Abstract
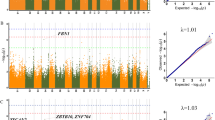
Backfat thickness (BFT) and average daily gain (ADG) are two important economic traits in commercial swine production. Identifying QTLs and uncovering the molecular mechanism for BFT and ADG would greatly help to speed up the breeding progress. In current breeding program, EBV for these two traits are calculated and formulated a comprehensive breeding index, which then be used to improve pig performance. Using Illumina PorcineSNP60 BeadChip, a pilot genomewide association studies (GWAS) for BFT and ADG in 83 Duroc pigs were performed. A total of 31 genome-wise significant SNPs were detected to be associated with BFT on SSC 4, 9, 11, 12 and 14, ten of which were coincident with previously reported QTL regions. There are two genome-wise loci prominently associated with ADG on SSC2 and SSC13, respectively. The two loci on SSC2 are well overlapped with the QTL regions previously reported. All the 31 significant SNPs associated with BFT are verified on 219 outbreed pigs, six SNPs reach an extreme significant level and seven SNP reaches a significant level, CACNA1E and ACBD6 are chosen as positional candidate genes. Our findings not only confirmed previously findings, but also revealed a number of novel SNPs associated with BFT and ADG. Two positional candidate genes CACNA1E and ACBD6 were identified for further study. These results would facilitate the identification of causative genes for BFT and ADG.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Dobney, K., Ervynck, A., and Rowley-Conwy, P., Pigs and Humans: 10000 Years of Interaction, Oxford: Oxford University Press, 2007.
Andersson, L. and Georges, M., Domestic-animal genomics: deciphering the genetics of complex traits, Nat. Rev. Genet., 2004, vol. 5, pp. 202–212.
Bidanel, J.P. and Rothschild, M., Current status of quantitative trait locus mapping in pigs, Pig News Inform., 2002, vol. 23, no. 2, pp. 39N–54N.
Gray, R.C., Tribble, L.F., Day, B.N., et al., Results of five generation of selection for low Backfat Thickness in swine, J. Anim. Sci., 1968, vol. 27, no. 2, pp. 331–335.
Hetzer, H.O. and Harvey, W.R., Selection for high and low fatness in swine, J. Anim. Sci., 1967, vol. 26, no. 6, pp. 1244–1251.
Jiang, L., Liu, J., Sun, D., et al., Genome wide association studies for milk production traits in Chinese Holstein population, PLoS One, 2010, vol. 5, no. 10. e13661
Maes, D.G.D., Janssens, G.P.J., Delputte, P., et al., Back fat measurements in sows from three commercial pig herds: relationship with reproductive efficiency and correlation with visual body condition scores, Livest. Prod. Sci., 2004, vol. 91, nos. 1–2, pp. 57–67.
Hutchens, L.K., Hintz, R.L., and Johnson, R.K., Genetic and phenotypic relationships between pubertal and growth characteristics of gilts, J. Anim. Sci., 1981, vol. 53 no. 4, p. 22.
Daw, E.W., Heath, S.C., and Lu, Y., Single-nucleotide polymorphism versus microsatellite markers in a combined linkage and segregation analysis of a quantitative trait, BMC Genet., 2005, vol. 6, suppl. 1. doi: 10.1186/1471-2156-1186-S1181-S1132
Amills, M., Vidal, O., Varona, L., et al., Polymorphism of the pig 2,4-dienoyl CoA reductase 1 gene (DECR1) and its association with carcass and meat quality traits, J. Anim. Sci., 2005, vol. 83, pp. 493–498.
Brym, P., Kamin-ski, S., and Wójcik, E., Nucleotide sequence polymorphism within exon 4 of the bovine prolactin gene and its associations with milk performance traits, J. Appl. Genet., 2005, vol. 46, no. 2, pp. 179–185.
Haegeman, A., Williams, J.L., Law, A., et al., Mapping and SNP analysis of bovine candidate genes for meat and carcass quality, Anim. Genet., 2003, vol. 34, no. 5, pp. 349–353.
Horin, P., Osickova, J., Necesankova, M., et al., Single nucleotide polymorphisms of interleukin-1 beta related genes and their associations with infection in the horse, Anim. Genomics Anim. Health, 2008, vol. 132, pp. 347–351.
Vallet, J.L., Freking, B.A., Leymaster, K.A., et al., Allelic variation in the erythropoietin receptor gene is associated with uterine capacity and litter size in swine, Anim. Genet., 2005, vol. 36, no. 2, pp. 97–103.
Becker, D., Wimmers, K., Luther, H., et al., A genome-wide association study to detect QTL for commercially important traits in Swiss Large White boars, PLoS One, 2013, vol. 8, no. 2. e55951
Ramos, A.M., Crooijmans, R.P.M.A., Affara, N.A., et al., Design of a high density SNP genotyping assay in the pig using SNPs identified and characterized by next generation sequencing technology, PLoS One, 2009, vol. 4, no. 8. e6524
Barrett, J.C., Fry, B., Maller, J., et al., Haploview: analysis and visualization of LD and haplotype maps, Bioinformatics, 2005, vol. 21, no. 2, pp. 263–265.
Kent, W.J., BLAT-the BLAST-like alignment tool, Genome Res., 2002, vol. 12, pp. 656–664.
Reich, D.E. and Lander, E.S., On the allelic spectrum of human disease, Trends Genet., 2001, vol. 17, no. 9, pp. 502–510.
Dragos-Wendrich, M., Moser, G., Bartenschlager, H., and Geldermann, H., Linkage and QTL mapping for Sus scrofa chromosome 11, J. Anim. Breed. Genet., 2003, vol. 120, suppl. 1, pp. 89–94.
Liu, G., Kim, J.J., Jonas, E., et al., Combined line-cross and half-sib QTL analysis in Duroc-Pietrain population, Mamm. Genome, 2008, vol. 19, no. 6, pp. 429–438.
Liu, G., Jennen, D.G.J., Tholen, E., et al., A genome scan reveals QTL for growth, fatness, leanness and meat quality in a Duroc-Pietrain resource population. Anim. Genet., 2007, vol. 38, no. 3, pp. 241–252.
Soupene, E., Serikov, V., and Kuypers, F.A., Characterization of an acyl-coenzyme A binding protein predominantly expressed in human primitive progenitor cells, J. Lipid Res., 2008, vol. 49, no. 5, pp. 1103–1112.
Holmkvist, J., Tojjar, D., Almgren, P., et al., Polymorphisms in the gene encoding the voltage-dependent Ca2+ channel Cav2.3 (CACNA1E) are associated with type 2 diabetes and impaired insulin secretion, Diabetologia, 2007, vol. 50, no. 12, pp. 2467–2475.
Dyson, M.C., Alloosh, M., Vuchetich, J.P., et al., Components of metabolic syndrome and coronary artery disease in female Ossabaw swine fed excess atherogenic diet, Comp. Med., 2006, vol. 56, no. 11, pp. 35–45.
Witczak, C.A., Mokelke, E.A., Boullion, R., et al., Noninvasive measures of body fat percentage in male Yucatan swine, Comp. Med., 2005, vol. 55, no. 7, pp. 445–451.
Norton, S.A., Zavy, M.T., Maxwell, C.V., et al., Insulin, growth hormone, glucose, and fatty acids in gilts selected for rapid vs. slow growth rate, Am. J. Physiol. 1989, vol. 257, pp. E554–E560.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Both authors contributed equally to this study and should be considered as co-first authors.
Rights and permissions
About this article
Cite this article
Long, Y., Ruan, G.R., Su, Y. et al. Genome-wide association study identifies QTLs for EBV of Backfat Thickness and average daily gain in Duroc pigs. Russ J Genet 50, 1308–1315 (2014). https://doi.org/10.1134/S102279541410007X
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S102279541410007X