Abstract
Non-Hodgkin lymphoma (NHL) is a heterogeneous group of cancers that differ in pathogenesis and prognosis. The main methods of treating NHL include chemotherapy, immunochemotherapy, and radiation therapy. However, a significant proportion of these tumors are chemoresistant or rapidly recur after a short chemotherapy-induced remission. In this regard, the search for alternative cytoreductive therapeutic methods is relevant. Aberrant expression of microRNA (miRNA) is one of the mechanisms responsible for the emergence and progression of malignant lymphoid neoplasms. We analyzed the profile of miRNA expression in the biopsy material from lymph nodes affected by diffuse large B-cell lymphoma (DLBCL). The key material of the study was histological preparations of lymph nodes obtained by excisional diagnostic biopsy and treated using conventional histomorphological formalin fixation methods. The study group consisted of patients with DLBCL (n = 52); the control group consisted of patients with reactive lymphadenopathy (RL) (n = 40). It was shown that the miR-150 expression level in DLBCL was reduced by more than 12 times (p = 3.6 × 10‒15) compared with RL. Bioinformatics analysis revealed the involvement of miR-150 in the regulation of hematopoiesis and lymphopoiesis. The data we obtained allow us to consider miR-150 as a promising therapeutic target with great potential in clinical practice.
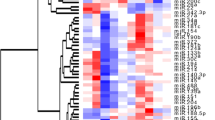
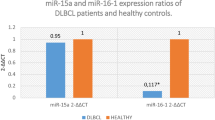
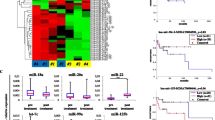
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
According to the classification of hematolymphoid tumors WHO-HAEM5 published by the World Health Organization in 2022, lymphomas are a heterogeneous group of cancers that differ in prognosis and pathogenesis. Non-Hodgkin lymphomas (NHLs) are divided into B- and T-cell lymphomas. The main subtypes of NHL include the following: diffuse large B-cell lymphoma (DLBCL), follicular lymphoma, Burkitt lymphoma, chronic lymphocytic leukemia, mucosa-associated lymphoid tissue tumors, mantle cell lymphoma, and marginal zone lymphoma. The classification of lymphomas is based on a combination of data on morphology, immunophenotype, specific genetic abnormalities, and clinical features of the disease. In patients with the diagnosed disease, the clinical course is extremely variable: from slow to aggressive. Thus, their treatment strategies and response to therapy also vary greatly, as do their clinical outcomes. Chemotherapy, immunochemotherapy, and radiation therapy are considered to be the main treatments for NHL, but a significant proportion of this type of tumors is chemoresistant or recurs after the treatment [1]. In this regard, the search for new therapeutic approaches remains relevant.
Advances in the study of molecular mechanisms underlying lymphomagenesis have led to the development of targeted drugs [2]. In recent years, microRNAs (miRNAs) have been considered as such drugs [3]. Aberrant miRNA expression has been described for most types of malignant tumors, including lymphomas [4]. In vitro experiments showed that the use of long interfering RNA for simultaneous inhibition of 13 onco-miRNAs significantly reduced cell proliferation, induced cell cycle arrest and apoptosis in DLBCL cell lines, mainly due to an increase in expression of PTEN, p27Kip1, TIMP3, RECK proteins and suppression of p38/MAPK, Survivin, CDK4, c-Myc [5]. H. Due et al. [6] reported that an increase in the miR-155 level in DLBCL cell lines induced changes in the sensitivity of these cells to vincristine. In addition, transfection of miR-197 and miR-187 mimetics into DLBCL cell lines increased their sensitivity to doxorubicin, enhancing apoptosis [7, 8]. Increased miR-10a and miR-26a expression inhibited proliferation and induced apoptosis of DLBCL cells [9, 10]. Increased miR-223-3p expression reduced cell proliferation and accelerated cell apoptosis in mantle cell lymphoma in vitro and in vivo through modulation of the CHUK/NF-κB2 signaling pathway [11]. miR-373 significantly slowed down growth of T-cell lymphoma cells [12]. M. Morales-Martinez et al. [13] noted that miR-7 regulates NHL chemosensitivity through negative regulation of YY1 and KLF4 genes [13]. It was shown that miR-150 can be considered as a potential therapeutic sensitizer that regulates the PI3K/AKT/mTOR pathway in the treatment of NK/T-cell lymphoma [14]. K. Musilova et al. [15] showed that transfection of miR-150 into follicular lymphoma cells led to a significant decrease in the proportion of cells in the S phase, which correlated with a decrease in tumor cell proliferation.
The possibility of delivering synthetic miRNA mimetics or inhibitors has opened new therapeutic perspectives. MRX34 (miR-34 mimetic), mesomiR-1 (miR-16 mimetic) and cobomarsen (anti-miR-155) showed antitumor activity in phase I clinical trials. These studies were not specifically designed for NHL, but included patients with B-cell NHLs [3, 16, 17].
These data confirm the therapeutic potential of miRNA. Personalized combined therapy can directly address the main problem in the treatment of tumors of various localization, drug resistance.
The aim of the work was to determine the miRNA expression profile in the material of DLBCL lymph nodes and to search for miRNAs that can be used as targeted drugs that allow maximum personalization of the therapy in the future.
EXPERIMENTAL
Studied groups. The experimental group included patients with DLBCL (n = 52). The control group consisted of patients with reactive lymphadenopathy (RL) (n = 40). Written informed consent was obtained from each patient, all data were anonymized. The study was approved by the Ethics Committee of the Novosibirsk State Medical University.
The study was carried out on histological preparations of tumor lymph node biopsies, which were treated using the classical methods of fixation in formalin, dehydration in isopropyl alcohol, defatting with xylene and impregnated with paraffin.
RNA extraction. To isolate nucleic acids from formalin-fixed paraffinized samples, they were deparaffinized. To do this, 1 mL of mineral oil was added to a tube containing three paraffin sections of lymph node tissue with a thickness of 15 μm, and thoroughly mixed for 10 s on a vortex (BioSan, Latvia), then the tube was transferred to a thermoshaker and incubated at 65°С and stirring frequency of 1300 rpm for 2 min. The resulting suspension was centrifuged at 13 000–15 000 g for 4 min, and the supernatant was removed; 1 mL of 96% ethanol was added to the precipitate, vortexed for 10 s, and centrifuged at 13 000–15 000 g for 4 min. The supernatant was removed and 1 mL of 70% ethanol was added to the precipitate, and the tube was centrifuged at the same speed for 2 min. The resulting precipitate was used to isolate nucleic acids.
Isolation of nucleic acids from the samples was carried out using the RealBest Extraction 100 reagent kit (Vector-Best, Russia).
miRNA selection. Based on data analysis by E. Sebestyén et al. [18], miRNAs for which number of copies in the sample exceeded 100 units that were present in at least 80% of the samples studied by the authors were selected for the study. Twenty-nine miRNAs met these criteria: miR-124-3p, -144-5p, -15а-5р, -16-5p, -196b-5p, -221-3p, -29b-3p, -148b-3p, -150-5p, ‑18a-5p, -183-5p, -185-5p, -205-5p, -20a-5p, -23a-3p, -23b-3p, -26b-5p, -30b-5p, -34a-5p, -451a, -9-5p, ‑128-3p, -141-3p, -200b-3p, -574-3p, -96-5p, let-7a-5p, let-7c-5p, and let-7f-5p. For normalization, the geometric mean of CT values of three miRNAs was used: miR-378-3p, -191-5p, and -103a-3p, which were also selected based on the literature data [19‒21]. All oligonucleotides were synthesized at Vector-Best. Depending on the system, the E value (reaction efficiency) varied from 92.5 to 99.7%.
Reverse transcription (RT). cDNA was synthesized in a 30 µL reaction mixture containing 3 µL of isolated RNA, 16.2 µL of 40% trehalose solution, 3 µL of 10× RT buffer (500 mM Tris-HCl, pH 8.3 (at 25°C), 500 mM KCl, 40 mM MgCl2), 3 µL of 4 mM solution of deoxynucleoside triphosphates, 3 µL of 10% BSA solution, 0.32 µL of reverse transcriptase (Vector-Best), 1.5 µL of 10 µM solution of the corresponding primer for RT. The mixture was incubated for 15 min at 16°C and 15 min at 42°C followed by inactivation for 2 min at 95°C.
Real time PCR. miRNA expression levels were assessed by real-time PCR on a CFX96 thermocycler (Bio-Rad Laboratories, United States). The reaction mixture with a volume of 30 µL contained 3 µL of cDNA sample, PCR buffer (Vector-Best), 0.5 µM of each primer, and 0.25 µM of a probe. Reaction conditions: 2 min at 50°C, 2 min at 94°C and 50 cycles of denaturation (10 s at 94°C), annealing and chain elongation (20 s at 60°C).
Statistical analysis. Statistical analysis was performed using the Statistica v13.1 program and the Mann–Whitney U test. P values < 0.05 were considered statistically significant.
RESULTS
Expression Analysis of Studied miRNAs in Clinical Samples
Expression levels of 29 miRNAs were analyzed by real-time RT-PCR in 52 DLBCL samples and 40 RL samples: miR-124, -144, -15a, -16, -196b, -221, -29b, -148b, -150, -18a, -183, -185, -205, -20a, -23a, -23b, -26b, -30b, -34a, -451a, -9, -128, -141, -200b, -574, ‑96, let-7a, let-7c, and let-7f. A statistically significant change in expression was revealed in DLBCL samples for 21 miRNAs compared with RL: a decrease for miR-26b, -30b, -150, -451a, -574, -144, -15a, -16, ‑196b, -221, ‑29b, -23a, -23b, -148, -128, let-7a, let-7c, let-7f, and an increase for miR-124, -9 and -96 (p < 0.05) (Table 1).
The most significant differences, by more than 2 times, in miRNA expression were registered for ten representatives: miR-26b, -150, -451a, -196b, -221, ‑29b, -23b, -128, let-7a, and let-7c. The distribution of the relative expression levels of these miRNAs, including the median value and interquartile range, is shown in Fig. 1.
The relative levels of differentially expressed miRNAs in DLBCL and RL samples. The horizontal black line within the box is the median, the box is the interquartile range, error bars are the range without outliers, outliers are indicated by circles. For all miRNAs presented, differences in expression were statistically significant (p < 0.05 according to the Mann–Whitney test).
Thus, miR-150 was found to have the largest range of expression changes among the studied miRNAs: its level was reduced by more than 12 times in DLBCL cells compared with nontumor tissue. Apparently, aberrant expression of this miRNA is associated with tumor development.
Bioinformatics Analysis of miRNA-150 Target Genes
The understanding of gene regulation mechanisms is one of the main tasks of molecular biology and bioinformatics. Experimentally confirmed pathways that are significantly enriched in miR-150 targets and disorders that can be associated with DLBCL (Table 2) were identified using the miRPathDB 2.0 resource (https://mpd.bioinf.uni-sb.de/).
DISCUSSION
Lymphocytes are among the most important cellular components of the immune system. Most lymphomas originate from B lymphocytes that have undergone neoplastic transformation. B-cell lymphoma subtypes are heterogeneous in terms of genetic and clinical characteristics [22]. The development of B cells is a complex process involving both transcription factors and cytokines, as well as miRNAs [23]. S. Koralov et al. [24] showed that the Dicer ribonuclease, a key participant in miRNA processing, is involved in the development of B lymphocytes, which suggests that a complex signaling cascade is involved in the regulation of this biological process.
We analyzed expression levels of 29 miRNAs in DLBCL and RL samples and found the most significant differences, more than 2 times, for ten miRNAs: miR-26b, -150, -451a, -196b, -221, -29b, -23b, -128, let-7a, and let-7c. According to the literature data, these miRNAs are involved in the regulation of normal hematopoiesis; therefore, their aberrant expression may be involved in the development of both myeloid and lymphoid tumors [25‒31]. It should be noted that miR-150 expression is reduced by more than 12 times (p = 3.6 × 10‒15) in DLBCL samples compared with tissues of nontumor nodules. Thus, miR-150 may be one of the key participants in the malignant transformation of B lymphocytes.
A number of works reflecting the role of miRNAs in B-cell differentiation and the development of B-cell lymphomas have been published [32]. miR-150 controls B-cell differentiation by acting on the c-Myb transcription factor [33]. c-Myb plays an important role during the development of B cells, in maintaining their proliferation, as well as in the cell cycle control of hematopoietic cells [34]. Disturbances in each of these biological processes contribute to the progression of tumors of various geneses, characterizing c-Myb as an important link in the development of both solid and hematological tumors. Studies on the mechanisms of carcinogenesis in Burkitt lymphoma revealed the key role of ZDHHC11 and ZDHHC11B genes in maintaining the MYC‒miR-150‒MYB pathway, which provide tumor development [35]. Burkitt lymphoma, like DLBCL, is an aggressive B-cell lymphoma. The MYC, MYB, and ZDHHC11 genes were also shown to be involved in DLBCL oncogenesis [36]. Thus, it is possible that similar regulatory pathways, in which miR-150 is involved, may be engaged in the development of DLBCL. M. Wang et al. [37] identified miR-150 as a tumor suppressor that reduces proliferation of Burkitt lymphoma cells, identifying c-Myb and Survivin as targets for this miRNA. In NK/T-cell lymphoma cells, increased expression of miR-150 correlates with increased apoptosis and reduced cell proliferation, which confirms the role of this miRNA as a suppressor; at the same time, DKC1 and AKT2 genes were identified as its direct targets [38]. In addition, miR-150 is involved in the development of T cells by regulating NOTCH3 expression, as well as the AKT3/Bim signaling pathway [39, 40]. A decrease in miR-150 expression promotes multiple organ invasion and metastasis of T-cell lymphoma due to increased expression of the target, CCR6 [41].
From a fundamental point of view, miR-150 can be considered a regulator of the development of lymphomas. In addition, the potential of this miRNA as a clinical and biological marker has already been shown. Thus, M. Mraz et al. [42] found that in patients with chronic lymphocytic leukemia, an increased level of miR-150 in the blood correlates with a longer overall survival. A H. Wang et al. [43] showed that reduced miR-150 expression correlates with shorter overall survival and progression-free periods in patients with primary gastrointestinal DLBCL.
As noted above, miRNAs are important regulators of normal hematopoiesis. In particular, miR-150 regulates terminal erythropoiesis in humans [44]. J. Lu et al. [45] noted that miR-150 modulates the development of megakaryocyte-erythroid progenitors, while the level of its expression is significantly reduced under conditions of an increased demand for erythropoiesis. Very often, in patients with DLBCL, the disease is accompanied by anemia, which allows us to consider miR-150 as one of the potential regulators of this physiological process as well. Thus, C. Apple et al. [46] found differential expression of miR-150, miR-223, miR-15a, and miR-24 in the bone marrow of patients with hip joint injury, noting the important role of these miRNAs in erythropoietic dysfunction associated with anemia.
Analysis of miRNA expression levels can be used as biomarkers not only in tissues, but also in blood plasma. H. Fayyad-Kazan et al. [47] believe that plasma levels of miR-150 and miR-342 can be considered as promising biomarkers in the diagnosis of acute myeloid leukemia.
Thus, it can be assumed that miR-150 mediates many pathophysiological processes by regulating expression of its target genes. Based on the data we obtained, miR-150 can be considered as a promising therapeutic target with great potential in clinical practice.
REFERENCES
Alaggio R., Amador C., Anagnostopoulos I., Attygalle A.D., Araujo I.B.O., Berti E., Bhagat G., Borges A.M., Boyer D., Calaminici M., Chadburn A., Chan J.K.C., Cheuk W., Chng W.J., Choi J.K., Chua-ng S.S., Coupland S.E., Czader M., Dave S.S., de Jong D., Du M.Q., Elenitoba-Johnson K.S., Ferry J., Geyer J., Gratzinger D., Guitart J., Gujral S., Harris M., Harrison C.J., Hartmann S., Hochhaus A., Jansen P.M., Karube K., Kempf W., Khoury J., Kimura H., Klapper W., Kovach A.E., Kumar S., Lazar A.J., Lazzi S., Leoncini L., Leung N., Leventaki V., Li X.Q., Lim M.S., Liu W.P., Louissaint A. Jr, Marcogliese A., Medeiros L.J., Michal M., Miranda R.N., Mitteldorf C., Montes-Moreno S., Morice W., Nardi V., Naresh K.N., Natkunam Y., Ng S.B., Oschlies I., Ott G., Parrens M., Pulitzer M., Rajkumar S.V., Rawstron A.C., Rech K., Rosenwald A., Said J., Sarkozy C., Sayed S., Saygin C., Schuh A., Sewell W., Siebert R., Sohani A.R., Tooze R., Traverse-Glehen A., Vega F., Vergier B., Wechalekar A.D., Wood B., Xerri L., Xiao W. 2022. The 5th edition of the World Health Organization classification of haematolymphoid tumours: lymphoid neoplasms. Leukemia. 36, 1720–1748.
Wang L., Qin W., Huo Y.J., Li X., Shi Q., Rasko J.E.J., Janin A., Zhao W.L. 2020. Advances in targeted therapy for malignant lymphoma. Signal Transduct. Target. Ther. 5 (1), 15.
Fuertes T., Ramiro A.R., de Yebenes V.G. 2020. miRNA-based therapies in B cell non-Hodgkin lymphoma. Trends Immunol. 41 (10), 932‒947.
Peng Y., Croce C.M. 2016. The role of microRNAs in human cancer. Signal Transduct. Target. Ther. 1, 15004.
Su Y., Sun B., Lin X., Zhao X., Ji W., He M., Qian H., Song X., Yang J., Wang J., Chen J. 2016. Therapeutic strategy with artificially-designed i-lncRNA targeting multiple oncogenic microRNAs exhibits effective antitumor activity in diffuse large B-cell lymphoma. Oncotarget. 7 (31), 49143‒49155.
Due H., Schönherz A.A., Ryø L., Primo M.N., Jespers-en D.S., Thomsen E.A., Roug A.S., Xiao M., Tan X., Pang Y., Young K.H., Bøgsted M., Mikkelsen J.G., Dybkær K. 2019. MicroRNA-155 controls vincristine sensitivity and predicts superior clinical outcome in diffuse large B-cell lymphoma. Blood Adv. 3 (7), 1185‒1196.
Yang J.M., Jang J.Y., Jeon Y.K., Paik J.H. 2018. Clinicopathologic implication of microRNA-197 in diffuse large B cell lymphoma. J. Transl. Med. 16 (1), 162.
Huang F., Jin Y., Wei Y. 2016. MicroRNA-187 induces diffuse large B-cell lymphoma cell apoptosis via targeting BCL6. Oncol. Lett. 11 (4), 2845‒2850.
Fan Q., Meng X., Liang H., Zhang H., Liu X., Li L., Li W., Sun W., Zhang H., Zen K., Zhang C.Y., Zhou Z., Chen X., Ba Y. 2016. miR-10a inhibits cell proliferation and promotes cell apoptosis by targeting BCL6 in diffuse large B-cell lymphoma. Protein Cell. 7 (12), 899‒912.
Farina F.M., Inguscio A., Kunderfranco P., Cortesi A., Elia L., Quintavalle M. 2017. MicroRNA-26a/cyclin-dependent kinase 5 axis controls proliferation, apoptosis and in vivo tumor growth of diffuse large B-cell lymphoma cell lines. Cell Death Dis. 8 (6), e2890.
Yuan J., Zhang Q., Wu S., Yan S., Zhao R., Sun Y., Tian X., Zhou K. 2021. MiRNA-223-3p affects mantle cell lymphoma development by regulating the CHUK/NF-ƘB2 signaling pathway. OncoTargets Ther. 14, 1553‒1564.
Tian Y.Y., Jia C.M., Li Y., Wang Y., Jiang L., Liu A.C. 2016. Restoration of microRNA-373 suppresses growth of human T-cell lymphoma cells by repressing CCND1. Eur. Rev. Med. Pharmacol. Sci. 20 (21), 4435‒4444.
Morales-Martinez M., Vega G.G., Neri N., Nambo M.J., Alvarado I., Cuadra I., Duran-Padilla M.A., Huerta-Yepez S., Vega M.I. 2020. MicroRNA-7 regulates migration and chemoresistance in non-Hodgkin lymphoma cells through regulation of KLF4 and YY1. Front. Oncol. 10, 588893.
Wu S.J., Chen J., Wu B., Wang Y.J., Guo K.Y. 2018. MicroRNA-150 enhances radiosensitivity by inhibiting the AKT pathway in NK/T cell lymphoma. J. Exp. Clin. Cancer Res. 37, 18.
Musilova K., Devan J., Cerna K., Seda V., Pavlasova G., Sharma S., Oppelt J., Pytlik R., Prochazka V., Prouzova Z., Trbusek M., Zlamalikova L., Liskova K., Kruzova L., Jarosova M., Mareckova A., Kornauth C., Simonitsch-Klupp I., Schiefer A.I., Merkel O., Mocikova H., Burda P., Machova Polakova K., Kren L., Mayer J., Zent C.S., Trneny M., Evans A.G., Janikova A., Mraz M. 2018. miR-150 downregulation contributes to the high-grade transformation of follicular lymphoma by upregulating FOXP1 levels. Blood. 132 (22), 2389‒2400.
Witten L., Slack F.J. 2020. miR-155 as a novel clinical target for hematological malignancies. Carcinogenesis. 41, 2–7.
Chakraborty C., Sharma A.R., Sharma G., Lee S.S. 2020. Therapeutic advances of miRNAs: a preclinical and clinical update. J. Adv. Res. 28, 127‒138.
Sebestyén E., Nagy Á., Marosvári D., Rajnai H., Kajtár B., Deák B., Matolcsy A., Brandner S., Storhoff J., Chen N., Bagó A.G., Bödör C., Reiniger L. 2022. Distinct miRNA expression signatures of primary and secondary central nervous system lymphomas. J. Mol. Diagn. 24 (3), 224‒240.
Costé É., Rouleux-Bonnin F. 2020. The crucial choice of reference genes: identification of miR-191-5p for normalization of miRNAs expression in bone marrow mesenchymal stromal cell and HS27a/HS5 cell lines. Sci. Rep. 10 (1), 17728.
de Leeuw D.C., van den Ancker W., Denkers F., de Menezes R.X., Westers T.M., Ossenkoppele G.J., van de Loosdrecht A.A., Smit L. 2013. MicroRNA profiling can classify acute leukemias of ambiguous lineage as either acute myeloid leukemia or acute lymphoid leukemia. Clin. Cancer Res. 19, 2187–2196.
Ragni E., Colombini A., De Luca P., Libonati F., Viganò M., Perucca Orfei C., Zagra L., de Girolamo L. 2021. miR-103a-3p and miR-22-5p are reliable reference genes in extracellular vesicles from cartilage, adipose tissue, and bone marrow cells. Front. Bioeng. Biotechnol. 9, 632440.
Esmeray E., Küçük C. 2020. Genetic alterations in B cell lymphoma subtypes as potential biomarkers for noninvasive diagnosis, prognosis, therapy, and disease monitoring. Turk. J. Biol. 44 (1), 1‒14.
Souza O.F., Popi A.F. 2022. Role of microRNAs in B‑cell compartment: development, proliferation and hematological diseases. Biomedicines. 10 (8), 2004.
Koralov S.B., Muljo S.A., Galler G.R., Krek A., Chakraborty T., Kanellopoulou C., Jensen K., Cobb B.S., Merkenschlager M., Rajewsky N., Rajewsky K. 2008. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell. 132 (5), 860‒874.
Veryaskina Yu.A., Titov S.E., Kovynev I.B., Fedorova S.S., Pospelova T.I., Zhimulev I.F. 2021. microRNAs in the myelodysplastic syndrome. Acta Nat. 13 (2), 4‒15.
Emmrich S., Rasche M., Schöning J., Reimer C., Keihani S., Maroz A., Xie Y., Li Z., Schambach A., Reinhardt D., Klusmann J.H. 2014. miR-99a/100~125b tricistrons regulate hematopoietic stem and progenitor cell homeostasis by shifting the balance between TGFβ and Wnt signaling. Genes Dev. 28 (8), 858‒874.
Velu C.S., Baktula A.M., Grimes H.L. 2009. Gfi1 regulates miR-21 and miR-196b to control myelopoiesis. Blood. 113 (19), 4720‒4728.
Felli N., Fontana L., Pelosi E., Botta R., Bonci D., Facchiano F., Liuzzi F., Lulli V., Morsilli O., Santoro S., Valtieri M., Calin G.A., Liu C.G., Sorrentino A., Croce C.M., Peschle C. 2005. MicroRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc. Natl. Acad. Sci. U. S. A. 102 (50), 18081‒18086.
Lee J.Y., Kim M., Heo H.R., Ha K.S., Han E.T., Park W.S., Yang S.R., Hong S.H. 2018. Inhibition of microRNA-221 and 222 enhances hematopoietic differentiation from human pluripotent stem cells via c-KIT upregulation. Mol. Cells. 41 (11), 971‒978.
Kurkewich J.L., Boucher A., Klopfenstein N., Baskar R., Kapur R., Dahl R. 2017. The Mirn23a and Mirn23b microRNA clusters are necessary for proper hematopoietic progenitor cell production and differentiation. Exp. Hematol. 59, 14‒29.
Pelosi A., Careccia S., Lulli V., Romania P., Marziali G., Testa U., Lavorgna S., Lo-Coco F., Petti M.C., Calabretta B., Levrero M., Piaggio G., Rizzo M.G. 2013. miRNA let-7c promotes granulocytic differentiation in acute myeloid leukemia. Oncogene. 32 (31), 3648‒3654.
Zheng B., Xi Z., Liu R., Yin W., Sui Z., Ren B., Miller H., Gong Q., Liu C. 2018. The function of microRNAs in B-cell development, lymphoma, and their potential in clinical practice. Front. Immunol. 9, 936.
Xiao C., Calado D.P., Galler G., Thai T.H., Patterson H.C., Wang J., Rajewsky N., Bender T.P., Rajewsky K. 2007. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell. 131 (1), 146‒159.
Nakata Y., Shetzline S., Sakashita C., Kalota A., Rallapalli R., Rudnick S.I., Zhang Y., Emerson S.G., Gewirtz A.M. 2007. c-Myb contributes to G2/M cell cycle transition in human hematopoietic cells by direct regulation of cyclin B1 expression. Mol. Cell Biol. 27 (6), 2048‒2058.
Dzikiewicz-Krawczyk A., Kok K., Slezak-Prochazka I., Robertus J.L., Bruining J., Tayari M.M., Rutgers B., de Jong D., Koerts J., Seitz A., Li J., Tillema B., Guikema J.E., Nolte I.M., Diepstra A., Visser L., Kluiver J., van den Berg A. 2017. ZDHHC11 and ZDHHC11B are critical novel components of the oncogenic MYC-miR-150-MYB network in Burkitt lymphoma. Leukemia. 31 (6), 1470‒1473.
Ziel-Swier L.J.Y.M., Liu Y., Seitz A., de Jong D., Koerts J., Rutgers B., Veenstra R., Razak F.R.A., Dzikiewicz-Krawczyk A., van den Berg A., Kluiver J. 2022. The role of the MYC/miR-150/MYB/ZDHHC11 network in Hodgkin lymphoma and diffuse large B-cell lymphoma. Genes (Basel). 13 (2), 227.
Wang M., Yang W., Li M., Li Y. 2014. Low expression of miR-150 in pediatric intestinal Burkitt lymphoma. Exp. Mol. Pathol. 96 (2), 261‒266.
Watanabe A., Tagawa H., Yamashita J., Teshima K., Nara M., Iwamoto K., Kume M., Kameoka Y., Takahashi N., Nakagawa T., Shimizu N., Sawada K. 2011. The role of microRNA-150 as a tumor suppressor in malignant lymphoma. Leukemia. 25 (8), 1324‒1334.
Ghisi M., Corradin A., Basso K., Frasson C., Serafin V., Mukherjee S., Mussolin L., Ruggero K., Bonanno L., Guffanti A., De Bellis G., Gerosa G., Stellin G., D’Agostino D.M., Basso G., Bronte V., Indraccolo S., Amadori A., Zanovello P. 2011. Modulation of microRNA expression in human T-cell development: targeting of NOTCH3 by miR-150. Blood. 117 (26), 7053‒7062.
Sang W., Sun C., Zhang C., Zhang D., Wang Y., Xu L., Zhang Z., Wei X., Pan B., Yan D., Zhu F., Yan Z., Cao J., Loughran T.P. Jr., Xu K. 2016. MicroRNA-150 negatively regulates the function of CD4(+) T cells through AKT3/Bim signaling pathway. Cell Immunol. 306–307, 35‒40.
Ito M., Teshima K., Ikeda S., Kitadate A., Watanabe A., Nara M., Yamashita J., Ohshima K., Sawada K., Tagawa H. 2014. MicroRNA-150 inhibits tumor invasion and metastasis by targeting the chemokine receptor CCR6, in advanced cutaneous T-cell lymphoma. Blood. 123 (10), 1499‒1511.
Mraz M., Chen L., Rassenti L.Z., Ghia E.M., Li H., Jepsen K., Smith E.N., Messer K., Frazer K.A., Kipps T.J. 2014. miR-150 influences B-cell receptor signaling in chronic lymphocytic leukemia by regulating expression of GAB1 and FOXP1. Blood. 124 (1), 84‒95.
Wang X., Kan Y., Chen L., Ge P., Ding T., Zhai Q., Yu Y., Wang X., Zhao Z., Yang H., Liu X., Li L., Qiu L., Qian Z., Zhang H., Wang Y., Zhao H. 2020. miR-150 is a negative independent prognostic biomarker for primary gastrointestinal diffuse large B-cell lymphoma. Oncol. Lett. 19 (5), 3487‒3494.
Sun Z., Wang Y., Han X., Zhao X., Peng Y., Li Y., Peng M., Song J., Wu K., Sun S., Zhou W., Qi B., Zhou C., Chen H., An X., Liu J. 2015. miR-150 inhibits terminal erythroid proliferation and differentiation. Oncotarget. 6 (40), 43033‒43047.
Lu J., Guo S., Ebert B.L., Zhang H., Peng X., Bosco J., Pretz J., Schlanger R., Wang J.Y., Mak R.H., Dombkowski D.M., Preffer F.I., Scadden D.T., Golub T.R. 2008. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev. Cell. 14 (6), 843‒853.
Apple C.G., Miller E.S., Kannan K.B., Stortz J.A., Loftus T.J., Lopez M.C., Parvataneni H.K., Patrick M., Hagen J.E., Baker H.V., Efron P.A., Mohr A.M. 2021. The role of bone marrow microRNA (miR) in erythropoietic dysfunction after severe trauma. Surgery. 169 (5), 1206‒1212.
Fayyad-Kazan H., Bitar N., Najar M., Lewalle P., Fayyad-Kazan M., Badran R., Hamade E., Daher A., Hussein N., ElDirani R., Berri F., Vanhamme L., Burny A., Martiat P., Rouas R., Badran B. 2013. Circulating miR-150 and miR-342 in plasma are novel potential biomarkers for acute myeloid leukemia. J. Transl. Med. 11, 31.
Funding
The work was supported by the Russian Foundation of Basic Research (project no. 19-34-60024) and the Russian Science Foundation (project no. 20-14-00074).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare that they have no conflicts of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals performed by any of the authors.
Statement of compliance with standards of research involving humans as subjects. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants involved in the study.
Additional information
Translated by D. Novikova
Abbreviations: NHL, non-Hodgkin lymphoma; DLBCL, diffuse large B-cell lymphoma; miRNA, microRNA; RL, reactive lymphadenopathy.
Rights and permissions
About this article
Cite this article
Veryaskina, Y.A., Titov, S.E., Kovynev, I.B. et al. MicroRNA Expression Profiling of Diffuse Large B-Cell Lymphoma. Mol Biol 57, 475–481 (2023). https://doi.org/10.1134/S0026893323030159
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893323030159