chin. Chem. Soc., 2004, vol. 51,pp. 125–134. https://doi.org/10.1002/jccs.200400020
Shivanika, C., Kumar, D., Ragunathan, V., Tiwari, P.,and Sumitha, A., J. Biomol. Struct. Dyn., 2020,vol. 2020, Article ID: 1815584. https://doi.org/10.1080/07391102.2020.1815584
Latif, M., Ahmed, T., Hossain, M.S., Chaki, B., Abdou, A., and Kudrat-E-Zahan, M., Russ. J. Gen. Chem., 2023, vol. 93, pp. 389–397. https://doi.org/10.1134/S1070363223020214
Meng, X.Y., Zhang, H.X., Mezei, M., and Cui, M.,Curr. Comput. Aided Drug Des., 2011, vol. 7,pp. 146–157. https://doi.org/10.2174/157340911795677602
Abd El-Lateef, H.M., Khalaf, M.M., Kandeel, M.,Amer, A.A., Abdelhamid, A.A., and Abdou, A.,Appl. Organomet. Chem., 2023, vol. 37, Article ID: e7134. https://doi.org/10.1002/aoc.7134
El-Saghier, A.M., Enaili, S.S., Abdou, A., Alzahrani, A.Y.A., Ben-Moussa, S., Gad, M.A., and Kadry, A.M., J. Agric. Food Chem., 2024, vol. 2024, Article ID: 3c09703 https://doi.org/10.1021/acs.jafc.3c09703
Abdel-Raheem, Sh.A.A., Fouad, M.R., Gad, M.A.,Kamal El-Dean, A.M., and Tolba, M.S., J. Environ. Chem. Eng., 2023, vol. 11, Article ID: 110839. https://doi.org/10.1016/j.jece.2023.110839
Elhady, O.M., Mansour, E.S., Elwassimy, M. M.,Zawam, S.A., Drar, A.M., and Abdel-Raheem, Sh.A.A., Curr. Chem. Lett., 2022, vol. 11, pp. 285–290. https://doi.org/10.5267/j.ccl.2022.3.006
Abdel-Raheem, Sh.A.A., Drar, A.M., Hussein, B.R.M.,and Moustafa, A.H., Curr. Chem. Lett., 2023, vol. 12,pp. 695–704. https://doi.org/10.5267/j.ccl.2023.5.005
Ahmed, A.A., Mohamed, S.K., and Abdel-Raheem, Sh.A.A., Curr. Chem. Lett., 2022, vol. 11,pp. 359–370. https://doi.org/10.5267/j.ccl.2022.6.001
Shamsan, A.Q.S., Fouad, M.R., Yacoob, W.A.R.M., Abdul-Malik, M.A., and Abdel-Raheem, Sh.A.A.,Curr. Chem. Lett., 2023, vol.12, pp. 431–438. https://doi.org/10.5267/j.ccl.2022.11.003
Mohamed, S.K., Mague, J.T., Akkurt, M., Alfayomy, A.M.,Abou-Seri, S.M., Abdel-Raheem, Sh.A.A., and AbdulMalik, M.A., Acta Cryst., 2022, vol. 78, pp. 846–850. https://doi.org/10.1107/S205698902200603X
Fouad, M.R., Shamsan, A.Q.S., and Abdel-Raheem, Sh.A.A., Curr. Chem. Lett., 2023, vol. 12,pp. 185–192. https://doi.org/10.5267/j.ccl.2022.8.006
Abdel-Raheem, Sh.A.A., Kamal El-Dean, A.M., Abdul-Malik, M.A., Marae, I.S., Bakhite, E.A., Hassanien, R., El-Sayed, M.E.A., Zaki, R.M., Tolba, M.S., Sayed, A.S.A., and Abd-Ella, A.A.,Rev. Roum. Chem., 2022, 67, vol. pp. 305–309. https://doi.org/10.33224/rrch.2022.67.4-5.09
Gad, M.A., Bakry, M.M.S., Tolba, E.F.M., Alkhaibari, A.M., Mashlawi, A.M., Thabet, M.A., AlTaifi, E.A., and Bakhite, E.A., Chem. Biodiver., 2024, vol. 2024, Article ID: e202400451. https://doi.org/10.1002/cbdv.202400451
Bondar, O.V., Karwt, R., Mohammed, T., Pavelyev, R.S.,Pugachev, M.V., Ygaiev, Be.B., Kayumov, A.R., Aimaletdinov, A.M., and Shtyrlin, Y.G., Russ. J. Bioorg.Chem., 2023, vol. 49, pp. 797–814. https://doi.org/10.1134/s106816202304009x
El-Masry, T., Abdl-Mohsen, S.A., and Mohamed, S.A.,Russ. J. Bioorg. Chem., 2021, vol. 47, pp. 561–571. https://doi.org/10.1134/S1068162021020102
Elewa, S., Mansour, E., and Nassar, I.F., Russ. J. Bioorg. Chem., 2020, vol. 46, pp. 382–392. https://doi.org/10.1134/S1068162020030061
Ali, M.A., Salah, H., Gad, M.A., Youssef, M.A.M.,and Elkanzi, N.A.A., ACS Omega, 2022, vol. 7,pp. 40091–40097. https://doi.org/10.1021/acsomega.2c04814
Bakry, M.M.S., Mohammed, L.H., Dabour, N.A.,and Gad, M. A., Curr. Chem. Lett., 2024, vol. 13,pp. 173–186. https://doi.org/10.5267/j.ccl.2023.7.003
El-Saghier, A.M., Abosella, L., Aborahma, G.A., Elakesh, E.O., Abdelhamid, A.A., and Gad, M.A.,Sci. Rep., 2023, vol. 13, p. 13089. https://doi.org/10.1038/s41598-023-39868-y
Abdelhamid, A.A., Elwassimy, M.M., Aref, S. A., andGad, M.A., Biotechnol. Rep., 2019, vol. 24, pp. 394–401. https://doi.org/10.1016/j.btre.2019.e00394
Abdelhamid, A.A, Salama, K.S.M., Elsayed, A.M.,Gad, M.A., and El-Remaily, M.A.A.A., ACS Omega,2022, vol. 7, pp. 3990–4000. https://doi.org/10.1021/acsomega.1c05049
Abd El-Lateef, H.M., Khalaf, M.M., Gouda, M.,Kandeel, M., Amer, A.A., Abdelhamid, A.A.,Drar, A.M., and Gad, M.A., ACS Omega, 2023,vol. 8, pp. 29685–29692. https://doi.org/10.1021/acsomega.3c03831
Bakhite, E.A., Marae, I.S., Gad, M.A., Mohamed, Sh.K.,Mague, J.T., and Abuelhassan, S., J. Agric. Food Chem.,2022, vol. 70, pp. 9637–9644. https://doi.org/10.1021/acs.jafc.2c02776
Abdelhamid, A.A., Aref, S.A., Ahmed, N.A., Elsaghier, A.M.M., Abd El Latif, F.M., Al-Ghamdi, S.N., andGad, M.A., ACS Omega, 2023, vol. 8, pp. 709–717. https://doi.org/10.1021/acsomega.2c05977
El-Gaby, M.S.A., Hussein, M.F., Faraghally, A.F.,Drar, A.M. and Gad, M.A., Curr. Chem. Lett., 2023,vol. 12, pp. 599–606. https://doi.org/10.5267/j.ccl.2023.2.003
El-Saghier, A.M., Enaili, S.S., Kadry, A.M., Abdou, A., and Gad, M.A., Sci. Rep., 2023, vol. 13,p. 19142. https://doi.org/10.1038/s41598-023-46602-1
El-Gaby, M.S.A., Bakry, M.M.S., Hussein, M.F., Faraghally, A.F., Khalil, A.M., Gad, M.A., and Drar, A.M., Curr. Chem. Lett., 2023, vol. 12, pp. 529–536. https://doi.org/10.5267/j.ccl.2023.3.003
Drar, A.M., Abdel-Raheem, Sh.A.A., Moustafa, A.H.,and Hussein, B.R.M., Curr. Chem. Lett., 2023, vol. 12,pp. 499–508. https://doi.org/10.5267/j.ccl.2023.5.005
Kamel, M.S., Aboelez, M.O., Elnagar, M.R.,Shokr, E.K., Selim, H.M.R.M., Abdel-Ghany, H.E.,Drar, A.M., Belal, A., El Hamd, A.M., and ElRemaily, M.A., Chem. Select, 2022, vol. 7, Article ID:e202203191. https://doi.org/10.1002/slct.202203191
Khodery, A., Mansour, E.S., Elhady, O.M., andDrar, A.M., J. Pest Manag., 2024, vol. 70, pp. 30–35. https://doi.org/10.1080/09670874.2021.1943048
Abbott, W.S., J. Econ. Entomol., 1925, vol. 18,pp. 265–267. https://doi.org/10.1093/jee/18.2.265a
Abd El-Lateef, H.M., Khalaf, M.M., Gouda, M.,Gad, M.A., Abdelhamid, A.A., Ismail, A.F., Amer A.A.,and Drar, A.M., Chem. Biod., 2024, vol. 2024, Article ID:e2024002. https://doi.org/10.1002/cbdv.202400218
Sun, Y.P., J. Econ. Entomol., 1950, vol. 43, pp. 45–53. https://doi.org/10.1093/jee/43.1.45
Scholz, C., Knorr, S., Hamacher, K., and Schmidt, B., J. Chem. Inf. Model., 2015, vol. 55, pp. 398–406. https://doi.org/10.1021/ci500681r
Johnson, T.O., Ojo, O.A., Ikiriko, S., Ogunkua, J., Akinyemi, G.O., Rotimi, D.E., Oche, J.R., and Adegboyega, A.E., Biochem. Biophys. Rep., 2021, vol. 28,Article ID: 101175. https://doi.org/10.1016/j.bbrep.2021.101175
Shaaban, S., Abdou, A., Alhamzani, A.G., Abou-Krisha, M.M., Al-Qudah, M.A., Alaasar, M., Youssef, I., and Yousef, T.A., Life, 2023, vol. 13, Article ID: 912. https://doi.org/10.3390/life13040912
Jarad, A.J., Dahi, M.A., Al-Noor, T.H., El-ajaily, M.M.,AL-Ayash, S.R., and Abdou, A., J. Mol. Struct., 2023,vol. 1287, Article ID: 135703. https://doi.org/10.1016/j.molstruc.2023.135703
Shaaban, S., Al-Faiyz, Y.S., Alsulaim, G.M., Alaasar, M.,Amri, N., Ba-Ghazal, H., Al-Karmalawy, A.A., andAbdou, A., Inorganics, 2023, vol. 11, Article ID: 321. https://doi.org/10.3390/inorganics11080321
Funding
This work was supported by regular institutional funding, and no additional grants were obtained.
Author information
Authors and Affiliations
Contributions
The author MSAEl-G comprehends the task, complies with all preparatory instructions, collects data, does formal analysis, composes the first draft, and edits. The authors GAMEl-HA and MAMAR assembling the initial draft, composing the criticism, editing, and arranging every recently created compound; writing (original draft), formal analysis, resources, supervision, research, and editing are among the tasks assigned to authors AA and MMSB. The author AMD, wrote the first draft, revised it, and made adjustments. The author MAG formal analysis, data length, resources, editing, and producing creative drafts.
Corresponding authors
Ethics declarations
This article does not contain any studies involving patients or animals as test objects.
Informed consent was not required for this article. No conflict of interest was declared by the authors.
Additional information
Publisher's Note. Pleiades Publishing remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
El-Gaby, M.S.A., Ali, G.A.M.EH., Reheim, M.A.M.A. et al. Insecticidal Evaluation, Molecular Docking, and Structure-Activity Relationship Study of Some Synthesized Thiazole-Owing Hydrazone Derivatives Against Spodoptera frugiperda under Laboratory Conditions. Russ J Bioorg Chem 50, 1037–1048 (2024). https://doi.org/10.1134/S1068162024030282
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1068162024030282