Abstract
This lecture presents classical information and new data on the molecular events of the “basic” (core) cell cycle (CC) of plants. The impact of water deficit, CO2, light, and temperature on CC is briefly examined. Data on the regulation of cell proliferation by auxins, cytokinins, abscisic acid, gibberellins, brassinosteroids, and ethylene are presented. Commonality and peculiarities of the effect of phytohormones on CC in various organs and tissues are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Cell division is the basis for the existence of any multicellular organism, including higher plants. It is worth remembering here characteristic of this process, which was given by a contemporary of his discovery, Edmund Wilson: “due to cell division, in all higher forms at least, not only the continuity of life, but also the maintenance of the species, is effected; for through this beautiful mechanism the cell hands on to its descendants an exact duplicate of the idioplasm by which its own organization is determined.” [1, p. 178].
The specificity of plants is that they develop and grow mainly postembryonally, forming two types of organs: axial organs, such as roots and shoots, with indeterminate growth and, therefore, with a theoretically unlimited growth potential, and organs, such as leaves and flowers, with deterministic growth and a fixed final size. Cells of organs with deterministic growth stop dividing at a certain stage of ontogeny and proceed to elongation, the period of which is also determined [2]. Axial organs grow throughout the life of the plant. Such constant and long-term growth is due to the existence of apical meristems, the cells of which retain the ability to continuously proliferate. Leaving the meristematic zone, cells proceed to elongation and differentiation, and the transition to elongation (growth at a relatively higher rate) or growth inhibition occurs simultaneously for cells of all tissues located at the same distance from the meristem [3]. Therefore, in the whole plant, cell growth and proliferation must be coordinated, which is constantly observed and is an obvious fact. However, elucidation of the mechanisms of coordination of cell proliferation and growth still remains one of the most important tasks of experimental plant biology [4].
The specificity of organ growth undoubtedly affects the spatiotemporal characteristics of cell proliferation and the features of its regulation. In this lecture, for the most part, we will consider the molecular mechanics of the “basic” (core) cell cycle and its regulation under the influence of external factors and phytohormones.
CELL CYCLE: BASIC DEFINITIONS
Under cell proliferation (from lat. proles—offspring and fero—I carry), it is customary to understand their appearance through reproduction by division. The proliferative or more commonly referred to as the mitotic cycle (MC) is a complex of interrelated and chronologically determined events occurring in the cell during the period of time from the beginning of preparation for division, throughout mitosis itself, and until the completion of all processes associated with division in two daughter cells. Simplifying, we can give the following definition of MC: a time-ordered sequence of events occurring between two mitoses. This year marks the 70th anniversary of the publication of the classic work by Alma Howard and Stephen Pelc [5], in which they studied the incorporation of 32P into the DNA of Vicia faba root cells using autoradiography and revealed that the period of DNA synthesis occupies part of the interphase and is separated by time intervals from the beginning and end of mitosis. The authors proposed to divide the MC into four periods: mitosis proper (M), presynthetic period (G1), DNA synthesis (replication) period (S) and postsynthetic (or premitotic) period (G2).
In continuously multiplying populations of cells (embryonic tissues, meristems, etc.), the next MC begins immediately after the end of the previous one; that is, its duration coincides with the entire period of the cell’s existence as well as with its life (or cell) cycle (CC). However, in a full-grown multicellular organism, after the completion of the next round of divisions, some cells do not start preparing for the following MC but proceed to differentiation and the performance of tissue-specific functions. The portion of such cells can be quite significant, and the period of their life during which these cells perform specific functions in the absence of proliferation is not attributed to MC but is included in CC. The ratio of the number of proliferating (dividing) cells to the total number of cells in the population is called the “growth fraction” or “proliferative pool.” The terms MC and CC are often considered the same for those populations where these cycles coincide. We will use the term CC, however, emphasizing the existence of differences in certain cases.
The physiological role of the phase G1 consists in the implementation of growth and the processes of primary cell metabolism associated with it. At this stage, the processes of transcription and translation, the preparation of cells for DNA synthesis, are actively going on. Phase G1 is the most variable of all CC phases. There is a relationship between the intensity of cell divisions and the duration of the G1 period: as a rule, it lengthens with a decrease in the proliferative potential of cells.
In the S period, doubling (replication) of the nuclear DNA occurs, which is subsequently evenly distributed between two daughter cells. The content of DNA in the nucleus is usually expressed by the value “C.” In 1950, Hewson Swift introduced the designation “C,” referring to the “Constant”, DNA content in an organism various tissues' cells [6]. For the 1C value, we accept the amount of DNA in the unreplicated haploid set of chromosomes [7]. In plants, the amount of nuclear DNA varies greatly. The smallest amount of DNA (0.1 pg) per 1C was found in Genlisea margaretae [8], and the largest, 152.2 pg, belongs to Paris japonica [9]. “C” values have been determined for many plant species and are available, for example, from the Kew Botanical Garden database. Based on the data available in the literature, it can be concluded that the duration of the S period is the result of the interaction of several factors, such as the content of nuclear DNA and the length of individual replicons, the number and size of replicon families, and the synchronism of processes occurring in individual replicon families [10, 11].
Period G2 is usually shorter than G1- and S phases, and the variations in its duration are usually less than those of the G1 period. Events of the G2 phase are associated with the completion of the S period, reorganization of the cytoskeleton, and preparation for mitosis. In mitosis, an equal distribution of the doubled chromosomal material between daughter cells occurs due to the spindle apparatus functioning. Mitosis is the shortest (on average 1–3 h) and the least variable period of CC.
MOLECULAR EVENTS OF THE CELL CYCLE
Many processes in cell life are controlled by the reversible phosphorylation of certain target proteins. In humans, approximately 2% of all functionally active genes encode protein kinases, while that in Arabidopsis thaliana is approximately 4%, which certainly confirms the important role of protein kinases for animal and plant cells [12]. The presence of a large number of conditionally lethal mutant strains in the cell cycle, cdc (cell division cycle), of yeast-derived mutants has formed one of the directions in the study of molecular mechanisms of proliferation control. This direction is associated with the isolation of genes capable of correcting defects in cdc yeast mutants. Central to these genes are cdc2 in Schizosaccharomyces pombe or CDC28 in Saccharomyces cerevisiae, encoding a 34-kD protein (p34cdc2), a serine/threonine protein kinase whose activity, which in turn depends on phosphorylation or dephosphorylation of its own amino acid residues, is necessary for yeast cells to pass the boundaries between G1 and S, as well as G2 and M, periods of CC. It has been shown that the human gene, similar to cdc2+ S. pombe, is also (along with CDC28+) able to complement the mutation cdc2– in S. pombe [13]. Subsequently, several groups of proteins were found that are detected only in certain periods of CC, called cyclins. In response to proliferative stimuli, such as growth factors, forms of D-cyclins appear in mammalian cells, which interact with protein kinases Cdk4 and/or Cdk6 (Cdk, cyclin-dependent kinases), which leads to the phosphorylation of suppressor proteins, such as pRb (tumor growth suppressor, first discovered in retinoblastoma cells) and similar proteins. This leads to the activation of transcription factors (TF) of the E2F type, which are released from the complex with suppressor proteins, and the expression of genes necessary for the transition from G1 to S period in the CC [14]. Further, on the border of G1/S, cyclin E is expressed and binds to Cdk2. The activity of this complex is necessary to cross the indicated boundary and start DNA replication. Progress of the S period requires the interaction of Cdk2 with cyclin A; cyclin B works in mitosis. The functioning of Cdk–cyclin complexes is regulated by two families of Cdk inhibitors both under normal conditions and (mainly) under stress, DNA damage, telomere dysfunction, etc. Representatives of the INK4 family specifically bind to Cdk4 and Cdk6 and prevent the functioning of D-type cyclins, while proteins of the Cip/Kip family inhibit the Cdk2–cyclin E, Cdk2–cyclin A, Cdk1–cyclin A, and Cdk1–cyclin B complexes [14].
But what about plants? Back in the 1960s, Jack Van’t Hof demonstrated, on the separated tips of Pisum sativum roots, that the absence of sucrose in the medium or treatment with puromycin leads to the arrest of CC in the G1 or G2 periods, which is consistent with the paradigm of the main control points of the CC during the transitions G1/S and G2/M and indicates the need for protein synthesis for their successful passage [15]. Moreover, the cascade of protein kinase and protein phosphatase reactions that control CC is also universal for plant cells. According to modern concepts [16–19], the CC progression is ensured by the coordinated interaction of many regulatory proteins, among which the key positions, as in mammals, are occupied by cyclins (CYC) and serine/threonine protein kinases, called cyclin-dependent kinases (CDKs), expressed in certain time periods of CC. Plants have the largest number of CC regulatory proteins and their unique representatives; for example, only plants have B-type CDKs [20]. In A. thaliana, 13 proteins are assigned to CDK and 49 proteins are assigned to CYC, although not all of them are “seen” in the processes of CC regulation [20–22]. Each type has subclasses, such as A. thaliana—CYCA2;3 is the third representative of the second subclass of A-type cyclins—although it has not yet been shown that all plant species have all types of CYC and CDK. Even within the same subclass, CYCs differ not only in the “schedule” of their work in CC but also in the type and number of CDKs, inhibitors, and structural proteins with which they can interact, indicating a significant functional diversification of CC control mechanisms. Thus, for example, 92 potential variants of CDK–CYC complexes were found in A. thaliana according to the interactome data [21].
However, there are three types of cyclins for which a role in CC regulation has been postulated: CYCA, CYCB, and CYCD [23]. Specific CDK–CYC complexes regulate the transition at major CC checkpoints (G1/S and G2/M).
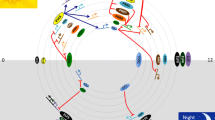
Complexes CDKA;1-CYCD ensure the transition from the G1 to S period by phosphorylation of the RBR1 (RETINOBLASTOMA-RELATED 1) protein with its subsequent proteolysis and, thereby, release TF E2Fa and E2Fb, which, in combination with dimerization proteins (DPa), provide “G1-transcription wave” [24], activating the expression G1/S-phase genes (Figs. 1a, 1b). It should be noted here that RB-like proteins (RBR) are an early acquisition in the course of evolution and are present in gymnosperms and angiosperms, unicellular algae, ferns, liverworts, and mosses. RBR1 regulates not only the G1/S transition but also functions as part of DREAM complexes, activating gene expression in the G2 period and genes not associated with CC and is also involved in the preservation of stem cells and the regulation of asymmetric divisions [19, 25].
Cyclins (CYC) and cyclin-dependent protein kinases (CDK) are the main regulators of CC. (a) Many CYCs and CDKs are expressed at specific times of CC and their combinatorial interaction regulates its progression. (b) Expression of genes supporting the G1/S transition of CC is provided by the functioning of the CDKA;1–CYCD complexes, which phosphorylate the RBR1 protein with its subsequent proteolysis, release of TF E2Fa and E2Fb, which, in combination with DPa proteins, provide a “G1-transcription wave.” (c) Many genes whose expression is associated with G2 and M periods of CC contain MSA-motifs (mitosis-specific activator) in the promoter region, which recognize TF MYB3R (mitotic CDK–CYC complexes phosphorylate and activate MYB3R), providing a “G2-transcription wave.” Gene expression of CDK and CYC and the functioning of their corresponding proteins are color-coded: pink, yellow, blue, and green, respectively, for G1-, S-, G2-, and M periods. Genes and their products that are constantly expressed during CC are marked in light lilac. KNOLLE is a gene of A. thaliana required for cytokinesis that encodes a protein similar to syntaxins. ORC (Origin Recognition Complex), CDC6 (Cell Division Control 6), and MCM (Minichromosome Maintenance) are genes encoding proteins of pre-replicative complexes (according to Komaki and Sugimoto [111] with additions and modifications).
CDKA;1-CYCA3 complexes control the passage of S phases, CDKB1–CYCA2/B2/B3 provide passage of the G2 period and initiation of mitosis, while CDKB2–CYCB1 and CDKA–CYCD3;1 are responsible for the passage of mitosis (Fig. 1a). It has recently been found that CYCB1 plays a key role in the organization of the mitotic network of microtubules, forming active complexes with CDKB2;2 and phosphorylating of the microtubule-associated proteins [26].
Data obtained on a synchronized cell culture of A. thaliana using DNA microarrays [27] reliably confirm the “G2-transcription wave.” Many genes whose expression is associated with the G2/M CC transition (Fig. 1c) contain the so-called MSA (mitosis-specific activator) motifs in the promoter region that recognize TFs of the MYB3R type (such as MYB3R1 and MYB3R4 from A. thaliana). Genes whose expression is activated in this way include, for example, CYCB1 and CYCB2, as well as KNOLLE, which activity is necessary for the formation of the cell plate after the completion of mitosis [28]. In turn, mitotic CDK–CYC complexes phosphorylate and activate MYB3R [29].
CDK function is accurately regulated: phosphorylation of Tre160 (or an equivalent threonine residue) by CDK-activating kinases (CAKs) leads to CDK activation [30]. Phosphorylation of the N-terminal tyrosine residue by protein kinase WEE1 leads to CDK inactivation, while WEE1 is usually more active in DNA damage (replication stress). It is assumed that WEE1 delays the passage of S phase to ensure DNA repair (Fig. 2) [31]. In addition, in maize and tomato, WEE1 activity is associated with endoreduplication [32].
Main points of the functioning of activators and inhibitors of CDK and the exits of cells from the MC. G0 is a state of proliferative quiescence of cells inherent in differentiated, aging, and stem cells. In G0, exit from G1 and G2 periods is possible. Under certain conditions, such as hormonal stimuli, cells can return from G0 to MC. CAK are CDK-activating protein kinases; WEE is CDK-inactivating protein kinase; KRP and SIM are inhibitors of CYC–CDK complexes (according to Velappan et al. [112] with additions and modifications).
Additionally, CDK activity is specifically inhibited by CKIs (CDK Inhibitors). In plants, Kip-related proteins (KRP or CDK inhibitors, or ICKs) contain a conserved domain similar to that of mammalian Cip/Kip family proteins. In A. thaliana, KRP proteins encode seven genes [33] and inhibit CDKs in G1/S (Fig. 2), which leads to MC arrest and commits cells to endoreduplication [16, 21, 32]. A second family of CDK inhibitors known as SIAMESE (SIM proteins) has been discovered in connection with the analysis of A. thaliana plant mutants with impaired development of leaf trichomes [34]. Many representatives of SMR (SIM-related) proteins can interact with CYCD and CDKA;1 [21, 34], but SIM proteins interact in general with CDKB1;1 [21, 32] (Fig. 2). The ability of SMR proteins to regulate CDKA–CYCD complexes is probably important for adequate control of CC under the action of biotic and abiotic stressors [35]. Interestingly, for example, SMR4 interacts with CYCD3;1 and slows down G1 in the last asymmetric division preceding the symmetrical division forming stomatal guard cells [36].
One of the earliest events in CC is the licensing of DNA (more precisely, chromatin), the process of active access to chromatin and “anchoring” of proteins and protein complexes on it, including DNA replication factors and chromatin-modifying proteins [24]. Replication of the eukaryotic genome requires the activation of thousands of replication origins (ORIs) to which initiation complexes must bind. These primary regulatory events, the association of pre-replication complexes (pre-RCs) with each potential origin of replication, are activated in the late telophase almost immediately after the distribution of the two newly formed nuclei into daughter cells. Sequential assembly of a conserved multisubunit complex containing six ORC (Origin Recognition Complex) proteins, ATP binding protein CDC6 (Cell Division Control 6), chromatin licensing and DNA replication factor 1 (CDT1), and a heterohexamer of six MCM helicase proteins (Minichromosome Maintenance) occurs [19, 37].
In the early G1 period, CDK activity should be low, which is necessary to license replication origins. Later, CDK activity increases due to an increase in the level of required CYCs of the G1 period (see Fig. 1a), reaching a threshold sufficient to inactivate RBR1 and start (firing) the work of replicons, that is, the start of the S period. Progress of the S period is provided by specific CDK–CYC complexes; in the G2 period, CYCs of the M period are actively transcribed, the activity of mitotic CDKs increases, and mitosis begins. Then, to complete mitosis, degradation (proteolysis involving 26S-proteasomes) of mitotic CYCA1, CYCB1, and CYCB2 classes is required. These CYCs contain a specific sequence of amino acid residues (D-box, destruction box) that determines the sensitivity to ubiquitination and is recognized by APC/C (Anaphase-Promoting Complex/Cyclosome) E3 ubiquitin ligase, which leads to the attachment of ubiquitin and rapid proteolytic degradation of mitotic CYCs [28]. As a result, CDK activity again falls to the level of the early G1 period (Fig. 3).
Scheme of regulation of MC and endocycles. In G1, CDK activity increases by increasing the level of CYCs required for this, reaching a threshold sufficient to the start of the S period. Further, the activity of mitotic CDKs increases and mitosis begins. Mitosis is completed by degradation (proteolysis involving 26S-proteasomes) of mitotic CYCs, as a result of which CDK activity drops to the level of early G1. During endocycles, CDK activity does not reach the level necessary to trigger mitosis, while maintaining the activity of CDKA;1 in complexes with CYC D-type and CYCA3, which ensures the passage of the S period. The successive the GE and S periods of endocycles end with a drop in APC/C activity. The cell goes into proliferative quiescence (G0) (according to Breuer et al. [41] with modifications).
In the M period, parallel to the segregation of chromosomes, there is the preparation for the physical separation of the two daughter cells (cytokinesis). Despite the relative short duration of the M period (2–3 h), its implementation is highly coordinated and involves a huge number of molecular participants and events leading to significant, including visible, structural rearrangements of the cell (chromatin condensation, formation of the spindle apparatus, destruction of the nuclear envelope, segregation of chromosomes, phragmoplast formation, etc.) [37, 38].
Cells can leave the MC and go into the G0 state (proliferative quiescence), which is characteristic of differentiated, aging, and stem cells (Fig. 2), because, as already noted, cell growth and proliferation must be coordinated in the whole plant. The consequence of this interdependence is different modifications of CC, which are most suitable for specific stages of development and physiological needs of certain types of tissues or cells [39]. One of the most common modifications of CC is endoreduplication (synonyms: endoreplication, endocycles, endoploidization), in which multiple rounds of DNA replication occur without subsequent chromosome segregation and cytokinesis [17, 32, 40, 41]. Endoreduplication is usually seen in large, metabolically active, or highly specialized cells, although it also occurs in cells that do not fit this description.
Endoreduplication cycles are modified MCs in which the stages of DNA replication and mitosis are changed. It is important that, during endocycles, CDK activity does not reach the level required to trigger mitosis (Fig. 3). This occurs due to a decrease in the transcription of pre-mitotic and mitotic CYCA, CYCB, and CDKB as well as a decrease in CDKB activity under the influence of inhibitors of the SMR family; at the same time, CDKA;1 activity is preserved due to the formation of complexes with CYC of D‑type and CYCA3, which ensures the passage of the S period [34, 40]. Endocycles consist of successive specific GE- and S periods (Fig. 3), while oscillations in antiphase of the activities of CYC–CDK complexes and APC/C ubiquitin ligase are observed. At the end of endocycles, APC/C activity decreases, which allows the cell to go into a state of proliferative quiescence, G0; the presence and activity of CYC–CDK complexes remains unclear [41].
It turned out that such a protein kinase as CDKG2 has a positive effect on endoreduplication, while CDKB1;1 directly phosphorylates CDKG2 and reduces its stability. The molecular mechanism of action of CDKG2 is still unclear [42]. Instability of the plastid genome caused by mutations reca1why1why3 or ciprofloxacin also induces endocycles. The signal goes through SOG1 (TF that responds to DNA damage) and leads to activation of SMR5 and SMR7 [43]. Interestingly, SOG1 is also involved in the induction of endoreduplication in A. thaliana caused by strong salinity [44]. Under these conditions, oxidative stress develops, breaks are formed in the double-stranded DNA molecule, and SOG1 is activated, which leads to a change in the level of mRNA and proteins: a decrease in CDKB1;1, CDKB2;1, and CYCB1;1 and a simultaneous increase in WEE1, CCS52A (APC/C activator), and E2Fa.
For plants, mixoploidy (endopolyploidy), i.e., the presence of cells with different DNA contents in one tissue, is a widespread phenomenon [45]. For example, a population of leaf epidermal cells of A. thaliana is mixoploid [46] and, in the Col-0 ecotype, contains nuclei with DNA amounts of 2C (36%), 4C (48%), 8C (15%), and 16C (1%) (Fig. 4).
Mixoploidy of A. thaliana (L.) Heynh. leaf epidermis. Micrograph of Bénédicte Desvoyes (after Gutierrez [113] with modifications).
At present, the physiological significance of endopolyploidy is not yet fully understood and, therefore, is being intensively studied. Possibly, endopolyploidization is a way to accelerate plant growth under specific environmental conditions since there is a certain relationship between the amount of nuclear DNA and cell size [46, 47]. For cultivated in vitro cells, mixoploidy is also a widespread phenomenon, which can be determined both by specific cultivation conditions (nutrient medium components, temperature, etc.) and by the ploidy of the initial explant.
EXTERNAL FACTORS IN REGULATION OF PLANT CELL CYCLE
Higher plants have a eukaryotic type of CC. This means that the work of a genetically determined program is modulated by external factors and stimuli. External factors such as water availability, nutrients, CO2, light, and temperature affect cell growth and division [48]. However, changes in CC do not always lead to changes in the size of plant organs. Thus, in young A. thaliana seedlings, nitrate has been shown to induce SMR1/LGO (CDK inhibitor) gene expression as early as 3 days after germination, which leads to the induction of endoreduplication and increase in the amount of DNA and cell size. In lgo-2 mutant plants, whose cells are unable to support active endoreduplication, the cotyledon size is similar to that of wild-type plants, which is achieved through cells' division [49].
With water deficiency, the rate of cell division slows down, the size of meristems and the number of leaf cells decrease, and the proportion of cells located in the G1 period grows, which indicates an increase in its duration and/or a stop of the CC at the border of G1/S. At the same time, the duration of S, G2, and M periods changes to a lesser extent [48]. Water deficiency in many plant species leads to the arrest of both MC and endocycles due to a decrease in CDK activity, although endoredupication can be stimulated under mild water stress. For example, in A. thaliana mesophyll cells, this allows the plant to maintain leaf area during drought through a ploidy-mediated increase in cell size [17]. Drought quickly induces a high level of gene transcripts of SMR and KRP, CDK inhibitors, and inhibits the expression CDKA/B and CYC in young Arabidopsis leaves [50].
Increasing CO2 concentration accelerates the growth of many plant species; this is apparently determined by an increase in the productivity of photosynthesis and, consequently, the concentration of carbohydrates [51, 52]. At the same time, an increase in the size of leaves and roots is associated with an increase in the proportion of dividing cells in meristems.
The first cardinal change that an etiolated seedling encounters when it reaches the soil surface is light. We should recall here the classical data that illumination of etiolated seedlings causes a wave of divisions [53]. For example, already after 6 h of illumination of 3‑day-old etiolated A. thaliana seedlings, there is a coordinated increase in the expression of genes associated with translation and then genes, regulators of CC, involved both in the transition from G1 to S phase and from G2 to mitosis and a decrease in the expression of CDK inhibitor genes [54].
A decrease in the intensity of incident light causes a decrease in the final number of cells in the leaves of dicotyledonous plants [48]. At the same time, a similar effect was observed when the light intensity was reduced by 40% using neutral light filters or when the equivalent area of the photosynthetic surface of the leaf was covered. Both of these effects reduce the relative rate of cell division and do not affect the duration of a leaf’s proliferative activity. A decrease in the amount of absorbed light is accompanied by a decrease in photosynthesis and sugar content in leaf tissues [48]. An interesting recently established fact is that CDKA in A. thaliana, independently of CC, controls light-induced tropism and chloroplast movements, probably by influencing the organization of the cytoskeleton. This function requires low CDKA activity compared to its activity in MC [55].
Among environmental variables, the duration of the light phase is more predictable and has a fixed cyclical nature. The duration of the light period controls the growth and development of plants, ensuring the regulation of the expression of many genes through the work of the internal, light-dependent circadian clock. The circadian clock modulates CC time by rhythmically binding TF, TOC1/PRR1 (TIMING OF CAB EXPRESSION1/PSEUDO RESPONSE REGULATOR 1) to the gene promoter CDC6, one of the components of the pre-replicative complex [56].
Studies into the influence of the spectral characteristics of light on CC are intensively carried out on unicellular green algae. It has been shown that light in the blue region of the spectrum (400–500 nm) inhibits cell division of Protosiphon botryoides through a signal transmitted from a yet unidentified photoreceptor [57] and red light stimulates cell division of Chlamydomonas reinhardtii [58]. Continuous far red light eliminates the negative effect of continuous blue light on synchronous divisions in tobacco cell suspension culture, which is associated with phytochrome-induced auxin biosynthesis [59]. Light, in general, promotes cell proliferation by stimulating the activity of photoreceptors, which leads to the suppression of the activity of CC inhibitors. Light also affects endoreduplication, inhibits endocycles. For example, more endocycles run in the dark in hypocotyls A. thaliana, Brassica oleracea, and Pisum sativum [60].
The most studied environmental factor influencing CC is temperature. A lot of data shows that the duration of the total CC decreases (hence, the rate of cell division increases) with increasing temperature [48]. However, despite changes in the rate of cell division, the final number of cells in an organ does not change over a wide temperature range. In fact, an increase in the rate of cell division with an increase in temperature is compensated by a decrease in the duration of the period of cell proliferation [61]. It has been shown that growth retardation at low temperature is the result of a proportional increase in the duration of CC periods and the morphological characteristics of the meristem do not change at the minimum constant temperature. The concept of critical temperature points is proposed, on the basis of which is determined the optimal temperature for CC [62].
PHYTOHORMONES—REGULATORS OF PLANTS’ CELL CYCLE
Phytohormones are considered as intracellular pleiotropic factors that control almost all processes in plant cells. At the same time, in order to show the effect of a particular phytohormone, as a rule, they are used exogenously. The effect of phytohormones on the growth and development of plants, including cell proliferation, depends on the organ and tissue, the stage of ontogenesis, the endogenous ratio of phytohormones, the light regime, etc. All this can change the direction of phytohormones' action [63]. It has now been convincingly shown that different phytohormones can influence the same processes; however, paradoxically, their signaling pathways do not work redundantly. The modern paradigm determines that the signals of different phytohormones are integrated at the level of the gene network, and their transduction is associated with “communication” (cross-talk) of signaling pathways [64–66].
Auxins and Cytokinins Are the Most Important Regulators of the Cell Cycle
Participation of auxins and cytokinins (CK) in the regulation of CC is the most researchable topic. In A. thaliana, the key role of auxin in the formation of lateral roots is shown. The auxin concentration significantly increases in the founder cells of the pericycle, which leads to the induction of divisions of these cells, the formation of primordia, and the development of an active meristem, which subsequently forms a lateral root [67].
In A. thaliana plants, auxins (natural and synthetic) have been shown to induce CDKA;1 gene transcription in root apexes, while CK induces the transcription in the elongation zone [68]. Auxin promotes cell division in the root meristem and has a maximum concentration in the quiescent centre, where it is synthesized, a high concentration in the proliferative domain, and a relatively low concentration in the zone of cell elongation and differentiation. At the same time, the use of low concentrations of auxin (200 nM IAA) stimulates mitotic activity, which leads to an increase in the meristematic zone [65]. In addition, a double mutant with loss of auxin biosynthesis gene functions of WEI8/TAA1 and TAR2 (wei8tar2) does not have an identifiable meristem, and IAA treatment partially restores it [69]. In the transition domain of the root apex, where cells change from normal MC, ending in chromosome segregation and cytokinesis, to endoreduplication cycles, any disruption in auxin biosynthesis, transport, or signaling facilitates the transition from MC to endocycles, while an increase in auxin levels delays the onset of endocycles and the transition into the zone of cell elongation [65].
A fairly high basal level of transcripts and CDKA;1 protein is found in all living plant cells, even in non-actively dividing [68], which apparently reflects their unique ability to return to division. Auxin determines the possibility of cell division by stimulating the accumulation of a sufficient concentration of CDKA;1, which, after interacting with certain CYCD and CYCA, will provide its activity to pass the G1/S checkpoint and the S period itself (see Figs. 1, 2).
Cell proliferation in the root meristem is negatively affected by CKs. A decrease in CK synthesis and/or impaired signal transduction leads to an increase in the meristematic zone and a delay in the onset of endoreduplication [65]. It should be noted that there is another view on the mechanism of CK action in the meristematic zone of the root. It has been shown that trans-zeatin slows down the rate of root growth and the transition of cells to elongation, lengthening the MC. In other words, any changes in the concentration of endogenous CK or changes in CK signaling lead to changes in the duration of MC [70]. However, the dominant view is that CK have the opposite effect on MC in the apical meristems of shoots and roots [71].
Along with the root apical meristem, in vitro cultured protoplasts and plant cells are frequently used systems in studies of the role of auxins and CK in CC regulation. On protoplasts from alfalfa leaf cells, it was demonstrated that the number of dividing cells depends on the auxin concentration [72]. However, CK (zeatin) was necessary for the normal passage of the S period and completion of mitosis, which was expressed in an increase in the activity of the corresponding CDKs, namely CDKA;1 and CDKB1;1. In the absence of CK, the CDKA;1 protein was synthesized but did not exhibit phosphorylating activity [72]. The combined action of CK and auxin significantly increased transcription of CDKA;1 in cultivated tobacco mesophyll protoplasts [68]. A study on A. thaliana suspension cell culture into the effects of auxin (NAA), CK (kinetin), and/or sucrose showed that the expression of genes encoding components of the CC regulatory network responds differently depending on the combination of effectors [73]. Expression of CDKA;1, CYCA2;1, and CYCD was induced by sucrose, while CYCD3;1 was by kinetin and sucrose; expression of CDKB1;1 and CYCB1;1 significantly increased with the simultaneous action of NAA and kinetin. The level of CDK inhibitor gene transcripts of the KRP family significantly decreased under the influence of phytohormones and sucrose [73].
The combined effect of auxin and CK was also observed in experiments with alfalfa leaf explants. Treatment of leaf explants with 2,4-D slightly increased the general CDK activity, as determined by histone H1 phosphorylation, and the combination of CK and auxin significantly increased CDK activity [74]. The presented data show that the CK is necessary in the indicated objects for the passage of the S phase and G2/M transition. Moreover, the CK signal is also probably important for the transition between all phases of CC since in the synchronized suspension culture of tobacco cells (BY-2) there is an oscillation of the intracellular concentration of endogenous CK (mainly in the form of trans-zeatin) with a maximum before the G1/S and peaks before the S/G2, G2/M, and M/G1 boundaries [75].
The response of whole plants to exogenous phytohormones is complex and, in particular, depends on the organ and tissue and does not necessarily repeat the response of cells cultured in vitro. However, along with cell culture, the treatment of A. thaliana seedlings with zeatin caused an increase in CYCD3 mRNA levels in plant tissues committed to division [76]. Overexpression of CYCD3;1 in transgenic plants increases mitotic activity and reduces the number of endocycles. In cell culture, the overexpression of CYCD3;1 leads to G2 phase prolongation and delayed activation of mitotic genes, suggesting that CYCD3;1 ensures the formation of CDKA;1–CYCD complexes necessary for the transition from G1 to S period and activation of S-phase gene expression [77]. Interestingly, in A. thaliana mutant plants with a high content of endogenous CK, a threefold increase in the level of CYCD3 mRNA was noted compared to wild-type plants, while it is possible to obtain callus tissues and maintain their proliferation without exogenous CK from explants of A. thaliana transgenic plants with constitutive expression of CYCD3 [76].
Auxins and CK also affect CYC and CDK gene expression and CDK inhibitors in other experimental systems. After 1 day of cultivation on a medium that induces callus formation (medium with 2,4-D), in explants of hypocotyls of 5-week-old A. thaliana plants, increased expression of many genes associated with CC, including CYCB1;1, CYCB2;2, and the level of KRP1 and SMR2 transcripts (CDK inhibitors) decreased [78]. Similarly, treatment A. thaliana seedling with auxins and CK led to a decrease in KRP4 gene transcription [79].
In addition, auxins and CK also implement other ways to control CC. Auxins stabilize the level of the E2Fb protein, which regulates the passage of S and M periods, activating the expression of CDKA;1 and CDKB1;1. At a high level of E2Fb, the duration of CC is reduced [80]. It was shown that the F-box protein SKP2A (S-Phase Kinase-Associated Protin 2A) A. thaliana, which controls the stability of E2Fc/DPB, transcription repressors of genes that regulate CC, can directly bind to auxin, which leads to ubiquitination and proteolysis of SKP2A together with its targets, E2Fc/DPB, and this has a positive effect on the going through of CC [81]. That is, we can talk about the possibility of regulation of CC by auxin, bypassing the classical module of its signaling: Aux/IAA-ARF-SCFTIR1/AFB [82]. In this regard, it should be noted that the ABP1 (AUXIN BINDING PROTEIN 1), one of the first characterized proteins that binds auxin with high affinity and meets the criteria of the receptor, opens its functions in the regulation of CC. The BY-2 tobacco cell culture showed that ABP1 inactivation causes CC arrest in the G1 period and a sharp drop in CYCD3;1 expression [83]. In addition, it was reported that auxin, via transmembrane kinases, activates signal transduction through the Mitogen-Activated Protein Kinase (MAPK) cascade and controls the orientation of cell divisions during the lateral roots' formation [84].
In the transition zone of the A. thaliana root apex, CK activates two TFs, ARR1 (ARABIDOPSIS RESPONSE REGULATOR 1) and ARR12, which leads to the induction of SHY2/IAA3, repressors of auxin signal transduction. This inhibits auxin signaling and reduces the expression of auxin transporters, which leads to the arrest of cell division [71]. However, CK can have a direct effect on CC, which is not determined by the influence on auxin signal transduction. It was shown that CK-activated ARR2 directly binds to the promoter of the CCS52A1 gene and induces its expression. The CCS52A1 protein, an activator of APC/C E3 ubiquitin ligase, is expressed in the transition and elongation zones of the root and stimulates the degradation of mitotic cyclins, which leads to the triggering of endocycles [85].
However, for A. thaliana shoot apical meristem, it has recently been discovered that, during the G2/M transition, CK facilitates the movement of TF MYB3R4 into the nucleus, which activates the expression of mitotic genes that control the “proper” going through of mitosis and cytokinesis (see Fig. 1). The rapid accumulation of MYB3R4 in the nucleus depends on importin proteins and coincides with the transient peak of CK concentration [86]. There are also new players in the implementation of the CK signal. Thus, it was shown that CKG (CYTOKININ-RESPONSIVE GROWTH REGULATOR), known as TF bHLH137, controls MC and cell growth and works after ARR2, inducing transcription of WEE1 [87].
Abscisic Acid, Gibberellins and Brassinosteroids: Inhibitors and Stimulators of the Cell Cycle
The ambiguous effect of ABA on cell proliferation was discovered long ago. Along with the manifestation of an inhibitory effect, ABA stimulated growth, cell division, and DNA synthesis in some systems. As for many phytohormones, this is determined by the ABA concentration and the sensitivity of plant tissues and cells [88]. The undisputed favorite in explaining the negative effect of ABA on CC is the fact that, under its influence, gene expression of KRP/ICK, which encode CDKA inhibitors, increases. This leads to a disruption of the G1/S transition. In addition, ABA suppresses the expression of CYCB1;1 (check point G2/M) [89]. ABA contributes to the increase in the size of A. thaliana leaf epidermal cells in plants not showing water stress, which is associated with the stimulation of endoreduplication [90].
Treatment of the root apex with a low concentration of ABA (0.5 μM) increases the size of the meristem, while a high concentration (30 μM) leads to the opposite result [65]. Interestingly, the negative effect of ABA on meristem size can be partially eliminated by cotreatment with glutathione. ROS are probably involved in the implementation of this ABA effect [65]. Curiously, ABA treatment of the aerial parts of A. thaliana plants leads to an increase in the size of the root meristem and induces the expression of CYCB1;1::GFP (marker of mitosis). It is assumed that this is caused by increased basipetal auxin transport [91]. More recently, a new regulatory module has been discovered, where TF ABI4 directly inhibits the activity of promoters of at least two major cell cycle genes, CYCB1;1 and CDKB2;2 [92].
It is well known that ethylene and ABA mutually influence each other’s synthesis and there are points of “communication” of their signal-transduction pathways. However, unexpected was the result obtained on A. thaliana wild-type suspension cell cultures (Col-0) and mutants at one of the ethylene receptors, etr1-1 and the signal transduction component following the receptors, ctr1-1 (constitutive triple response1-1), in which the ethylene signaling pathway is constantly active. For wild-type cells, exogenous ABA acts as an inhibitor of DNA synthesis and cell proliferation. If the ethylene signal is not fully perceived (etr1-1), then ABA significantly enhances cell proliferation. When the ethylene signaling pathway is constitutively active (ctr1-1), cells more often switch to endoreduplication, but the addition of ABA promotes MC reactivation [93].
Gibberellins play a significant role in coordinating cell growth and proliferation under normal gravity, which does not work under microgravity [94]. In the root meristem, gibberellins are necessary both for the going through of MC and for cell growth in the elongation zone, stimulating the destruction of growth-repressing DELLA proteins [65]. In A. thaliana seeds, gibberellin GA4 induces CYCD1;1 expression and several DNA replication factors. During vegetative growth, DELLA proteins inhibit cell proliferation by inducing the expression of CC inhibitors: SMR1, SMR2, and KRP2. In response to osmotic stress, DELLAs induce cell differentiation and the onset of endoreduplication in leaves by suppressing the activity of genes encoding APC/C inhibitors [95].
Brassinosteroids, as a rule, act as positive regulators of CC [96]. Epibrassinolide increases the expression of CYCD3;1 and, thereby, enhances the intensity of cell division and can replace CK in callus and suspension cell cultures of A. thaliana [97]. At the same time, for A. thaliana leaf cells, brassinosteroids are important for the processes of division, elongation, and differentiation. The balance between cell proliferation and differentiation depends on the phytohormone concentration and the functioning of the BRI1-dependent (brassinosteroids' receptor) signaling system [98]. In the root apical meristem, the effect of brassinosteroids also depends on the concentration. Plants treated with 0.4–4 nM brassinolide have a shorter meristem than untreated plants, while 0.04 nM phytohormone increases the size of the meristematic zone. Also, the effect of brassinosteroids on proliferation or differentiation depends on the cell type: in the epidermis, induction of proliferation; in the stele, stimulation of differentiation [65].
Does Ethylene Inhibit or Stimulate Cell Proliferation?
In vegetative growth, ethylene appears to play a dual role, stimulating and inhibiting growth, depending on species, tissue and cell type, developmental stage, hormonal status, and environmental conditions [64]. When ethylene is considered as a “stress hormone” and stress other than flooding, it regularly functions as an inhibitor of cell elongation and division, in particular, in leaf tissues, which is of great adaptive importance [50].
At present, there is no unambiguous understanding of the role of ethylene in the control of CC. It has been shown that ethylene inhibits nuclear DNA replication and cell division in pea seedlings [99] and induces the programmed death of cultivated tobacco cells during certain periods of CC at concentrations (17 700–35 000 µL/L) that are three to four orders of magnitude higher than physiological ones [100]. Ethylene inhibits cell proliferation in the A. thaliana root meristem, in particular, by activating the expression of KRP1 [101]. It was shown, taking into account the ability of CK to induce ethylene biosynthesis and the negative effect of CK on the proliferation of root meristem cells, that ethylene only partially contributes but is not required for the manifestation of the effects of CK. However, ethylene inhibits the stimulating effect of CK on cell division and the growth of cotyledons in etiolated A. thaliana seedlings [102].
Nevertheless, ethylene can have a positive effect on CC events. Ethylene activates DNA synthesis and endoreduplication but inhibits cytokinesis in the epidermis of cucumber hypocotyls [103]. Ethylene induces cell division in the quiescent centre of the A. thaliana root meristem [104] and epidermal cells of etiolated cucumber hypocotyls after short-term exposure [105]. Ethylene activates the proliferation of poplar cambial cells [106] and cells of the quiescent centre of maize root meristem after removal of the root tip [107]. Ethylene, in cooperation with the receptor kinase PXY, promotes an increase in the division rate in the “vascular” meristem during the formation of the phloem [108]. On A. thaliana cultured cells of the wild-type Col-0 and mutant etr1-1 carrying a point mutation in the ethylene binding site of the ETR1 receptor (one of five receptors), it was shown that an inhibitor of ethylene binding to receptors (1-methylcyclopropene, 1-MCP) significantly reduces, and ethylene increases, the viability of cells of both genotypes, similarly affecting growth indices. Ethylene significantly increases the number of S-phase cells in etr1-1 culture, while 1-MCP reduces it. Other ethylene receptors, rather than ETR1, are probably responsible for the control of cell proliferation [109]. Exogenous ethylene stimulates the transition to S phase of A. thaliana cultured cells only when the production of endogenous ethylene is low. In addition, a significant correlation was shown between ethylene production and the growth rate of several suspension cell culture strains. At the same time, ethylene synthesis apparently precedes cell proliferation [110].
CONCLUSIONS
Note that we did not consider the effect of jasmonates, strigolactones, and karrikins on CC in this lecture. This was mainly due to limited space. On the other hand, jasmonates, strigolactones, and karrikins are, to a greater extent, “stress molecules”, and it is probably necessary to talk about their effect on CC separately. Nevertheless, if we summarize the above data, it becomes clear that the conservative molecular mechanics of CC still continues to be replenished with new TFs, a new role for previously annotated CYCs and CDKs, etc. As for the influence of phytohormones on CC, it is obvious that the effects depend on the organ, tissue, external conditions, gradients of other phytohormones, etc. There is no ideal, abstract cell or a standard effect of one or another phytohormone in relation to this cell. Cells cultivated in vitro are also not devoid of the epigenetic landscape of the original explant. Do different cells use common or unique CC regulation tools? The question remains open.
The research was carried out within the state assignment of the Ministry of Science and Higher Education of the Russian Federation (theme no. 122042600086-7).
This article does not contain any studies involving humans and animals as subjects. The authors declare that they have no conflicts of interest.
REFERENCES
Wilson, E.B., The Cell in Development and Inheritance, New York: Macmillan, 1911.
Vercruysse, J., Baekelandt, A., Gonzalez, N., and Inzé, D., Molecular networks regulating cell division during Arabidopsis leaf growth, J. Exp. Bot., 2020, vol. 71, p. 2365. https://doi.org/10.1093/jxb/erz522
Ivanov, V.B. and Dubrovsky, J.G., Longitudinal zonation pattern in plant roots: conflicts and solutions, Trends Plant Sci., 2013, vol. 18, p. 237. https://doi.org/10.1016/j.tplants.2012.10.002
Sablowski, R., Coordination of plant cell growth and division: collective control or mutual agreement?, Curr. Opin. Plant Biol., vol. 34, p. 54. https://doi.org/10.1016/j.pbi.2016.09.004
Howard, A. and Pelc, S.R., Synthesis of deoxyribonucleic acid in normal and irradiated cells and its relation to chromosome breakage, Heredity (Edinb.) Suppl., 1953, vol. 6, p. 261.
Swift, H.H., The constancy of desoxyribose nucleic acid in plant nuclei, Proc. Natl. Acad. Sci. USA, 1950, vol. 36, p. 643. https://doi.org/10.1073/pnas.36.11.643
Greilhuber, J., Dolezel, J., Lysák, M.A., and Bennett, M.D., The origin, evolution and proposed stabilization of the terms “genome size” and “C-value” to describe nuclear DNA contents, Ann. Bot., vol. 95, p. 255. https://doi.org/10.1093/aob/mci019
Kejnovsky, E., Leitch, I.J., and Leitch, A.R., Contrasting evolutionary dynamics between angiosperm and mammalian genomes, Trends Ecol. Evol., 2009, vol. 24, p. 572. https://doi.org/10.1016/j.tree.2009.04.010
Pellicer, J., Fay, M.F., and Leitch, I.J., The largest eukaryotic genome of them all?, Bot. J. Linn. Soc., 2010, vol. 164, p. 10. https://doi.org/10.1111/j.1095-8339.2010.01072.x
Bechhoefer, J. and Rhind, N., Replication timing and its emergence from stochastic processes, Trends Genet., 2012, vol. 28, p. 374. https://doi.org/10.1016/j.tig.2012.03.011
Greenberg, A. and Simon, I., S phase duration is determined by local rate and global organization of replication, Biology, 2022, vol. 11, p. 718. https://doi.org/10.3390/biology11050718
Lehti-Shiu, M.D. and Shiu, S.-H., Diversity, classification and function of the plant protein kinase superfamily, Philos. Trans. R. Soc. Lond., B, Biol. Sci., 2012, vol. 367, p. 2619. https://doi.org/10.1098/rstb.2012.0003
Cross, F., Roberts, J., and Weintraub, H., Simple and complex cell cycles, Annu. Rev. Cell Biol., 1989, vol. 5, p. 341. https://doi.org/10.1146/annurev.cb.05.110189.002013
Satyanarayana, A. and Kaldis, P., Mammalian cell-cycle regulation: several Cdks, numerous cyclins and diverse compensatory mechanisms, Oncogene, 2009, vol. 28, p. 2925. https://doi.org/10.1038/onc.2009.170
Van’t Hof, J., The regulation of cell division in higher plants, Brookhaven Symp. Biol., 1973, vol. 25, p. 152.
Polyn, S., Willems, A., and De Veylder, L., Cell cycle entry, maintenance, and exit during plant development, Curr. Opin. Plant Biol., 2015, vol. 23, p. 1. https://doi.org/10.1016/j.pbi.2014.09.012
Scholes, D.R. and Paige, K.N., Plasticity in ploidy: a generalized response to stress, Trends Plant Sci., 2015, vol. 20, p. 165. https://doi.org/10.1016/j.tplants.2014.11.007
Carneiro, A.K., Montessoro, P.D.F., Fusaro, A.F., Araújo, B.G., and Hemerly, A.S., Plant CDKs—driving the cell cycle through climate change, Plants, 2021, vol. 10, p. 1804. https://doi.org/10.3390/plants10091804
Sablowski, R. and Gutierrez, C., Cycling in a crowd: coordination of plant cell division, growth, and cell fate, Plant Cell, 2022, vol. 34, p. 193. https://doi.org/10.1093/plcell/koab222
Blomme, J., Inzé, D., and Gonzalez, N., The cell-cycle interactome: a source of growth regulators?, J. Exp. Bot., 2014, vol. 65, p. 2715. https://doi.org/10.1093/jxb/ert388
Van Leene, J., Hollunder, J., Eeckhout, D., Persiau, G., Van De Slijke, E., Stals, H., Van Isterdael, G., Verkest, A., Neirynck, S., Buffel, Y., De Bodt, S., Maere, S., Laukens, K., Pharazyn, A., Ferreira, P.C.G., et al., Targeted interactomics reveals a complex core cell cycle machinery in Arabidopsis thaliana, Mol. Syst. Biol., 2010, vol. 6, p. 397. https://doi.org/10.1038/msb.2010.53
Jia, R.-D., Guo, C.-C., Xu, G.-X., Shan, H.-Y., and Kong, H.-Z., Evolution of the cyclin gene family in plants, J. Syst. Evol., 2014, vol. 52, p. 651. https://doi.org/10.1111/jse.12112
Vandepoele, K., Raes, J., De Veylder, L., Rouzé, P., Rombauts, S., and Inzé, D., Genome-wide analysis of core cell cycle genes in Arabidopsis, Plant Cell, 2002, vol 14, p. 903. https://doi.org/10.1105/tpc.010445
Desvoyes, B., De Mendoza, A., Ruiz-Trillo, I., and Gutierrez, C., Novel roles of plant RETINOBLASTOMA-RELATED (RBR) protein in cell proliferation and asymmetric cell division, J. Exp. Bot., 2014, vol. 65, p. 2657. https://doi.org/10.1093/jxb/ert411
Desvoyes, B. and Gutierrez, C., Roles of plant retinoblastoma protein: cell cycle and beyond, EMBO J., 2020, vol. 39, p. e105802. https://doi.org/10.15252/embj.2020105802
Romeiro Motta, M., Zhao, X.A., Pastuglia, M., Belcram, K., Roodbarkelari, F., Komaki, M., Harashima, H., Komaki, S., Kumar, M., Bulankova, P., Heese, M., Riha, K., Bouchez, D., and Schnittger, A., B1-type cyclins control microtubule organization during cell division in Arabidopsis, EMBO Rep., 2022, vol. 23, p. e53995. https://doi.org/10.15252/embr.202153995
Menges, M., De Jager, S.M., Gruissem, W., and Murray, J.A.H., Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control, Plant J., 2005, vol. 41, p. 546. https://doi.org/10.1111/j.1365-313X.2004.02319.x
Ito, M., Expression of mitotic cyclins in higher plants: transcriptional and proteolytic regulation, Plant Biotechnol. Rep., 2014, vol. 8, p. 9. https://doi.org/10.1007/s11816-013-0297-9
Araki, S., Ito, M., Soyano, T., Nishihama, R., and Machida, Y., Mitotic cyclins stimulate the activity of c-Myb-like factors for transactivation of G2/M phase-specific genes in tobacco, J. Biol. Chem., 2004, vol. 279, p. 32979. https://doi.org/10.1074/jbc.M403171200
Umeda, M., Shimotohno, A., and Yamaguchi, M., Control of cell division and transcription by cyclin-dependent kinase-activating kinases in plants, Plant Cell Physiol., 2005, vol. 46, p. 1437. https://doi.org/10.1093/pcp/pci170
Pedroza-Garcia, J.A., Xiang, Y., and De Veylder, L., Cell cycle checkpoint control in response to DNA damage by environmental stresses, Plant J., 2022, vol. 109, p. 490. https://doi.org/10.1111/tpj.15567
De Veylder, L., Larkin, J.C., and Schnittger, A., Molecular control and function of endoreplication in development and physiology, Trends Plant Sci., 2011, vol. 16, p. 624. https://doi.org/10.1016/j.tplants.2011.07.001
De Veylder, L., Beeckman, T., Beemster, G.T., Krols, L., Terras, F., Landrieu, I., Van Der Schueren, E., Maes, S., Naudts, M., and Inzé, D., Functional analysis of cyclin-dependent kinase inhibitors of Arabidopsis, Plant Cell, 2001, vol. 13, p. 1653. https://doi.org/10.1105/tpc.010087
Churchman, M.L., Brown, M.L., Kato, N., Kirik, V., Hülskamp, M., Inzé, D., De Veylder, L., Walker, J.D., Zheng, Z., Oppenheimer, D.G., Gwin, T., Churchman, J., and Larkin, J.C., SIAMESE, a plant-specific cell cycle regulator, controls endoreplication onset in Arabidopsis thaliana, Plant Cell, 2006, vol. 18, p. 3145. https://doi.org/10.1105/tpc.106.044834
Peres, A., Churchman, M.L., Hariharan, S., Himanen, K., Verkest, A., Vandepoele, K., Magyar, Z., Hatzfeld, Y., Van Der Schueren, E., Beemster, G.T.S., Frankard, V., Larkin, J.C., Inzé, D., and De Veylder, L., Novel plant-specific cyclin-dependent kinase inhibitors induced by biotic and abiotic stresses, J. Biol. Chem., 2007, vol. 282, p. 25588. https://doi.org/10.1074/jbc.M703326200
Han, S.K., Herrmann, A., Yang, J., Iwasaki, R., Sakamoto, T., Desvoyes, B., Kimura, S., Gutierrez, C., Kim, E.-D., and Torii, K.U., Deceleration of the cell cycle underpins a switch from proliferative to terminal divisions in plant stomatal lineage, Dev. Cell, 2022, vol. 57, p. 569. https://doi.org/10.1016/j.devcel.2022.01.014
Shoaib, M., Nair, N., and Sørensen, C.S., Chromatin landscaping at mitotic exit orchestrates genome function, Front. Genet., 2020, vol. 11, p. 103. https://doi.org/10.3389/fgene.2020.00103
Buschmann, H. and Müller, S., Update on plant cytokinesis: rule and divide, Curr. Opin. Plant Biol., 2019, vol. 52, p. 97. https://doi.org/10.1016/j.pbi.2019.07.003
Jakoby, M. and Schnittger, A., Cell cycle and differentiation, Curr. Opin. Plant Biol., 2004, vol. 7, p. 661. https://doi.org/10.1016/j.pbi.2004.09.015
Maluszynska, J., Kolano, B., and Sas-Nowosielska, H., Endopolyploidy in plants, in Plant Genome Diversity, Greilhuber, J., Ed., Springer, 2013, vol. 2, p. 99. https://doi.org/10.1007/978-3-7091-1160-4_7
Breuer, C., Braidwood, L., and Sugimoto, K., Endocycling in the path of plant development, Curr. Opin. Plant Biol., 2014, vol. 17, p. 78. https://doi.org/10.1016/j.pbi.2013.11.007
Jiang, S., Wei, J., Li, N., Wang, Z., Zhang, Y., Xu, R., Zhou, L., Huang, X., Wang, L., Guo, S., Wang, Y., Song, C.-P., Qian, W., and Li, Y., The UBP14-CDKB1;1-CDKG2 cascade controls endoreduplication and cell growth in Arabidopsis, Plant Cell, 2022, vol. 34, p. 1308. https://doi.org/10.1093/plcell/koac002
Duan, S., Hu, L., Dong, B., Jin, H.L., and Wang, H.B., Signaling from plastid genome stability modulates endoreplication and cell cycle during plant development, Cell Rep., 2020, vol. 32, p. 108019. https://doi.org/10.1016/j.celrep.2020.108019
Mahapatra, K. and Roy, S., SOG1 transcription factor promotes the onset of endoreduplication under salinity stress in Arabidopsis, Sci. Rep., 2021, vol. 11, p. 11659. https://doi.org/10.1038/s41598-021-91293-1
Barow, M. and Meister, A., Endopolyploidy in seed plants is differently correlated to systematics, organ, life strategy and genome size, Plant Cell Environ., 2003, vol. 26, p. 571. https://doi.org/10.1046/j.1365-3040.2003.00988.x
Melaragno, J., Mehrotra, B., and Coleman, A., Relationship between endopolyploidy and cell size in epidermal tissue of Arabidopsis, Plant Cell, 1993, vol. 5, p. 1661. https://doi.org/10.1105/tpc.5.11.1661
Sugimoto-Shirasu, K. and Roberts, K., “Big it up”: endoreduplication and cell-size control in plants, Curr. Opin. Plant Biol., 2003, vol. 6, p. 544. https://doi.org/10.1016/j.pbi.2003.09.009
Granier, C., Cookson, S.J., Tardieu, F., and Muller, B., Cell cycle and environmental stresses, in: Cell cycle control and plant development. Annu. Plant Rev., v. 32, Inzé, D., Ed., Blackwell, 2007, p. 335. https://doi.org/10.1002/9781119312994.apr0346
Moreno, S., Canales, J., Hong, L., Robinson, D., Roeder, A.H., and Gutiérrez, R.A., Nitrate defines shoot size through compensatory roles for endoreplication and cell division in Arabidopsis thaliana, Curr. Biol., 2020, vol. 30, p. 1988. https://doi.org/10.1016/j.cub.2020.03.036
Tenorio Berrío, R., Nelissen, H., Inzé, D., and Dubois, M., Increasing yield on dry fields: molecular pathways with growing potential, Plant J., 2022, vol. 109, p. 323. https://doi.org/10.1111/tpj.15550
Kinsman, E.A., Lewis, C., Davies, M.S., Young, J.E., Francis, D., Vilhar, B., and Ougham, H.J., Elevated CO2 stimulates cells to divide in grass meristems: a differential effect in two natural populations of Dactylis glomerata, Plant Cell Environ., 1997, vol. 20, p. 1309. https://doi.org/10.1046/j.1365-3040.1997.d01-21.x
Taylor, G., Tricker, P.J., Zhang, F.Z., Alston, V.J., Miglietta, F., and Kuzminsky, E., Spatial and temporal effects of free-air CO2 enrichment (POPFACE) on leaf growth, cell expansion, and cell production in a closed canopy of poplar, Plant Physiol., 2003, vol. 131, p. 177. https://doi.org/10.1104/pp.011296
Maksymowych, R., Analysis of leaf development, in: Developmental and cell biology, Abercrombie, M., et al., Cambridge University Press, 1973, 109 p.
López-Juez, E., Dillon, E., Magyar, Z., Khan, S., Hazeldine, S., de Jager, S.M., Murray, J.A.H., Beemster, G.T.S., Bögre, L., and Shanahan, H., Distinct light-initiated gene expression and cell cycle programs in the shoot apex and cotyledons of Arabidopsis, Plant Cell, 2008, vol. 20, p. 947. https://doi.org/10.1105/tpc.107.057075
Bao, L., Inoue, N., Ishikawa, M., Gotoh, E., The, O.-K., Higa, T., Morimoto, T., Ginanjar, E.F., Harashima, H., Noda, N., Watahiki, M., Hiwatashi, Y., Sekine, M., Hasebe, M., Wada, M., and Fujita, T., A PSTAIRE-type cyclin-dependent kinase controls light responses in land plants, Sci. Adv., 2022, vol. 8, p. eabk2116. https://doi.org/10.1126/sciadv.abk2116
Fung-Uceda, J., Lee, K., Seo, P.J., Polyn, S., De Veylder, L., and Mas, P., The circadian clock sets the time of DNA replication licensing to regulate growth in Arabidopsis, Dev. Cell., 2018, vol. 45, p. 101. https://doi.org/10.1016/j.devcel.2018.02.022
Nishihama, R., and Kohchi, T., Evolutionary insights into photoregulation of the cell cycle in the green lineage, Curr. Opin. Plant Biol., 2013, vol. 16, p. 630. https://doi.org/10.1016/j.pbi.2013.07.006
Beel, B., Prager, K., Spexard, M., Sasso, S., Weiss, D., Müller, N., Heinnickel, M., Dewez, D., Ikoma, D., Grossman, A.R., Kottke, T., and Mittag, M., A flavin binding cryptochrome photoreceptor responds to both blue and red light in Chlamydomonas reinhardtii, Plant Cell, 2012, vol. 24, p. 2992. https://doi.org/10.1105/tpc.112.098947
Qiao, F., Petrasek, J., and Nick, P., Light can rescue auxin-dependent synchrony of cell division in a tobacco cell line, J. Exp. Bot., 2010, vol. 61, p. 503. https://doi.org/10.1093/jxb/erp319
Okello, R.C.O., de Visser, P.H.B., Heuvelink, E., Marcelis, L.F.M., and Struik, P.C., Light mediated regulation of cell division, endoreduplication and cell expansion, Environ. Exp. Bot., 2016, vol. 121, p. 39. https://doi.org/10.1016/j.envexpbot.2015.04.003
Granier, C. and Tardieu, F., Is thermal time adequate for expressing the effects of temperature on sunflower leaf development?, Plant Cell Environ., 1998, vol. 21, p. 695. https://doi.org/10.1046/j.1365-3040.1998.00319.x
Grif, V.G., Ivanov, V.B., and Machs, E.M., Cell cycle and its parameters in flowering plants, Tsitologiia, 2002, vol. 44, p. 936.
Thimann, K.V., Antagonisms and similarities between cytokinins, abscisic acid and auxin (mini review), in Physiology and Biochemistry of Cytokinins in Plants, Kaminek, M., Ed., SPB Academic Publishing, 1992, p. 395.
Van de Poel, B., Smet, D., and Van Der Straeten, D., Ethylene and hormonal cross talk in vegetative growth and development, Plant Physiol., 2015, vol. 169, p. 61. https://doi.org/10.1104/pp.15.00724
Zluhan-Martínez, E., López-Ruíz, B.A., García-Gómez, M.L., García-Ponce, B., de la Paz Sánchez, M., Álvarez-Buylla, E.R., and Garay-Arroyo, A., Integrative roles of phytohormones on cell proliferation, elongation and differentiation in the Arabidopsis thaliana primary root, Front. Plant Sci., 2021, vol. 12, p. 659155. https://doi.org/10.3389/fpls.2021.659155
Jiang, K., Guo, H., and Zhai, J., Interplay of phytohormones and epigenetic regulation: a recipe for plant development and plasticity, J. Integr. Plant Biol., 2022, vol. 65, p. 381. https://doi.org/10.1111/jipb.13384
Péret, B., De Rybel, B., Casimiro, I., Benková, E., Swarup, R., Laplaze, L., Beeckman, T., and Bennett, M.J., Arabidopsis lateral root development: an emerging story, Trends Plant Sci., 2009, vol. 14, p. 399. https://doi.org/10.1016/j.tplants.2009.05.002
Hemerly, A.S., Ferreira, P., de Almeida Engler, J., Van Montagu, M., Engler, G., and Inzé, D., cdc2a expression in Arabidopsis is linked with competence for cell division, Plant Cell., 1993, vol. 5, p. 1711. https://doi.org/10.1105/tpc.5.12.1711
Brumos, J., Robles, L.M., Yun, J., Vu, T.C., Jackson, S., Alonso, J.M., and Stepanova, A.N., Local auxin biosynthesis is a key regulator of plant development, Dev. Cell., 2018, vol. 47, p. 306. https://doi.org/10.1016/j.devcel.2018.09.022
Ivanov, V.B. and Filin, A.N., Cytokinins regulate root growth through its action on meristematic cell proliferation but not on the transition to differentiation, Funct. Plant Biol., 2017, vol. 45, p. 215. https://doi.org/10.1071/FP16340
Schaller, G.E., Street, I.H., and Kieber, J.J., Cytokinin and the cell cycle, Curr. Opin. Plant Biol., 2014, vol. 21, p. 7. https://doi.org/10.1016/j.pbi.2014.05.015
Pasternak, T., Miskolczi, P., Ayaydin, F., Mészáros, T., Dudits, D., and Fehér, A., Exogenous auxin and cytokinin dependent activation of CDKs and cell division in leaf protoplast-derived cells of alfalfa, Plant Growth Regul., 2000, vol. 32, p. 129. https://doi.org/10.1023/A:1010793226030
Richard, C., Lescot, M., Inzé, D., and De Veylder, L., Effect of auxin, cytokinin, and sucrose on cell cycle gene expression in Arabidopsis thaliana cell suspension cultures, Plant Cell. Tissue Organ Cult., 2002, vol. 69, p. 167. https://doi.org/10.1023/A:1015241709145
Mészáros, T., Miskolczi, P., Ayaydin, F., Pettkó-Szandtner, A., Peres, A., Magyar, Z., Horváth, G.V., Bakó, L., Fehér, A., and Dudits, D., Multiple cyclin-dependent kinase complexes and phosphatases control G2/M progression in alfalfa cells, Plant Mol. Biol., 2000, vol. 43, p. 595. https://doi.org/10.1023/a:1006412413671
Hartig, K. and Beck, E., Endogenous cytokinin oscillations control cell cycle progression of tobacco BY-2 cells, Plant Biol., 2005, vol. 7, p. 33. https://doi.org/10.1055/s-2004-830474
Riou-Khamlichi, C., Huntley, R., Jacqmard, A., and Murray, J.A., Cytokinin activation of Arabidopsis cell division through a D-type cyclin, Science, 1999, vol. 283, p. 1541. https://doi.org/10.1126/science.283.5407.1541
Menges, M., Samland, A.K., Planchais, S., and Murray, J.A.H., The D-Type Cyclin CYCD3;1 Is Limiting for the G1-to-S-Phase Transition in Arabidopsis, Plant Cell, 2006, vol. 18, p. 893. https://doi.org/10.1105/tpc.105.039636
Chen, C.C., Fu, S.F., Lee, Y.I., Lin, C.Y., Lin, W.C., and Huang, H.J., Transcriptome analysis of age-related gain of callus-forming capacity in Arabidopsis hypocotyls, Plant Cell Physiol., 2012, vol. 53, p. 1457. https://doi.org/10.1093/pcp/pcs090
Cho, H.-J., Kwon, H.-K., and Wang, M.-H., Expression of Kip-related protein 4 gene (KRP4) in response to auxin and cytokinin during growth of Arabidopsis thaliana, BMB Rep., 2010, vol. 43, p. 273. https://doi.org/10.5483/bmbrep.2010.43.4.273
Magyar, Z., De Veylder, L., Atanassova, A., Bakó, L., Inzé, D., and Bögre, L., The role of the Arabidopsis E2FB transcription factor in regulating auxin-dependent cell division, Plant Cell, 2005, vol. 17, p. 2527. https://doi.org/10.1105/tpc.105.033761
Jurado, S., Abraham, Z., Manzano, C., López-Torrejón, G., Pacios, L.F., and Del Pozo, J.C., The Arabidopsis cell cycle F-box protein SKP2A binds to auxin, Plant Cell, 2010, vol. 22, p. 3891. https://doi.org/10.1105/tpc.110.078972
Del Pozo, J.C. and Manzano, C., Auxin and the ubiquitin pathway. Two players-one target: the cell cycle in action, J. Exp. Bot., 2014, vol. 65, p. 2617. https://doi.org/10.1093/jxb/ert363
Sauer, M. and Kleine-Vehn, J., AUXIN BINDING PROTEIN1: the outsider, Plant Cell, 2011, vol. 23, p. 2033. https://doi.org/10.1105/tpc.111.087064
Huang, R., Zheng, R., He, J., Zhou, Z., Wang, J., Xiong, Y., and Xu, T., Noncanonical auxin signaling regulates cell division pattern during lateral root development, Proc. Natl. Acad. Sci. USA, 2019, vol. 116, p. 21285. https://doi.org/10.1073/pnas.1910916116
Takahashi, N., Kajihara, T., Okamura, C., Kim, Y., Katagiri, Y., Okushima, Y., Matsunaga, S., Hwang, I., and Umeda, M., Cytokinins control endocycle onset by promoting the expression of an APC/C activator in Arabidopsis roots, Curr. Biol., 2013, vol. 23, p. 1812. https://doi.org/10.1016/j.cub.2013.07.051
Yang, W., Cortijo, S., Korsbo, N., Roszak, P., Schiessl, K., Gurzadyan, A., Wightman, R., Jönsson, H., and Meyerowitz, E., Molecular mechanism of cytokinin-activated cell division in Arabidopsis, Science, 2021, vol. 371, p. 1350. https://doi.org/10.1126/science.abe2305
Park, J., Lee, S., Park, G., Cho, H., Choi, D., Umeda, M., Choi, Y., Hwang, D., and Hwang, I., CYTOKININ-RESPONSIVE GROWTH REGULATOR regulates cell expansion and cytokinin-mediated cell cycle progression, Plant Physiol., 2021, vol. 186, p. 1734. https://doi.org/10.1093/plphys/kiab180
Humplík, J.F., Bergougnoux, V., and Van Volkenburgh, E., To stimulate or inhibit? That is the question for the function of abscisic acid, Trends Plant Sci., 2017, vol. 22, p. 830. https://doi.org/10.1016/j.tplants.2017.07.009
Sun, L.R., Wang, Y.B., He, S.B., and Hao, F.S., Mechanisms for abscisic acid inhibition of primary root growth, Plant Signal. Behav., 2018, vol. 13, p. e1500069. https://doi.org/10.1080/15592324.2018.1500069
Tanaka, Y., Nose, T., Jikumaru, Y., and Kamiya, Y., ABA inhibits entry into stomatal-lineage development in Arabidopsis leaves, Plant J., 2013, vol. 74, p. 448. https://doi.org/10.1111/tpj.12136
Xie, Q., Essemine, J., Pang, X., Chen, H., and Cai, W., Exogenous application of abscisic acid to shoots promotes primary root cell division and elongation, Plant Sci., 2020, vol. 292, p. 110385. https://doi.org/10.1016/j.plantsci.2019.110385
Luo, X., Xu, J., Zheng, C., Yang, Y., Wang, L., Zhang, R., Ren, X., Wei, S., Aziz, U., Du, J., Liu, W., Tan, W., and Shu, K., Abscisic acid inhibits primary root growth by impairing ABI4-mediated cell cycle and auxin biosynthesis, Plant Physiol., 2022, vol. 191, p. 265. https://doi.org/10.1093/plphys/kiac407
Novikova, G.V., Stepanchenko, N.S., Zorina, A.A., Nosov, A.V., Rakitin, V.Y., Moshkov, I.E., and Los, D.A., Coupling of cell division and differentiation in Arabidopsis thaliana cultured cells with interaction of ethylene and ABA signaling pathways, Life, 2020, vol. 10, p. 15. https://doi.org/10.3390/life10020015
Shtin, M., Dello Ioio, R., and Del Bianco, M., It’s time for a change: the role of gibberellin in root meristem development, Front. Plant Sci., 2022, vol. 13, p. 882517. https://doi.org/10.3389/fpls.2022.882517
Claeys, H., De Bodt, S., and Inzé, D., Gibberellins and DELLAs: central nodes in growth regulatory networks, Trends Plant Sci., 2014, vol. 19, p. 231. https://doi.org/10.1016/j.tplants.2013.10.001
Oh, M.-H., Honey, S.H., and Tax, F.E., The control of cell expansion, cell division, and vascular development by brassinosteroids: a historical perspective, Int. J. Mol. Sci., 2020, vol. 21, p. 1743. https://doi.org/10.3390/ijms21051743
Hu, Y., Bao, F., and Li, J., Promotive effect of brassinosteroids on cell division involves a distinct CycD3-induction pathway in Arabidopsis, Plant J., 2000, vol. 24, p. 693. https://doi.org/10.1046/j.1365-313x.2000.00915.x
Zhiponova, M.K., Vanhoutte, I., Boudolf, V., Betti, C., Dhondt, S., Coppens, F., Mylle, E., Maes, S., González-García, M., Caño-Delgado, A.I., Inzé, D., Beemster, G.T.S., De Veylder, L., and Russinova, E., Brassinosteroid production and signaling differentially control cell division and expansion in the leaf, New Phytol., 2013, vol. 197, p. 490. https://doi.org/10.1111/nph.12036
Apelbaum, A. and Burg, S.P., Effect of ethylene on cell division and deoxyribonucleic acid synthesis in Pisum sativum, Plant Physiol., 1972, vol. 50, p. 117. https://doi.org/10.1104/pp.50.1.117
Herbert, R.J., Vilhar, B., Evett, C., Orchard, C.B., Rogers, H.J., Davies, M.S., and Francis, D., Ethylene induces cell death at particular phases of the cell cycle in the tobacco TBY-2 cell line, J. Exp. Bot., 2001, vol. 52, p. 1615. https://doi.org/10.1093/jxb/52.361.1615
Street, I.H., Aman, S., Zubo, Y., Ramzan, A., Wang, X., Shakeel, S.N., Kieber, J.J., and Schaller, G.E., Ethylene inhibits cell proliferation of the Arabidopsis root meristem, Plant Physiol., 2015, vol. 169, p. 338. https://doi.org/10.1104/pp.15.00415
Stoynova-Bakalova, E., Bakalov, D.V., and Baskin, T.I., Ethylene represses the promoting influence of cytokinin on cell division and expansion of cotyledons in etiolated Arabidopsis thaliana seedlings, PeerJ., 2022, vol. 10, p. e14315. https://doi.org/10.7717/peerj.14315
Dan, H., Imaseki, H., Wasteneys, G.O., and Kazama, H., Ethylene stimulates endoreduplication but inhibits cytokinesis in cucumber hypocotyl epidermis, Plant Physiol., 2003, vol. 133, p. 1726. https://doi.org/10.1104/pp.103.025783
Ortega-Martínez, O., Pernas, M., Carol, R.J., and Dolan, L., Ethylene modulates stem cell division in the Arabidopsis thaliana root, Sci., 2007, vol. 317, p. 507. https://doi.org/10.1126/science.1143409
Kazama, H., Dan, H., Imaseki, H., and Wasteneys, G.O., Transient exposure to ethylene stimulates cell division and alters the fate and polarity of hypocotyl epidermal cells, Plant Physiol., 2004, vol. 134, p. 1614. https://doi.org/10.1104/pp.103.031088
Love, J., Björklund, S., Vahala, J., Hertzberg, M., Kangasjärvi, J., and Sundberg, B., Ethylene is an endogenous stimulator of cell division in the cambial meristem of Populus, Proc. Natl. Acad. Sci. USA, 2009, vol. 106, p. 5984. https://doi.org/10.1073/pnas.0811660106
Bystrova, E.I., Zhukovskaya, N.V., Rakitin, V.J., and Ivanov, V.B., Role of ethylene in activation of cell division in quiescent center of excised maize roots, Russ. J. Dev. Biol., 2015, vol. 46, p. 60. https://doi.org/10.1134/S1062360415020034
Etchells, J.P., Provost, C.M., and Turner, S.R., Plant vascular cell division is maintained by an interaction between PXY and ethylene signaling, PLoS Genet., 2012, vol. 8, p. e1002997. https://doi.org/10.1371/journal.pgen.1002997
Fomenkov, A.A., Nosov, A.V., Rakitin, V.Y., Mamaeva, A.S., and Novikova, G.V., Cytophysiological characteristics of Arabidopsis thaliana cultivated cells with disable perception of ethylene signal by the ETR1 receptor, Russ. J. Plant Physiol., 2014, vol. 61, p. 598. https://doi.org/10.1134/S1021443714050070
Fomenkov, A.A., Nosov, A.V., Rakitin, V.Y., Sukhanova, E.S., Mamaeva, A.S., Sobol’kova, G.I., Nosov, A.M., and Novikova, G.V., Ethylene in the proliferation of cultured plant cells: regulating or just going along?, Russ. J. Plant Physiol., 2015, vol. 62, p. 815. https://doi.org/10.1134/S1021443715060059
Komaki, S and, Sugimoto, K., Control of the plant cell cycle by developmental and environmental cues, Plant Cell Physiol., 2012, vol. 53, p. 953. https://doi.org/10.1093/pcp/pcs070
Velappan, Y., Signorelli, S., and Considine, M.J., Cell cycle arrest in plants: what distinguishes quiescence, dormancy and differentiated G1?, Ann. Bot., 2017, vol. 120, p. 495. https://doi.org/10.1093/aob/mcx082
Gutierrez, C., The Arabidopsis cell division cycle, Arabidopsis Book, 2009, vol. 7, p. e0120. https://doi.org/10.1199/tab.0120
Author information
Authors and Affiliations
Corresponding authors
Additional information
Abbreviations: CC, cell cycle; MC, mitotic cycle; TF, transcription factor(s); CK, cytokinin(s); APC/C, Anaphase-Promoting Complex/Cyclosome; CDK, cyclin-dependent kinase(s); CYC, cyclin(s); DELLA domain, Asp-Glu-Leu-Leu-Ala; KRP, Kip-related proteins (Kip-like proteins–CDK inhibitors); RBR1, RETINOBLASTOMA-RELATED 1 (retinoblastoma-like protein 1); SIM, SIAMESE proteins (family of CDK inhibitors); SMR, SIM-related proteins.
Rights and permissions
About this article
Cite this article
Nosov, A.V., Fomenkov, A.A. Plant Cell Cycle: Molecular Events, Regulation by External Factors and Phytohormones. Russ J Plant Physiol 70, 91 (2023). https://doi.org/10.1134/S1021443722700054
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1134/S1021443722700054