Abstract
The “Mendelian code” hypothesis postulates a relationship between Mendelian (monogenic) and common pathologies. In this hypothesis, polymorphisms in the genes of Mendelian diseases may have a significant contribution to predisposition to common diseases in which the same biochemical pathways may be involved. In this review a group of genes encoding various proteins participating in the DNA repair, with a particular focus on the BRCA1-associated genome surveillance complex (BASC), is presented through the prism of the “Mendelian code” hypothesis. Here we discuss (1) their main functions in the repair of DNA double-strand breaks (ATM, MRE11, NBN, RAD50, BRCA1, and BLM) and mismatch repair (MSH2, MSH6, MLH1, PMS2, RF-C, and PCNA); (2) the mitochondrial involvement of these proteins; (3) the involvement of BASC proteins in the development of an adaptive immune response. For 13 out of 16 BASC protein encoding genes, mutations leading to monogenic diseases have already been described; for 11, there are associations with common diseases or individual biological processes. Patients with mutations in the genes of the BASC complex and patients with severe combined immunodeficiency share similar symptoms. Polymorphisms within DNA repair genes may play a role in the development of common diseases through the involvement of the immune response. The pleiotropic effects of these genes suggest their participation in the development of various conditions, both in health and pathology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
“MENDELIAN CODE” OF COMMON DISEASES
Common diseases are a major public health problem around the world, therefore, the formation of new strategies for the prevention and treatment of widespread pathologies remains the focus of attention of researchers. The hereditary component of such diseases is complex, so genetic research in this area is still relevant today, although approaches to its study have changed significantly.
In recent years, the “Mendelian code,” according to which there is a connection between Mendelian and common diseases hypothesis has attracted considerable interest. Many of the common variants contributing to the formation of complexly inherited diseases are localized in the genes causal for monogenic pathologies [1]. The phenotypic effects of mutations leading to syndromic or monogenic pathology are extremely strong, while polymorphisms bring weak effects, modified by both environmental factors and genetic background. Therefore, polymorphisms in the genes of Mendelian diseases may be significant for common diseases in which the same biochemical pathways are involved. This idea has been confirmed and developed in a number of studies. For example, it has been shown that more than 23% of genes in which mutations lead to highly penetrant Mendelian diseases are also associated with common diseases, and the OR values for high risk variants of such genes are higher than for high risk variants of genes that do not lead to monogenic diseases, but are associated with common diseases [2]. It is known that carrying out genome-wide association studies (GWAS) makes it possible to identify substitutions in close proximity to genes of Mendelian diseases, and the phenotypic manifestations of complexly inherited pathologies partially overlap with manifestations of monogenic diseases [3].
A look through the prism of the “Mendelian code” hypothesis can give a second wind to the candidate approach in the study of the genetic component of common diseases. Our attention was drawn to a group of genes for proteins of DNA repair systems. Disturbance of DNA repair processes leads to a number of monogenic and oncological diseases [4, 5]. The involvement of gene polymorphism of DNA repair systems in the development of pathological conditions of various etiologies is currently being actively studied, but many questions remain. Most attention is focused on identifying the role of genes of DNA repair systems in the development of oncopathology. It is known that mutations in some genes of proteins of various DNA repair systems cause the development of a number of hereditary oncological diseases. Polymorphismsin the genes of this system are associated with oncopathology, sensitivity to chemicals, and radiation. The study of these phenomena most often includes the involvement of genetic polymorphism of these gene systems [5]. At the same time, the participation of polymorphism of genes of the repair systems in the formation of a predisposition to a wide range of complexly inherited pathologies, such as diseases of the cardiovascular system [6–9], mental [10, 11], metabolic [12], immunological [13–15], and other disorders has been studied (including by GWAS).
Hundreds of proteins are involved in DNA repair, according to GenOntology, the products of 511 genes are involved in DNA repair in humans (GO: 0006281, “Homo sapiens,” “experimental evidence,” and “UniProt” filters) [16]. In general, the proteins of the repair systems form multiprotein complexes that allow DNA damage to be corrected, and can also form complexes outside their own repair systems [17, 18]. In this review, the participation of polymorphisms of genes of the BRCA1-associated genome surveillance complex (BASC) in the formation of a predisposition to common diseases is considered. The BASC contains tumor suppressors and DNA damage repair proteins, namely, BRCA1, MSH2, MSH6, MLH1, PMS2, RF-C, PCNA, ATM, BLM, MRE11, NBN, and RAD50. The BASC is a dynamic system that changes its composition both throughout the cell cycle and within subcellular domains. The proteins of this complex (Fig. 1) participate as components in various repair systems and are also capable of forming small stable subcomplexes independent of BRCA1 [19].
BASC. The network was built using the on-line resource STRING v: 11.0; the lines connecting proteins in the network represent experimentally proven interactions, co-localization, and co-expression. In addition to the BASC, these proteins are included in protein complexes (all protein complexes are underlined) such as MRN (MRE11, RAD50, and NBN), MRX (MRE11 and RAD50), RF-C (RFC1, RFC2, RFC3, RFC4, and RFC5), and MMR (MLH1, PMS2, MSH2, MSH6, PCNA, and RF-C). Along with BRCA1, ATM, and BLM, the protein complexes MRN and MRX are involved in homologous recombination processes, and MRX and ATM are involved in non-homologous end joining processes. The RF-C heteropentamer and PCNA are involved in DNA replication and excisional repair. The MMR complex performs mismatch repair.
BASC PROTEINS AND DNA DOUBLE-STRAND BREAK REPAIR
Immediately after the DNA double-strand break, the nearby histone H2AX quickly becomes phosphorylated (by ATM kinase or other kinases of this family), forming gH2AX [20]. Phosphorylation of histone is necessary for coordinating the assembly of the repair complex, as well as for additional recruitment of the necessary enzymes [21–27]. Nibrin (NBN) in the MRN complex (including the MRE11, RAD50, and NBN proteins) binds to gH2AX and ATM, which promotes the accumulation of DNA repair factors in the chromatin surrounding damage, thereby activating the repair of DNA double-strand breaks [28–30]. The MRN complex is the only sensor for ATM activation in telomere dysfunction [31]. It plays a central role in the activation of ATM kinase at sites of DNA double-strand breaks [32]Fig. 1.
The ATM protein kinase, a member of the P13/P14 family , plays an important role in the signaling pathways for double-strand breaks in the DNA of higher eukaryotes. ATM indirectly controls the presence of DNA double-strand breaks through induced changes in the chromatin structure [33]. The ATM protein can be found both in the nucleus and in the cytoplasm, including in peroxisomes, vesicles, and mitochondria. The ATM kinase has hundreds of targets in a number of signaling pathways involved in the maintenance of cellular and redox homeostasis and regulation of mitochondrial functions [34–36]. It is assumed that in response to the appearance of DNA double-strand breaks, ATM is activated by two independent pathways, one involving TP53BP1, and the other NBN [37]. In addition, ATM can be activated in mitochondria in response to oxidative stress regardless of the cellular response to DNA damage, DNA single-strand breaks, and changes in chromatin structure [38–41]. Mutations in the ATM gene lead to ataxia-telangiectasia (Table 1), which, in addition to the main neurodegenerative disorders, is characterized by vascular changes and frequent respiratory infections. Associations of ATM gene variants with longevity [42, 43], schizophrenia [10], coronary atherosclerosis, diabetes mellitus [6, 44], insulin resistance [45], and response to metformin therapy in type 2 diabetes mellitus [46–48] have been described.
The MRN complex, consisting of the MRE11, NBN, and RAD50 proteins, is involved not only in the repair of DNA double-strand breaks, but is also associated with telomere formation and verification of DNA damage [49]. This complex is involved in signaling pathways that mediate the development of innate immunity [50]. Mutations in the NBN gene (NBS1) lead to Nijmegen chromosomal breakage syndrome, and mutations in the RAD50 gene lead to a disease similar to Nijmegen syndrome (Nijmegen breakage syndrome-like disorder) (Table 1). Moreover, polymorphisms of the NBN gene are associated with autoimmune diseases [13], metabolic disorders [12], and aging processes [51]. Variants of the RAD50 gene are associated with bronchial asthma [14, 52, 53] and cardiovascular pathology [54]. MRE11 gene mutations have been described in ATLD1 (Ataxia-telangiectasia-like disorder 1 (Table 1)) [56, 57], and the association of variants of this gene with myocardial infarction [7] and immune aging of T-cells [58] has also been described.
For effective signal transmission through MRN and ATM, a number of other factors are required, including TP53BP1 and BRCA1, the presence of which in the complex determines the pathway along which the repair of DNA double-strand breaks will go, either homologous recombination (HR), mediated by BRCA1, or non-homologous end joining (NHEJ) mediated by TP53BP1 [59, 60]. BRCA1 is a nuclear phosphoprotein that plays an important role in maintaining genomic stability in general, and as a tumor suppressor. The functions of BRCA1 are extremely diverse: it interacts with the components of the histone deacetylase complex, causing this enzyme to participate in the processes of transcription, DNA repair, and recombination [61], and participates in the assembly of the mitotic spindle [62]. By binding to RAD50, BRCA1 blocks the exonuclease activity of the MRN complex [63, 64]. Of all the genes of the BASC proteins, BRCA1 is the most actively studied - mutations in this gene are responsible for about 40% of hereditary breast cancer cases and more than 80% of cases of hereditary breast cancer with ovarian cancer (Table 1) [4, 5]. Polymorphisms of BRCA1 are associated with different types of inflammatory response [5, 65].
One of the components of the homologous recombination system, the product of the BLM gene, interacts with topoisomerase 3α and two helicases of the same subfamily as BLM (DExH-box-containing DNA and RNA helicases). BLM possesses DNA-dependent ATPase and DNA-helicase activity; it is involved in the processes of recombination, repair, replication, segregation of sister chromatids in mitosis, resolving the Holliday structure and untwisting G‑quadruplexes, recombination-mediated telomere elongation, and regulation of gene expression [66–69]. Mutations in this gene cause the development of Bloom syndrome (Table 1), which is characterized by an excessively high frequency of recombination of homologous chromosomes and sister chromatids [70, 71]. By influencing cell viability and apoptosis, BLM is involved in cataract progression [72]. In a mouse model, BLM has been shown to ensure the proper development and functioning of B-lymphocytes of various subtypes [73]. The involvement of BLM in the pathogenesis of colorectal and prostate cancer has been shown [74, 75].
It should be noted that there is an association of polymorphisms in the NBN, MRE11, RAD50, ATM, BRCA1, and BLM genes with various oncological diseases [5, 67, 76–85].
BASC PROTEINS AND MISMATCH REPAIR
Another complex associated with BASC is that of proteins of the very complex system of postreplicative DNA mismatch repair (MMR). The main role of this complex (which includes at least 20 proteins) is to eliminate errors associated with DNA replication (in addition to such mismatches as G/T, G/G, and A/C, the following are also targets for this system: O6-methylguanine, coupled with C or T, cisplatin-induced CpG interchain crosslinks, UV-induced photoproducts, purine adducts of benzpyrene, aminofluorene derivatives, 8‑oxyguanine, insertions, and deletions) [86]. In addition, MMR inhibits recombination between non-identical sequences and affects many processes associated with DNA metabolism, including signaling of DNA damage, expansion of trinucleotide repeats, switching of synthesis of immunoglobulin classes, and somatic hypermutability [87]. MMR appears to play a dual role in the response to DNA damage (direct mismatch repair and signaling of DNA damage) and is also involved in the activation of apoptosis induction [88]. In vivo experiments have shown that MMR mediates the cellular response to telomere dysfunction by weakening the induction of the p21 protein [89].
Mismatches are recognized and bound by MutS-subunits, which are heteroduplexes of three types: the MutSα complex, formed by the MSH2 and MSH6 proteins, recognizes mostly single mismatches and small insertions/deletions; the MutSβ complex (MSH2/MSH3), links extended insertions/deletions; and the MutSγ complex (MSH4/MSH5), is involved in the meiotic recombination process [90–93]. MutS binds the MutL-subunit, and is also represented by several heterodimeric forms: MutLα, MutLβ, and MutLγ. The MLH1 and PMS2 proteins form the MutLα subunit. Physical interactions of MutLα with DNA polymerase III, which bind to the repair site, are shown. MLH1 can also form heterodimers with MLH3, forming the MutLγ subunit involved in the meiotic process. PMS2 and MLH3 have weak endonuclease activity, which is critical for the functioning of the MutL subunit. PMS2 cannot cut methylated DNA, therefore it is likely that only de novo disorders can be corrected in this way, but this has not yet been proven in human cells [17, 18, 93, 94]. MLH1 in combination with PMS1 (MutLβ) or PMS2 suppresses homologous recombination, especially when it comes to short homologous DNA stretches. In addition, the human MutL complex protects against the formation of genomic rearrangements by participating in processes that are not directly related to mismatch repair [95]. Thus, PMS2 is involved in spermatogenesis [96]. Mutations in the MLH1, MLH3, PMS2, PMS1, MSH2, and MSH6 genes can be the cause of some hereditary cancers (Table 1), such as Lynch syndrome (hereditary non-polyposis colorectal cancer) and Turcot syndrome (mismatch repair syndromic cancer, brain tumor polyposis syndrome) [4, 97–101]. Mutations in MLH1 and MSH2 can cause Muir-Torre syndrome (MRTES) and in MLH1, MLH3, MSH2, MSH3, and MSH6—endometrial cancer [97, 102–105]. Associations of polymorphisms of these genes have also been shown with other oncological diseases [5, 106–111]. In addition, associations of polymorphisms of the MLH1 and PMS2 genes with lifespan have been revealed [112, 113]. Mutations in the MSH5 gene have been described, leading to premature extinction of ovarian function [114, 115], as well as the association of polymorphisms with IGAD1 (IgA deficiency) [116]. The MSH4 gene is associated with male infertility [117]. In a mouse model, it was shown that MSH2 deficiency leads to a genome-wide increase in the degree of histone H3 methylation [118].
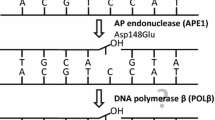
Another protein complex, replicative factor C (RF-C), which has ATPase activity in the presence of DNA and proliferating cell nuclear antigen (PCNA), plays an important role in mismatch repair. RF-C is a heteropentamer, the subunits of which are encoded by the RFC1, RFC2, RFC3, RFC4, and RFC5 genes [119]. RF-C interacts with the 5'-end of the DNA sequence and then binds PCNA, mediating its binding to DNA. PCNA is required for the assembly of the replication complex; it stabilizes the complex of template DNA and DNA polymerase (delta or epsilon), which provides processive DNA synthesis [120]. PCNA can also bind a number of proteins (ligase 1, methyltransferase, flap endonuclease 1, and others), attracting them, if necessary, to the replicative complex; it is involved in maintaining the viability of proliferating cells [121–123]. The endonuclease activity of MutLα is activated by PCNA [124]. RF-C is involved in the processes of replication, excisional nucleotide repair, mismatch repair, and maintenance of telomere stability [125–127]. Polymorphism of the PCNA, RFC1, RFC2, RFC3, RFC4, and RFC5 genes is actively studied in connection with oncopathologies [5, 119, 128–130]. Expansion of the pentanucleotide repeats in the RFC1 gene leads to autosomal recessive late age cerebellar ataxia (Table 1) [131]. An association of the polymorphisms in the RFC1 gene with immunological reactions was revealed [132]. The RFC2 gene is located in the deleted region in patients with Williams-Beuren syndrome (Table 1) [133, 134]. Mutations in the PCNA gene lead to the development of ATLD2 (Table 1), a disease similar to ataxia-telangiectasia [135]. The PCNA level is actively used as a protein marker of proliferating cells; therefore, a change in its level in various pathologies and in model systems has been noted in many works (see [5] and references therein). The involvement of PCNA in the pathogenesis of Parkinson’s disease has been shown [136, 137].
GENES OF BASC PROTEINS AND COMMON DISEASES
At present, there is no sufficiently convincing evidence of the participation of most genes of proteins of this functional class in the development of common diseases, although a lot of data have already been accumulated in this area. A. Ciccia and S.J. Elledge, investigating 40 syndromes caused by mutations in more than 80 genes, concluded that defects in DNA repair primarily affect the homeostasis of the nervous, immune, and reproductive systems; they can also lead to premature aging or a predisposition to cancer [34]. Indeed, from the data presented by these authors, it can be seen that pathology of the nervous system is observed in 62% of cases of syndromes caused by mutations in the genes of DNA repair proteins, an increased predisposition to oncopathology in 37%, disorders in the functioning of the immune system in 35%, and certain signs of premature aging in 25% of patients [34]. In the available sources, most information concerns the involvement of genes of DNA repair proteins in the development of oncopathologies (see, for example, DisGenet, and also [4, 5, 67, 76–85, 97–101, 106–111, 119, 128–130]). Therefore for this review, other aspects are more interesting to consider.
Mitochondrial Dysfunction
Chromosomal rearrangements and genome instability are markers of impaired DNA repair processes. However, a number of signs common to some syndromes associated with impaired DNA repair (ataxia-telangiectasia, Bloom syndrome, and Nijmegen syndrome), such as premature aging, growth retardation, insulin resistance, endocrine disorders, and immunodeficiency, can be caused by defects in antioxidant protection and, as a consequence, accumulation of oxidative DNA damage [138]. These processes are associated not only with the direct accumulation of mutations in the nuclei of somatic cells (correct DNA repair is especially important for neurons), but also with the accumulation of damage in mtDNA. Since mtDNA encodes important subunits of the mitochondrial respiratory chain, defects often lead to disruption of oxidative phosphorylation processes [34, 139]. Mitochondrial dysfunction can be the cause of a number of pathological conditions affecting primarily tissues with an active metabolism, including the central nervous system, skeletal muscles, heart, and number of other tissues, causing neurodegeneration, cardiovascular and metabolic diseases, aging, cancer, and other disorders [139, 140]. Disturbances in the ultrastructure and functions of mitochondria, and increased production of mitochondrial ROS are characteristic of ataxia-telangiectasia and ATLD, Bloom syndrome, Nijmegen syndrome, Cockayne syndrome, and pigmented xeroderma [36, 138].
Most ROS are generated during normal cellular metabolism, mainly in mitochondria, peroxisomes, and the endoplasmic reticulum [141]. The immediate physical proximity to the sources of ROS and the absence of histones leads to an increase in the rate of mutagenesis in mitochondria, which is 10–20 times higher than in the nuclear genome [140, 142]. The most stable oxidative damage to DNA, 8-oxyguanine, can form mismatches with adenine during subsequent replication, and these mismatches occur more often than “correct” pairs with cytosine [143]. Thus, a “mutator” phenotype is formed, i.e. DNA damage causing further mutations [144, 145]. Oxidative DNA damage is corrected primarily by BER (base excision repair); in mitochondria errors are also corrected primarily by BER. However, this is far from the only mechanism. It has now been proven that NER (nucleotide excision repair), MMR, and DNA double-strand break repair systems also actively function in mitochondria, both based on homologous recombination and non-homologous end joining, probably based on microhomology mediated end joining, MMEJ. Despite the fact that the details of the functioning of these systems in mitochondria and in the nucleus may differ [140, 145–147], the participation of proteins of the BASC in the repair processes in mitochondria is expected.
Several proteins of the BASC are found in mitochondria and are required for normal functioning. It has been shown in cellular and animal models that the loss of ATM kinase activity leads to a rapid change in mitochondrial homeostasis. In fact, this testifies to the important role of ATM in the functioning of mitochondria, independent of damage in both nuclear and mtDNA. It is assumed that an increase in the level of oxidative stress in mice with knockout of the ATM gene is associated with an increase in the level of ROS in mitochondria, possibly due to decreased activity of complex I [148]. When the MMR complex (MLH1 or MSH2) is deficient in the cell, the activity of complex I also decreases [124, 145], and the loss of MLH1 leads, in addition, to a significant decrease in the number of mtDNA copies. When assembling the MMR complex on DNA mismatches, MLH1 and ATM interact, which is necessary for the development of a further cellular response to DNA damage [149]. Mitochondrial dysfunction caused by a deficiency of proteins of the mismatch-repair system can either be mediated precisely by a violation of the MLH1/ATM interaction [145], or be an independent event with a still unknown implementation mechanism. The involvement of the MRN complex and the BLM gene product in the functioning of the MMEJ has been shown, however, evidence for the presence and functional significance in mitochondria has so far been obtained only for MRE11 and RAD50 [140, 150, 151]. The role of BRCA1 in maintaining the stability of both nuclear and mitochondrial genomes is considered universal [152]. The localization of other proteins of the BASC in mitochondria has not yet been established.
Thus, there are convincing data on the role of a number of genes of DNA repair systems in the development of mitochondrial dysfunction and response to oxidative stress. The participation of the ATM gene product in these processes has been studied most fully [153–155]. The involvement in the development of oxidative stress can explain the association of the ATM gene with the pathogenesis of a limited spectrum of diseases, in particular, cardiovascular or neurodegenerative diseases. In our opinion, a more universal mechanism by which proteins of genes of DNA repair systems can contribute to the development of common diseases of various etiologies is the modification of the immune response and inflammation.
Immune Response
The presence of concomitant immunological disorders in patients with monogenic diseases caused by mutations in the genes of DNA repair systems (along with increased radiosensitivity and a tendency to develop oncopathology) is well known. Thus, in patients with ataxia-telangiectasia (mutations in the ATM gene), an increased frequency of bacterial respiratory tract infections is observed from infancy due to a disturbance of the assembly of genes of immunoglobulins and T-cell receptors (TCR). Similar disorders were found in patients with Nijmegen (mutations in the NBN gene) and Bloom (mutations in the BLM gene) syndromes [156–158].
The formation of antigen-recognizing sites of immunoglobulins and TCRs during the maturation of T- and B-lymphocytes occurs as a result of V(D)J-recombination (rearrangement), which is somatic non-homologous recombination. Mutations in the genes of some proteins that carry out rearrangement lead to the development of severe combined immunodeficiency disorders (SCID) caused by disorders in the maturation and/or functioning of lymphocytes, such as, for example, Omenn syndrome. Disorder of rearrangement is not the only pathogenetic mechanism of SCID development; in addition, accumulation of toxic metabolites, impaired cytokine signaling, thymic abnormalities, and decreased survival of lymphopoietic progenitor cells have been described. However, it should be emphasized that increased radiosensitivity was observed only in patients with SCID, in whom the disease is caused by mutations in the genes of proteins involved in V(D)J-recombination [159–161]. Among the proteins of the BASC, ATM [162, 163], MRE11 [164], NBN [164–166], and RAD50 [164] are involved in such rearrangements.
At later stages of lymphocyte maturation, the formation of a further variety of antibodies is associated with two processes: somatic hypermutation (SHM) and immunoglobulin class switching (CSR) [167]. Violation of these processes leads to the development of rare primary immunodeficiencies, characterized by the absence of the production of the switched isotype (IgG, IgA or IgE) [168, 169]. The involvement of enzymes of the mismatch repair system in immunoglobulin class switching was described in a mouse model as early as 1999 [170]. Later, the involvement of the proteins of the MRN complex in this process was shown in humans [171]. It is known that CSR is carried out with the involvement of the MRN complex proteins, MRE11, RAD50, and NBN [171, 172], as well as MSH2, MLH1 [170], PMS2 [169, 170], ATM [173], and BRCA1 [174]. All proteins of the MMR system are also involved in somatic hypermutation processes [175, 176]. There are known mutations in the genes of the MMR system (MLH1 and MSH2) and the MRN complex (NBN and RAD50), leading to the development of IgAD and CVID antibody deficiency syndromes, as well as polymorphisms in the genes of various DNA repair systems associated with disorders of the process of immunoglobulin class switching [55].
As a result of the analysis of more than a million case histories, Blaire et al. compiled a list of more than 3000 highly correlated pairs of Mendelian disease/common diseases [1]. A query using the keyword “Immunodeficiency” yielded a list of two monogenic (Immunodeficiency with Increased IgM and SCID) and 50 common diseases. Moreover, in about a third of these diseases, inflammation is the leading or concomitant pathogenetic mechanism (for example, viral infection, rheumatoid arthritis, Crohn’s disease, psoriasis, bronchitis, acne, acute myocardial infarction, cataracts, osteoarthritis, and others).
“Mendelian Code”
A list of 3276 genes has been published, in which three groups are distinguished, genes for “complex and Mendelian diseases (CM),” “complex but not Mendelian (CNM)”, and “Mendelian but not complex (MNC)” [2]. Of the 16 genes of the BASC, eight are presented in this list, and only one of them, the ATM gene, belongs to the “CM” type, as it leads to ataxia-telangiectasia and is associated with rheumatoid arthritis. Two genes are categorized as “CNM” and are associated with asthma/atopic dermatitis (RAD50) and Alzheimer’s disease (RFC3). Five more genes are classified as MNC—MLH1, PMS2, BRCA1, BLM, and MSH2 [2]. Indeed, mutations leading to Mendelian pathology have been described in each of these five genes (Table 1). The questions remains whether these genes are truly not associated with complexly inherited diseases, or if such studies have simply not been carried out.
The development of monogenic diseases is associated with 13 out of 16 proteins of the BASC (Table 1). Mutations in the genes of seven of them lead either to the development of oncopathology (BRCA1, MLH1, PMS2, MSH2, and MSH6), or to an increased susceptibility to cancer (ATM and NBN). Mutations in eight genes are associated with neurological disorders (ATM, NBN, RAD50, MRE11, BRCA1 (with Fanconi anemia), BLM, RFC1, and PCNA), and nine, with immunological disorders as a part of the syndrome (immunodeficiency states—ATM, NBN, MRE11, and BLM; T-lymphomas in Turkot syndrome—MLH1, MSH2, MSH6, and PMS2) or as a result of other mutations in the same genes (NBN, RAD50, MLH1, and MSH2) [4]. Accordingly, it can be assumed that these genes are involved in the development of common diseases, which may be based on impaired immune processes, oxidative stress, and/or mitochondrial dysfunction.
Based on the hypothesis of the involvement of polymorphisms in the genes of proteins of DNA repair systems in the development of common diseases, we conducted a pilot study of coronary artery disease (CAD) and bronchial asthma (BA) and studied the variability of several genes of the BASC proteins (ATM, MLH1, PMS2, NBN, and MRE11) in these two pathologies. A number of associations with pathogenetically significant signs for the development of coronary artery disease (CAD) were revealed: body mass index and the presence of peripheral atherosclerosis (NBN and MLH1), levels of low-density lipoproteins (PMS2), dyslipidemia and myocardial mass index in patients with coronary artery disease (MRE11), and echocardiographic parameters (MLH1) [177–179]. In addition, associations of the ATM and MLH1 genes with BA have been revealed, it has been shown that the impact of environmental factors (smoking and parasitosis) significantly modifies the manifestation of associations [178, 180].
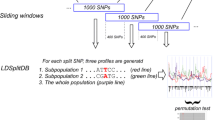
In addition, using the data presented in PheWeb, we analyzed the association of 16 genes of the BASC proteins with a wide range of phenotypes [181, 182]. PheWeb is a publicly available repository of phenome-wide association studies (PheWAS) of approximately 28 million SNPs with 1403 binary phenotypic traits. This analysis was carried out by the UK Biobank on 408 961 samples of Caucasian DNA using a specially adapted statistical approach [181, 182]. Table 2 shows associations with the highest levels of significance for each gene (a total of 71 pathological phenotypes of various etiologies; grouping of phenotypes and their names in the text are given according to PheWeb). Studies in the PheWAS format only began to be carried out in recent years due to a significant decrease in the cost of genotyping, but have not yet become widespread. Nevertheless, the PheWeb Internet resource has perhaps the most data.
The largest number of phenotypes associated with genes of the BASC proteins (Table 2) are related to disorders of the cardiovascular system, such as myocardial infarction, angina pectoris, various variants of coronary artery disease (associations of these phenotypes with the BLM gene are shown), coronary and cerebral atherosclerosis (BLM and MRE11), peripheral vascular disease (RFC4), hemorrhage (MSH6), and various types of hypertension (BLM, PCNA, and RAD50) (Table 2). Various sources also describe associations with both cardiovascular pathology in general and individual endophenotypes of the ATM, MRE11, and RAD50 genes [6–9, 54, 177–179]. There are also associations with a large group of phenotypes classified as dermatological disorders (Table 2) including alopecia (association with the NBN gene), diseases of the hair and hair follicles (MLH1), nails (MSH6), sebaceous glands (MLH1), disorders of the skin and subcutaneous tissue (BRCA1), and diffuse connective tissue diseases (RFC3 and MSH2). Most of these signs are found in progeria and/or physiological aging processes. It is noteworthy that when there are mutations in the ATM, BML, and NBN genes, premature aging is included in the complex of syndromic signs [138]. Associations of the NBN gene with aging processes [51] and associations of ATM, MLH1, and PMS2 with life expectancy [42, 43, 112, 113] are known. Other dermatological disorders (according to PheWeb, Table 2) include all variants of dermatitis, both of unspecified etiology and atopic/contact (RFC1), while asthma (RAD50) is classified as a respiratory system disease along with hemoptysis (PMS2), nasal polyps (RAD50), and other respiratory symptoms (PMS2) (Table 2). BA is associated with the RAD50 [14, 52, 53], ATM, and MLH1 genes [178, 180]. Both atopic dermatitis and atopic BA belong to allergic diseases (we mentioned them in the context of immunological pathologies). A number of associations of the genes under consideration with other immunological disorders are known including autoimmune diseases [13, 15], IgA deficiency [116], immune senescence of T-cells [58], etc. According to PheWeb, the BRCA1, MSH6, RFC1, and NBN genes are associated with some mental disorders (paranoid disorders, alcoholism, phobias, etc.) (Table 2), in particular the ATM gene is associated with schizophrenia [10]. The involvement of ATM kinase in the regulation of insulin response [35, 36] logically explains the associations of this gene with insulin resistance [6, 44–48], diabetes mellitus, and patient response to metformin treatment. According to PheWeb data (Table 2), there are associations of other genes of DNA repair proteins with metabolic disorders (hypothyroidism, MLH1), lipid metabolism disorders, and overweight (BLM and RFC2).
Thus, analysis of the results of associative studies of genes for DNA repair proteins (including information from PheWeb (Table 2)) allows us to make the following conclusions. A huge number of publications have been devoted to the role of genes of DNA repair proteins in the pathogenesis of neoplasia and carcinogenesis. In addition, associations of the genes discussed in this review with disorders of the cardiovascular system, digestive system, urogenital system, musculoskeletal system, hematopoietic system, sensory organs, endocrine disorders, metabolic disorders, mental disorders, neurological disorders, infectious diseases, respiratory diseases, dermatological diseases, traumatic injuries, and the body’s reactions to injuries and poisoning have been revealed. Accordingly, the considered genes have a wider scope than was classically suggested. All this indicates that genes of this functional class should be involved in the study of the genetic component of common diseases. The clearly pronounced pleiotropic effects of these genes explains their likelihood of participation in the development of various bodily states, both in health and disease, and, therefore, the association of markers of these genes with many diseases.
REFERENCES
Blair D.R., Lyttle C.S., Mortensen J.M., Bearden C.F., Jensen A.B., Khiabanian H., Melamed R., Rabadan R., Bernstam E.V., Brunak S., Jensen L.J., Nicolae D., Shah N.H., Grossman R.L., Cox N.J., et al. 2013. A nondegenerate code of deleterious variants in Mendelian loci contributes to complex disease risk. Cell. 155, 70–80.https://doi.org/10.1016/j.cell.2013.08.030
Spataro N., Rodrıguez J. A., Navarro A., Bosch E. 2017. Properties of human disease genes and the role of genes linked to Mendelian disorders in complex disease aetiology. Hum. Mol. Genet. 26, 489–500. https://doi.org/10.1093/hmg/ddw405
Freund M.K., Burch K.S., Shi H., Mancuso N., Kichaev G., Garske K.M., Pan D.Z., Miao Z., Mohlke K.L., Laakso M., Pajukanta P., Pasaniuc B., Arboleda V.A. 2018. Phenotype-specific enrichment of Mendelian disorder genes near GWAS regions across 62 complex traits. Am. J. Hum. Genet. 103, 535–552. https://doi.org/10.1016/j.ajhg.2018.08.017
OMIM–URL: https://omim.org/ Cited May 2020.
DisGeNet–URL: www.disgenet.org. Cited May 2020.
Li S., Zhang L., Chen T., Tian B., Deng X., Zhao Z., Yuan P., Dong B., Zhang Y., Mo X. 2011. Functional polymorphism rs189037 in the promoter region of ATM gene is associated with angiographically characterized coronary stenosis. Atherosclerosis. 219, 694–697. https://doi.org/10.1016/j.atherosclerosis.2011.08.040
Verschuren J.J., Trompet S., Deelen J., Stott D.J., Sattar N., Buckley B.M., Ford I., Heijmans B.T., Guchelaar H.J., Houwing-Duistermaat J.J., Slagboom P.E., Jukema J.W. 2013. Non-homologous end-joining pathway associated with occurrence of myocardial infarction: Gene set analysis of genome-wide association study data. PLoS One. 8, e56262. https://doi.org/10.1371/journal.pone.0056262
Wang L., Chu A., Buring J.E., Ridker P.M., Chasman D.I., Sesso H.D. 2014. Common genetic variations in the vitamin D pathway in relation to blood pressure. Am. J. Hypertens. 27, 1387–1395. https://doi.org/10.1093/ajh/hpu049
Guo Z., Kozlov S., Lavin M.F., Person M.D., Paull T.T. 2010. ATM activation by oxidative stress. Science. 330, 517–521. https://doi.org/10.1126/science.1192912
Zhang F., Xu Y., Liu P., Fan H., Huang X., Sun G., Song Y., Sham P.C. 2008. Association analyses of the interaction between the ADSS and ATM genes with schizophrenia in a Chinese population. BMC Med. Genet. 9, 119. https://doi.org/10.1186/1471-2350-9-119
Qian Y., Chen W., Wu J., Tao T., Bi L., Xu W., Qi H., Wang Y., Guo L. 2010. Association of polymorphism of DNA repair gene XRCC1 with sporadic late-onset Alzheimer’s disease and age of onset in elderly Han Chinese. J. Neurol. Sci. 295, 62–65. https://doi.org/10.1016/j.jns.2010.05.002
He C., Kraft P., Chasman D.I., Buring J.E., Chen C., Hankinson S.E., Pare G., Chanock S., Ridker P.M., Hunter D.J. 2010. A large-scale candidate-gene association study of age at menarche and age at natural menopause. Hum. Genet. 128, 515–527. https://doi.org/10.1007/s00439-010-0878-4
Lin Y.J., Lan Y.C., Wan L., Huang C.M., Lin C.W., Hsueh K.C., Chen D.Y., Lin T.H., Tsai F.J. 2010. The NBS1 genetic polymorphisms and the risk of the systemic lupus erythematosus in Taiwanese patients. J. Clin. Immunol. 30, 643–648. https://doi.org/10.1007/s10875-010-9427-0
Li X., Howard T.D., Zheng S.L., Haselkorn T., Peters S.P., Meyers D.A., Bleecker E.R. 2010 Genome-wide association study of asthma identifies RAD50-IL13 and HLA-DR/DQ regions. J. Allergy Clin. Immunol. 125, 328–335. https://doi.org/10.1016/j.jaci.2009.11.018
Souliotis V.L., Vougas K., Gorgoulis V.G., Sfikakis P.P. 2016. Defective DNA repair and chromatin organization in patients with quiescent systemic lupus erythematosus. Arthritis Res. Ther. 18, 182. https://doi.org/10.1186/s13075-016-1081-3
GenOntology. http://amigo.geneontology.org/amigo/ term/GO:0006281.
Kadyrov F.A., Dzantiev L., Constantin N., Modrich P. 2006. Endonucleolytic function of MutL alpha in human mismatch repair. Cell. 126, 297–308. https://doi.org/10.1016/j.cell.2006.05.039
Sacho E.J., Kadyrov F.A., Modrich P., Kunkel T.A., Erie D.A. 2008. Direct visualization of asymmetric adenine-nucleotide-induced conformational changes in MutL alpha. Mol. Cell. 29, 112–121. https://doi.org/10.1016/j.molcel.2007.10.030
Wang Y., Cortez D., Yazdi P., Neff N., Elledge S.J., Qin J. 2000. BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes Dev. 14, 927–939.
Daniel R., Ramcharan J., Rogakou E., Taganov K.D., Greger J.G., Bonner W., Nussenzweig A., Katz R.A., Skalka A.M. 2004. Histone H2AX is phosphorylated at sites of retroviral DNA integration but is dispensable for postintegration repair. J. Biol. Chem. 279, 45810–45814. https://doi.org/10.1074/jbc.M407886200
Paull T.T., Rogakou E.P., Yamazaki V., Kirchgessner C.U., Gellert M., Bonner W.M. 2000. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol. 10, 886–895. https://doi.org/10.1016/s0960-9822(00)00610-2
Redon C., Pilch D.R., Rogakou E.P., Orr A.H., Lowndes N.F., Bonner W.M. 2003. Yeast histone 2A serine 129 is essential for the efficient repair of checkpoint-blind DNA damage. EMBO J. 4, 678–684. https://doi.org/10.1038/sj.embor.embor871
Furuta T., Takemura H., Liao Z.Y., Aune G.J., Redon C., Sedelnikova O.A., Pilch D.R., Rogakou E.P., Celeste A., Chen H.T., Nussenzweig A., Aladjem M.I., Bonner W.M., Pommier Y. 2003. Phosphorylation of histone H2AX and activation of Mre11, Rad50, and Nbs1 in response to replication-dependent DNA double-strand breaks induced by mammalian DNA topoisomerase I cleavage complexes. J. Biol. Chem. 278, 20303–20312. https://doi.org/10.1074/jbc.M300198200
Lowndes N.F., Toh G.W. 2005. DNA repair: The importance of phosphorylating histone H2AX. Curr. Biol. 15, R99–R102. https://doi.org/10.1016/j.cub.2005.01.029
Morrison A.J., Shen X. 2005. DNA repair in the context of chromatin. Cell Cycle. 4, 568–571.
Ibuki Y., Toyooka T. 2015. Evaluation of chemical phototoxicity, focusing on phosphorylated histone H2AX. J. Radiat. Res. 56, 220–228. https://doi.org/10.1093/jrr/rru105
Georgoulis A., Vorgias C.E., Chrousos G.P., Rogakou E.P. 2017. Genome instability and γH2AX. Int. J. Mol. Sci. 18, E1979. https://doi.org/10.3390/ijms18091979
Kobayashi J., Tauchi H., Sakamoto S., Nakamura A., Morishima K., Matsuura S., Kobayashi T., Tamai K., Tanimoto K., Komatsu K. 2002. NBS1 localizes to gamma-H2AX foci through interaction with the FHA/BRCT domain. Curr. Biol. 12, 1846–1851. https://doi.org/10.1016/s0960-9822(02)01259-9
Roossink F., Wieringa H.W., Noordhuis M.G., ten Hoor K.A., Kok M., Slagter-Menkema L., Hollema H., de Bock G.H., Pras E., de Vries E.G., de Jong S., van der Zee A.G., Schuuring E., Wisman G.B., van Vugt M.A. 2012. The role of ATM and 53BP1 as predictive markers in cervical cancer. Int. J. Cancer. 131, 2056–2066. https://doi.org/10.1002/ijc.27488
Chen C., Zhang L., Huang N.J., Huang B., Kornbluth S. 2013. Suppression of DNA-damage checkpoint signaling by Rsk-mediated phosphorylation of Mre11. Proc. Natl. Acad. Sci. U. S. A. 110, 20605–20610. https://doi.org/10.1073/pnas.1306328110
Dimitrova N., de Lange T. 2009. Cell cycle-dependent role of MRN at dysfunctional telomeres: ATM signaling-dependent induction of nonhomologous end joining (NHEJ) in G1 and resection-mediated inhibition of NHEJ in G2. Mol. Cell. Biol. 29, 5552–5563. https://doi.org/10.1128/MCB.00476-09
Lavin M.F., Kozlov S., Gatei M., Kijas A.W. 2015. ATM-dependent phosphorylation of all three members of the MRN complex: From sensor to adaptor. Biomolecules. 5, 2877–2902. https://doi.org/10.3390/biom5042877
Zgheib O., Huyen Y., DiTullio R.A. Jr., Snyder A., Venere M., Stavridi E.S., Halazonetis T.D. 2005. ATM signaling and 53BP1. Radiother. Oncol. 76, 119–122. https://doi.org/10.1016/j.radonc.2005.06.026
Ciccia A., Elledge S.J. 2010. The DNA damage response: Making it safe to play with knives. Mol. Cell. 40, 179–204. https://doi.org/10.1016/j.molcel.2010.09.019
Shiloh Y., Ziv Y. 2013. The ATM protein kinase: Regulating the cellular response to genotoxic stress, and more. Nat. Rev. Mol. Cell. Biol. 14, 197–210. https://doi.org/10.1038/nrm3546
Choy K.R., Watters D.J. 2018. Neurodegeneration in ataxia-telangiectasia: Multiple roles of ATM kinase in cellular homeostasis. Dev. Dyn. 247, 33–46. https://doi.org/10.1002/dvdy.24522
Mochan T.A., Venere M., DiTullio R.A.Jr., Halazonetis T.D. 2003. 53BP1 and NFBD1/MDC1-Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage. Cancer Res. 63, 8586–8591.
Morita A., Tanimoto K., Murakami T., Morinaga T., Hosoi Y. 2014. Mitochondria are required for ATM activation by extranuclear oxidative stress in cultured human hepatoblastoma cell line Hep G2 cells. Biochem. Biophys. Res. Commun. 443, 1286–1290. https://doi.org/10.1016/j.bbrc.2013.12.139
Navrkalova V., Kafkova L.R., Divoky V., Pospisilova S. 2015. Oxidative stress as a therapeutic perspective for ATM-deficient chronic lymphocytic leukemia patients. Haematologica. 100, 994–996. https://doi.org/10.3324/haematol.2015.130260
Khoronenkova S.V. 2016. Mechanisms of noncanonocal ATM kinase activation. Usp. Biol. Khim. 56, 197–210. https://fbras.ru/wp-content/uploads/2017/10/ Khoronenkova-2016.pdf.
Berger N.D., Stanley F.K.T., Moore S., Goodarzi A.A. 2017. ATM-dependent pathways of chromatin remodelling and oxidative DNA damage responses. Philos. Trans. R. Soc. Lond. B. 372, 20160283. https://doi.org/10.1098/rstb.2016.0283
Chen T., Dong B., Lu Z., Tian B., Zhang J., Zhou J., Wu H., Zhang Y., Wu J., Lin P., Zhang J., Xu H., Mo X. 2010. A functional single nucleotide polymorphism in promoter of ATM is associated with longevity. Mech. Ageing Dev. 131, 636–640. https://doi.org/10.1016/j.mad.2010.08.009
Piaceri I., Bagnoli S., Tedde A., Sorbi S., Nacmias B. 2013. Ataxia-telangiectasia mutated (ATM) genetic variant in Italian centenarians. Neurol. Sci. 34, 573–575. https://doi.org/10.1007/s10072-012-1188-5
Schiekofer S., Bobak I., Kleber M.E., Maerz W., Rudofsky G., Dugi K.A., Schneider J.G. 2014. Association between a gene variant near ataxia telangiectasia mutated and coronary artery disease in men. Diab. Vasc. Dis. Res. 11, 60–63. https://doi.org/10.1177/1479164113514232
Joven J., Menendez J.A., Fernandez-Sender L., Espinel E., Rull A., Beltran-Debon R., Rodriguez-Gallego E., Riera-Borrull M., Pedro-Botet J., Alonso-Villaverde C., Camps J., Aragones G. 2013. Metformin: A cheap and well-tolerated drug that provides benefits for viral infections. HIV Med. 14, 233–240. https://doi.org/10.1111/hiv.12000
Florez J.C. 2011. Does metformin work for everyone? A genome-wide association study for metformin response. Curr. Diab. Rep. 11, 467–469. https://doi.org/10.1007/s11892-011-0220-0
GoDARTS and UKPDS Diabetes Pharmacogenetics Study Group; Wellcome Trust Case Control Consortium 2, Zhou K., Bellenguez C., Spencer C.C., Bennett A.J., Coleman R.L., Tavendale R., Hawley S.A., Donnelly L.A., Schofield C., Groves C.J., Burch L., Carr F., Strange A., et al. 2011. Common variants near ATM are associated with glycemic response to metformin in type 2 diabetes. Nat. Genet. 43, 117–120. https://doi.org/10.1038/ng.735
van Leeuwen N., Nijpels G., Becker M.L., Deshmukh H., Zhou K., Stricker B.H., Uitterlinden A.G., Hofman A., van’t Riet E., Palmer C.N., Guigas B., Slagboom P.E., Durrington P., Calle R.A., Neil A., et al. 2012. A gene variant near ATM is significantly associated with metformin treatment response in type 2 diabetes: A replication and meta-analysis of five cohorts. Diabetologia. 55, 1971–1977. https://doi.org/10.1007/s00125-012-2537-x
Haber J.E. 1998. The many interfaces of Mre11. Cell. 95, 583–586.
Bhattacharya S., Srinivasan K., Abdisalaam S., Su F., Raj P., Dozmorov I., Mishra R., Wakeland E.K., Ghose S., Mukherjee S., Asaithamby A. 2017. RAD51 interconnects between DNA replication, DNA repair and immunity. Nucleic Acids Res. 45, 4590–4605. https://doi.org/10.1093/nar/gkx126
Pereira-Lopes S., Tur J., Calatayud-Subias J.A., Lloberas J., Stracker T.H., Celada A. 2015. NBS1 is required for macrophage homeostasis and functional activity in mice. Blood. 126, 2502–2510. https://doi.org/10.1182/blood-2015-04-637371
Murk W., Walsh K., Hsu L.I., Zhao L., Bracken M.B., Dewan A.T. 2011. Attempted replication of 50 reported asthma risk genes identifies a SNP in RAD50 as associated with childhood atopic asthma. Hum. Hered. 71, 97–105. https://doi.org/10.1159/000319536
Chen J., Zhang J., Hu H., Jin Y., Xue M. 2015. Polymorphisms of RAD50, IL33 and IL1RL1 are associated with atopic asthma in Chinese population. Tissue Antigens. 86, 443–447. https://doi.org/10.1111/tan.12688
Mathur P., Kaga S., Zhan L., Das D.K., Maulik N. 2005. Antibody-array technique reveals overexpression of important DNA-repair proteins during cardiac ischemic preconditioning. J. Mol. Cell. Cardiol. 38, 99–102. https://doi.org/10.1016/j.yjmcc.2004.11.032
Offer S.M., Pan-Hammarstrom Q., Hammarstrom L., Harris R.S. 2010. Unique DNA repair gene variations and potential associations with the primary antibody deficiency syndromes IgAD and CVID. PLoS One. 5, e12260. https://doi.org/10.1371/journal.pone.0012260
Stewart G.S., Maser R.S., Stankovic T., Bressan D.A., Kaplan M.I., Jaspers N.G., Raams A., Byrd P.J., Petrini J.H., Taylor A.M. 1999. The DNA double-strand break repair gene hMRE11 is mutated in individuals with an ataxia-telangiectasia-like disorder. Cell. 99, 577–587.
Aquino J., Ribeiro V., Alonso I., Ramos F., Vasconcelos M. 2017. Ataxia-telangiectasia like: una adolescente portadora de una nueva variante del gen MRE11A (ataxia telangiectasia-like disorder—a child with a novel variant in MRE11A gene). Rev. Neurol. 65, 143–144.
Li Y., Shen Y., Hohensinner P., Ju J., Wen Z., Goodman S.B., Zhang H., Goronzy J.J., Weyand C.M. 2016. Deficient activity of the nuclease MRE11A induces T cell aging and promotes arthritogenic effector functions in patients with rheumatoid arthritis. Immunity. 45, 903–916. https://doi.org/10.1016/j.immuni.2016.09.013
Yun M.H., Hiom K. 2009. Understanding the functions of BRCA1 in the DNA-damage response. Biochem. Soc. Trans. 37, 597–604. https://doi.org/10.1042/BST0370597
Lee J.H., Goodarzi A.A., Jeggo P.A., Paull T.T. 2010. 53BP1 promotes ATM activity through direct interactions with the MRN complex. EMBO J. 29, 574–585. https://doi.org/10.1038/emboj.2009.372
Yarden R.I., Brody L.C. 1999. BRCA1 interacts with components of the histone deacetylase complex. Proc. Natl. Acad. Sci. U. S. A. 96, 4983–4988.
Joukov V., Groen A.C., Prokhorova T., Gerson R., White E., Rodriguez A., Walter J.C., Livingston D.M. 2006. The BRCA1/BARD1 heterodimer modulates ran-dependent mitotic spindle assembly. Cell. 127, 539–552.
Zhong Q., Chen C.F., Li S., Chen Y., Wang C.C., Xiao J., Chen P.L., Sharp Z.D., Lee W.H. 1999. Association of BRCA1 with the hRad50-hMre11-p95 complex and the DNA damage response. Science. 285, 747–750. https://doi.org/10.1126/science.285.5428.747
Paull T.T., Cortez D., Bowers B., Elledge S.J., Gellert M. 2001. Direct DNA binding by Brca1. Proc. Natl. Acad. Sci. U. S. A. 98, 6086–6091. https://doi.org/10.1073/pnas.111125998
Teoh H., Quan A., Creighton A.K., Bang A.K.W., Singh K.K., Shukla P.C., Gupta N., Pan Y., Lovren F., Leong-Poi H., Al-Omran M., Verma S. 2013. BRCA1 gene therapy reduces systemic inflammatory response and multiple organ failure and improves survival in experimental sepsis. Gene Ther. 20, 51–61. https://doi.org/10.1038/gt.2011.214
Johnson J.E., Cao K., Ryvkin P., Wang L.S., Johnson F.B. 2010. Altered gene expression in the Werner and Bloom syndromes is associated with sequences having G-quadruplex forming potential. Nucleic Acids Res. 38, 1114–1122. https://doi.org/10.1093/nar/gkp1103
Brosh R.M. Jr. 2013. DNA helicases involved in DNA repair and their roles in cancer. Nat. Rev. Cancer. 13, 542–558. https://doi.org/10.1038/nrc3560
Kitano K. 2014. Structural mechanisms of human RecQ helicases WRN and BLM. Front. Genet. 5, 366. https://doi.org/10.3389/fgene.2014.00366
Nguyen G.H., Tang W., Robles A.I., Beyer R.P., Gray L.T., Welsh J.A., Schetter A.J., Kumamoto K., Wang X.W., Hickson I.D., Maizels N., Monnat R.J. Jr, Harris C.C. 2014. Regulation of gene expression by the BLM helicase correlates with the presence of G-quadruplex DNA motifs. Proc. Natl. Acad. Sci. U. S. A. 111, 9905–9910. https://doi.org/10.1073/pnas.1404807111
Karow J.K., Constantinou A., Li J.L., West S.C., Hickson I.D. 2000. The Bloom’s syndrome gene product promotes branch migration of Holliday junctions. Proc. Natl. Acad. Sci. U. S. A. 97, 6504–6508. https://doi.org/10.1073/pnas.100448097
Lillard-Wetherell K., Machwe A., Langland G.T., Combs K.A., Behbehani G.K., Schonberg S.A., German J., Turchi J.J., Orren D.K., Groden J. 2004. Association and regulation of the BLM helicase by the telomere proteins TRF1 and TRF2. Hum. Mol. Genet. 13, 1919–1932. https://doi.org/10.1093/hmg/ddh193
Xiang J., Kang L., Gao H., Wu J., Qin B., Zhou T., Zhang G., Guan H. 2019. BLM can regulate cataract progression by influencing cell vitality and apoptosis. Exp. Eye Res. 178, 99–107. https://doi.org/10.1016/j.exer.2018.08.022
Babbe H., McMenamin J., Hobeika E., Wang J., Rodig S.J., Reth M., Leder P. 2009. Genomic instability resulting from Blm deficiency compromises development, maintenance, and function of the B cell lineage. J. Immunol. 182, 347–360. https://doi.org/10.4049/jimmunol.182.1.347
Qian X., Feng S., Xie D., Feng D., Jiang Y., Zhang X. 2017. RecQ helicase BLM regulates prostate cancer cell proliferation and apoptosis. Oncol. Lett. 14, 4206–4212. https://doi.org/10.3892/ol.2017.6704
Votino C., Laudanna C., Parcesepe P., Giordano G., Remo A., Manfrin E., Pancione M. 2017. Aberrant BLM cytoplasmic expression associates with DNA damage stress and hypersensitivity to DNA-damaging agents in colorectal cancer. J. Gastroenterol. 52, 327–340. https://doi.org/10.1007/s00535-016-1222-0
Rapakko K., Heikkinen K., Karppinen S.M., Erkko H., Winqvist R. 2007. Germline alterations in the 53BP1 gene in breast and ovarian cancer families. Cancer Lett. 245, 337–340. https://doi.org/10.1016/j.canlet.2006.01.021
Loizidou M.A., Michael T., Neuhausen S.L., Newbold R.F., Marcou Y., Kakouri E., Daniel M., Papadopoulos P., Malas S., Hadjisavvas A., Kyriacou K. 2009. DNA-repair genetic polymorphisms and risk of breast cancer in Cyprus. Breast Cancer Res. Treat. 115, 623–627. https://doi.org/10.1007/s10549-008-0084-4
Schuetz J.M., MaCarthur A.C., Leach S., Lai A.S., Gallagher R.P., Connors J.M., Gascoyne R.D., Spinelli J.J., Brooks-Wilson A.R. 2009. Genetic variation in the NBS1, MRE11, RAD50 and BLM genes and susceptibility to non-Hodgkin lymphoma. BMC Med. Genet. 10, 117. https://doi.org/10.1186/1471-2350-10-117
Pennington K.P., Walsh T., Harrell M.I., Lee M.K., Pennil C.C., Rendi M.H., Thornton A., Norquist B.M., Casadei S., Nord A.S., Agnew K.J., Pritchard C.C., Scroggins S., Garcia R.L., King M.C., Swisher E.M. 2014. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 20, 764–775. https://doi.org/10.1158/1078-0432.CCR-13-2287
Damiola F., Pertesi M., Oliver J., Le Calvez-Kelm F., Voegele C., Young E.L., Robinot N., Forey N., Durand G., Vallee M.P., Tao K., Roane T.C., Williams G.J., Hopper J.L., Southey M.C., et al. 2014. Rare key functional domain missense substitutions in MRE11A, RAD50, and NBN contribute to breast cancer susceptibility: Results from a Breast Cancer Family Registry case-control mutation-screening study. Breast Cancer Res. 16, R58. https://doi.org/10.1186/bcr3669
Janku F., Kaseb A.O., Tsimberidou A.M., Wolff R.A., Kurzrock R. 2014. Identification of novel therapeutic targets in the PI3K/AKT/mTOR pathway in hepatocellular carcinoma using targeted next generation sequencing. Oncotarget. 5, 3012–3022. https://doi.org/10.18632/oncotarget.1687
Ramus S.J., Song H., Dicks E., Tyrer J.P., Rosenthal A.N., Intermaggio M.P., Fraser L., Gentry-Maharaj A., Hayward J., Philpott S., Anderson C., Edlund C.K., Conti D., Harrington P., Barrowdale D., et al. 2015. Germline mutations in the BRIP1, BARD1, PALB2, and NBN genes in women with ovarian cancer. J. Natl. Cancer Inst. 107, pii: djv214. https://doi.org/10.1093/jnci/djv214
Smolarz B., Michalska M.M., Samulak D., Romanowicz H., Wojcik L. 2019. Polymorphism of DNA repair genes via homologous recombination (HR) in ovarian cancer. Pathol. Oncol. Res. 25, 1607–1614. https://doi.org/10.1007/s12253-019-00604-5
Kang Z., Zhu Y., Zhang Q.A., Dong L., Xu F., Zhang X., Guan M. 2019. Methylation and expression analysis of mismatch repair genes in extramammary Paget’s disease. J. Eur. Acad. Dermatol. Venereol. 33, 874–879. https://doi.org/10.1111/jdv.15404
Slavin T.P., Sun C.L., Chavarri-Guerra Y., Sedrak M.S., Katheria V., Castillo D., Herzog J., Dale W., Hurria A., Weitzel J.N. 2020. Older breast cancer survivors may harbor hereditary cancer predisposition pathogenic variants and are at risk for clonal hematopoiesis. J. Geriatr. Oncol. 11, 316–319. https://doi.org/10.1016/j.jgo.2019.09.004
Marra G., Schar P. 1999. Recognition of DNA alterations by the mismatch repair system. Biochem. J. 338, 1–13.
Jiricny J. 2006. The multifaceted mismatch-repair system. Nat. Rev. Mol. Cell. Biol. 7, 335–346. https://doi.org/10.1038/nrm1907
Luo Y., Lin F.T., Lin W.C. 2004. ATM-mediated stabilization of hMutL DNA mismatch repair proteins augments p53 activation during DNA damage. Mol. Cell. Biol. 24, 6430–6444. https://doi.org/10.1128/MCB.24.14.6430-6444.2004
Siegl-Cachedenier I., Munoz P., Flores J.M., Klatt P., Blasco M.A. 2007. Deficient mismatch repair improves organismal fitness and survival of mice with dysfunctional telomeres. Genes Dev. 21, 2234–2247. https://doi.org/10.1101/gad.430107
Paquis-Flucklinger V., Santucci-Darmanin S., Paul R., Saunieres A., Turc-Carel C., Desnuelle C. 1997. Cloning and expression analysis of a meiosis-specific MutS homolog: the human MSH4 gene. Genomics. 44, 188–194.
Bocker T., Barusevicius A., Snowden T., Rasio D., Guerrette S., Robbins D., Schmidt C., Burczak J., Croce C.M., Copeland T., Kovatich A.J., Fishel R. 1999. hMSH5: A human MutS homologue that forms a novel heterodimer with hMSH4 and is expressed during spermatogenesis. Cancer Res. 59, 816–822.
Snowden T., Shim K.S., Schmutte C., Acharya S., Fishel R. 2008. hMSH4-hMSH5 adenosine nucleotide processing and interactions with homologous recombination machinery. J. Biol. Chem. 283, 145–154. https://doi.org/10.1074/jbc.M704060200
Cannavo E., Gerrits B., Marra G., Schlapbach R., Jiricny J. 2007. Characterization of the interactome of the human MutL homologues MLH1, PMS1, and PMS2. J. Biol. Chem. 282, 2976–2986. https://doi.org/10.1074/jbc.M609989200
Raschle M., Marra G., Nystrom-Lahti M., Schar P., Jiricny J. 1999. Identification of hMutLbeta, a heterodimer of hMLH1 and hPMS1. J. Biol. Chem. 274, 32368–32375. https://doi.org/10.1074/jbc.274.45.32368
Siehler S.Y., Schrauder M., Gerischer U., Cantor S., Marra G., Wiesmüller L. 2009. Human MutL-complexes monitor homologous recombination independently of mismatch repair. DNA Repair (Amst.). 8, 242–252. https://doi.org/10.1016/j.dnarep.2008.10.011
van Oers J.M., Roa S., Werling U., Liu Y., Genschel J., Hou H. Jr., Sellers R.S., Modrich P., Scharff M.D., Edelmann W. 2010. PMS2 endonuclease activity has distinct biological functions and is essential for genome maintenance. Proc. Natl. Acad. Sci. U. S. A. 107, 13384–13389. https://doi.org/10.1073/pnas.1008589107
Hamilton S.R., Liu B., Parsons R.E., Papadopoulos N., Jen J., Powell S.M., Krush A.J., Berk T., Cohen Z., Tetu B, Burger P.C., Wood P.A., Taqi F., Booker S.V., Petersen G.M., et al. 1995. The molecular basis of Turcot’s syndrome. N. Engl. J. Med. 332, 839–847. https://doi.org/10.1056/NEJM199503303321302
Ou J., Rasmussen M., Westers H., Andersen S.D., Jager P.O., Kooi K.A., Niessen R.C., Eggen B.J., Nielsen F.C., Kleibeuker J.H., Sijmons R.H., Rasmussen L.J., Hofstra R.M. 2009. Biochemical characterization of MLH3 missense mutations does not reveal an apparent role of MLH3 in Lynch syndrome. Genes Chromosomes Cancer. 48, 340–350. https://doi.org/10.1002/gcc.20644
Senter L., Clendenning M., Sotamaa K., Hampel H., Green J., Potter J.D., Lindblom A., Lagerstedt K., Thibodeau S.N., Lindor N.M., Young J., Winship I., Dowty J.G., White D.M., Hopper J.L., et al. 2008. The clinical phenotype of Lynch syndrome due to germline PMS2 mutations. Gastroenterology. 135, 419–428. https://doi.org/10.1053/j.gastro.2008.04.026
Farrell M.P., Hughes D.J., Drost M., Wallace A.J., Cummins R.J., Fletcher T.A., Meany M.A., Kay E.W., de Wind N., Power D.G., Andrews E.J., Green A.J., Gallagher D.J. 2013. Multivariate analysis of MLH1 c.1664T>C (p.Leu555Pro) mismatch repair gene variant demonstrates its pathogenicity. Fam. Cancer. 12, 741–747. https://doi.org/10.1007/s10689-013-9652-9
Rossi L., Le Frere-Belda M.A., Laurent-Puig P., Buecher B., De Pauw A., Stoppa-Lyonnet D., Canlorbe G., Caron O., Borghese B., Colas C., Delhomelle H., Chabbert-Buffet N., Grandjouan S., Lecuru F., Bats A.S. 2017. Clinicopathologic characteristics of endometrial cancer in Lynch syndrome: A French multicenter study. Int. J. Gynecol. Cancer. 27, 953–960. https://doi.org/10.1097/IGC.0000000000000985
Risinger J.I., Umar A., Boyd J., Berchuck A., Kunkel T.A., Barrett J.C. 1996. Mutation of MSH3 in endometrial cancer and evidence for its functional role in heteroduplex repair. Nat. Genet. 14, 102–105.
Simpkins S.B., Bocker T., Swisher E.M., Mutch D.G., Gersell D.J., Kovatich A.J., Palazzo J.P., Fishel R., Goodfellow P.J. 1999. MLH1 promoter methylation and gene silencing is the primary cause of microsatellite instability in sporadic endometrial cancers. Hum. Mol. Genet. 8, 661–666. https://doi.org/10.1093/hmg/8.4.661
Stefansson I., Akslen L.A., MacDonald N., Ryan A., Das S., Jacobs I.J., Salvesen H.B. 2002. Loss of hMSH2 and hMSH6 expression is frequent in sporadic endometrial carcinomas with microsatellite instability: A population-based study. Clin. Cancer Res. 8, 138–143.
Taylor N.P., Powell M.A., Gibb R.K., Rader J.S., Huettner P.C., Thibodeau S.N., Mutch D.G., Goodfellow P.J. 2006. MLH3 mutation in endometrial cancer. Cancer Res. 66, 7502–7508. https://doi.org/10.1158/0008-5472.CAN-06-0248
Chen C.C., Yang S.Y., Liu C.J., Lin C.L., Liaw Y.F., Lin S.M., Lee S.D., Chen P.J., Chen C.J., Yu M.W. 2005. Association of cytokine and DNA repair gene polymorphisms with hepatitis B-related hepatocellular carcinoma. Int. J. Epidemiol. 34, 1310–1318. https://doi.org/10.1093/ije/dyi191
Rubio-Del-Campo A., Salinas-Sanchez A.S., Sanchez-Sanchez F., Gimenez-Bachs J.M., Donate-Moreno M.J., Pastor-Navarro H., Carrion-Lopez P., Escribano J. 2008. Implications of mismatch repair genes hMLH1 and hMSH2 in patients with sporadic renal cell carcinoma. BJU Int. 102, 504–509. https://doi.org/10.1111/j.1464-410X.2008.07581.x
Zhang Y., Shu Y.M., Wang S.F., Da B.H., Wang Z.H., Li H.B. 2010. Stabilization of mismatch repair gene PMS2 by glycogen synthase kinase 3beta is implicated in the treatment of cervical carcinoma. BMC Cancer. 10, 58. https://doi.org/10.1186/1471-2407-10-58
Vilkin A., Niv Y. 2011. Association between hMLH1 hypermethylation and JC virus (JCV) infection in human colorectal cancer (CRC). Clin. Epigenetics. 2, 1–5. https://doi.org/10.1007/s13148-010-0013-3
Shinsato Y., Furukawa T., Yunoue S., Yonezawa H., Minami K., Nishizawa Y., Ikeda R., Kawahara K., Yamamoto M., Hirano H., Tokimura H., Arita K. 2013. Reduction of MLH1 and PMS2 confers temozolomide resistance and is associated with recurrence of glioblastoma. Oncotarget. 4, 2261–2270. https://doi.org/10.18632/oncotarget.1302
Vageli D.P., Zaravinos A., Daniil Z., Dahabreh J., Doukas S.G., Spandidos D.A., Gourgoulianis K.I., Koukoulis G.K. 2013. hMSH2 and hMLH1 gene expression patterns differ between lung adenocarcinoma and squamous cell carcinoma: correlation with patient survival and response to adjuvant chemotherapy treatment. Int. J. Biol. Markers. 27, e400–4. https://doi.org/10.5301/JBM.2012.9420
Kim D.J., Yi S.M., Lee S.Y., Kang H.S., Choi Y.H., Song Y.W., Park S.C. 2006. Association between the MLH1 gene and longevity. Hum. Genet. 119, 353–354. https://doi.org/10.1007/s00439-006-0148-7
Han J., Ryu S., Moskowitz D.M., Rothenberg D., Leahy D.J., Atzmon G., Barzilai N., Suh Y. 2013. Discovery of novel non-synonymous SNP variants in 988 candidate genes from 6 centenarians by target capture and next-generation sequencing. Mech. Ageing Dev. 134, 478–485. doihttps://doi.org/10.1016/j.mad.2013.01.005
Mandon-Pepin B., Touraine P., Kuttenn F., Derbois C., Rouxel A., Matsuda F., Nicolas A., Cotinot C., Fellous M. 2008. Genetic investigation of four meiotic genes in women with premature ovarian failure. Eur. J. Endocrinol. 158, 107–115. https://doi.org/10.1530/EJE-07-0400
Guo T., Zhao S., Zhao S., Chen M., Li G., Jiao X., Wang Z., Zhao Y., Qin Y., Gao F., Chen Z.J. 2017. Mutations in MSH5 in primary ovarian insufficiency. Hum. Mol. Genet. 26, 1452–1457. https://doi.org/10.1093/hmg/ddx044
Sekine H., Ferreira R.C., Pan-Hammarstrom Q., Graham R.R., Ziemba B., de Vries S.S., Liu J., Hippen K., Koeuth T., Ortmann W., Iwahori A., Elliott M.K., Offer S., Skon C., Du L., et al. 2007. Role for Msh5 in the regulation of Ig class switch recombination. Proc. Natl. Acad. Sci. U. S. A. 104, 7193–7198. https://doi.org/10.1073/pnas.0700815104
Aston K.I., Krausz C., Laface I., Ruiz-Castane E., Carrell D.T. 2010. Evaluation of 172 candidate polymorphisms for association with oligozoospermia or azoospermia in a large cohort of men of European descent. Hum. Reprod. 25, 1383–1397. https://doi.org/10.1093/humrep/deq081
Herberg M., Siebert S., Quaas M., Thalheim T., Rother K., Hussong M., Altmuller J., Kerner C., Galle J., Schweiger M.R., Aust G. 2019. Loss of Msh2 and a single-radiation hit induce common, genome-wide, and persistent epigenetic changes in the intestine. Clin. Epigenetics. 11, 65. https://doi.org/10.1186/s13148-019-0639-8
Li Y., Gan S., Ren L., Yuan L., Liu J., Wang W., Wang X., Zhang Y., Jiang J., Zhang F., Qi X. 2018. Multifaceted regulation and functions of replication factor C family in human cancers. Am. J. Cancer Res. 8, 1343–1355. PMID: 30210909
Okumura K., Nogami M., Taguchi H., Dean F.B., Chen M., Pan Z.Q., Hurwitz J., Shiratori A., Murakami Y., Ozawa K., Eki T. 1995. Assignment of the 36.5‑kDa (RFC5), 37-kDa (RFC4), 38-kDa (RFC3), and 40-kDa (RFC2) subunit genes of human replication factor C to chromosome bands 12q24.2-q24.3, 3q27, 13q12.3-q13, and 7q11.23. Genomics. 25, 274–278. https://doi.org/10.1016/0888-7543(95)80135-9
Moldovan G.L., Pfander B., Jentsch S. 2007. PCNA, the maestro of the replication fork. Cell. 129, 665–679. https://doi.org/10.1016/j.cell.2007.05.003
Witko-Sarsat V., Mocek J., Bouayad D., Tamassia N., Ribeil J.A., Candalh C., Davezac N., Reuter N., Mouthon L., Hermine O., Pederzoli-Ribeil M., Cassatella M.A. 2010. Proliferating cell nuclear antigen acts as a cytoplasmic platform controlling human neutrophil survival. J. Exp. Med. 207, 2631–2645. https://doi.org/10.1084/jem.20092241
Cazzalini O., Sommatis S., Tillhon M., Dutto I., Bachi A., Rapp A., Nardo T., Scovassi A.I., Necchi D., Cardoso M.C., Stivala L.A., Prosperi E. 2014. CBP and p300 acetylate PCNA to link its degradation with nucleotide excision repair synthesis. Nucleic Acids Res. 42, 8433–8448. https://doi.org/10.1093/nar/gku533
Rashid S. 2017. Targeting the mitochondria for the treatment of MLH1-deficient disease. PhD Thesis, Queen Mary University of London. http://qmro. qmul.ac.uk/xmlui/handle/123456789/30924.
KEGG PATHWAY Database. https://www.genome. jp/kegg/pathway.html. Cited May 2020.
Uchiumi F., Ohta T., Tanuma S. 1996. Replication factor C recognizes 5'-phosphate ends of telomeres. Biochem. Biophys. Res. Commun. 229, 310–315. https://doi.org/10.1006/bbrc.1996.1798
Ohashi E., Tsurimoto T. 2017. Functions of multiple clamp and clamp-loader complexes in eukaryotic. In: DNA Replication. From Old Principles to New Discoveries. Eds. Masai H., Foiani M. Adv. Exp. Med. Biol. 135–162. https://doi.org/10.1007/978-981-10-6955-0_7
Liao Y.H., Ren J.T., Zhang W., Zhang Z.Z., Lin Y., Su F.X., Jia W.H., Tang L.Y., Ren Z.F. 2017. Polymorphisms in homologous recombination repair genes and the risk and survival of breast cancer. J. Gene Med. 19, e2988. https://doi.org/10.1002/jgm.2988
Hua Q., Gu X., Chen X., Song W., Wang A., Chu J. 2019. IL-8 is involved in radiation therapy resistance of esophageal squamous cell carcinoma via regulation of PCNA. Arch. Biochem. Biophys. 676, 108158. https://doi.org/10.1016/j.abb.2019.108158
Ye X., Ling B., Xu H., Li G., Zhao X., Xu J., Liu J., Liu L. 2020. Clinical significance of high expression of proliferating cell nuclear antigen in non-small cell lung cancer. Medicine (Baltimore). 99, e19755. https://doi.org/10.1097/MD.0000000000019755
Cortese A., Simone R., Sullivan R., Vandrovcova J., Tariq H., Yau W.Y., Humphrey J., Jaunmuktane Z., Sivakumar P., Polke J., Ilyas M., Tribollet E., Tomaselli P.J., Devigili G., Callegari I., et al. 2019. Biallelic expansion of an intronic repeat in RFC1 is a common cause of late-onset ataxia. Nat. Genet. 51, 649–658. https://doi.org/10.1038/s41588-019-0372-4
Arora M., Lindgren B., Basu S., Nagaraj S., Gross M., Weisdorf D., Thyagarajan B. 2010. Polymorphisms in the base excision repair pathway and graft-versus-host disease. Leukemia. 24, 1470–1475. https://doi.org/10.1038/leu.2010.139
Peoples R., Perez-Jurado L., Wang Y.K., Kaplan P., Francke U. 1996. The gene for replication factor C subunit 2 (RFC2) is within the 7q11.23 Williams syndrome deletion. Am. J. Hum. Genet. 58, 1370–1373. PMID: 8651315
Martindale D.W., Wilson M.D., Wang D., Burke R.D., Chen X., Duronio V., Koop B.F. 2000. Comparative genomic sequence analysis of the Williams syndrome region (LIMK1-RFC2) of human chromosome 7q11.23. Mamm. Genome. 11, 890–898. https://doi.org/10.1007/s003350010166
Baple E.L., Chambers H., Cross H.E., Fawcett H., Nakazawa Y., Chioza B.A., Harlalka G.V., Mansour S., Sreekantan-Nair A., Patton M.A., Muggenthaler M., Rich P., Wagner K., Coblentz R., Stein C.K., et al. 2014. Hypomorphic PCNA mutation underlies a human DNA repair disorder. J. Clin. Invest. 124, 3137–3146. https://doi.org/10.1172/JCI74593
Zhang Z., Zhang Z., Wang H., Zhang G., Hu D., Xiong J., Xiong N., Wang T., Cao X., Mao L. 2014. Proliferating cell nuclear antigen binds DNA polymerase-β and mediates 1-methyl-4-phenylpyridinium-induced neuronal death. PLoS One. 9, e106669. https://doi.org/10.1371/journal.pone.0106669
Li D.W., Li G.R., Zhang B.L., Feng J.J., Zhao H. 2016. Damage to dopaminergic neurons is mediated by proliferating cell nuclear antigen through the p53 pathway under conditions of oxidative stress in a cell model of Parkinson’s disease. Int. J. Mol. Med. 37, 429–435. https://doi.org/10.3892/ijmm.2015.2430
Maciejczyk M., Mikoluc B., Pietrucha B., Heropolitanska-Pliszka E., Pac M., Motkowski R., Car H. 2017. Oxidative stress, mitochondrial abnormalities and antioxidant defense in ataxia-telangiectasia, Bloom syndrome and Nijmegen breakage syndrome. Redox Biol. 11, 375–383. https://doi.org/10.1016/j.redox.2016.12.030
Greaves L.C., Reeve A.K., Taylor R.W., Turnbull D.M. 2012. Mitochondrial DNA and disease. J. Pathol. 226, 274–286. https://doi.org/10.1002/path.3028
Vasileiou P.V.S., Mourouzis I., Pantos C. 2017. Principal aspects regarding the maintenance of mammalian mitochondrial genome integrity. Int. J. Mol. Sci. 18, 1821. https://doi.org/10.3390/ijms18081821
Gorrini C., Harris I., Mak T. 2013. Modulation of oxidative stress as an anticancer strategy. Nat. Rev. Drug Discov. 12, 931–947. https://doi.org/10.1038/nrd4002
Song S, Pursell Z.F., Copeland W.C., Longley M.J., Kunkel T.A., Mathews C.K. 2005. DNA precursor asymmetries in mammalian tissue mitochondria and possible contribution to mutagenesis through reduced replication fidelity. Proc. Natl. Acad. Sci. U. S. A. 102, 4990–4995. https://doi.org/10.1073/pnas.0500253102
Kalam M.A., Haraguchi K., Chandani S., Loechler E.L., Moriya M., Greenberg M.M., Basu A.K. 2006. Genetic effects of oxidative DNA damages: comparative mutagenesis of the imidazole ring-opened formamidopyrimidines (Fapy lesions) and 8-oxo-purines in simian kidney cells. Nucleic Acids Res. 34, 2305–2315. https://doi.org/10.1093/nar/gkl099
Martin S.A., Hewish M., Sims D., Lord C.J., Ashworth A. 2011. Parallel high-throughput RNA interference screens identify PINK1 as a potential therapeutic target for the treatment of DNA mismatch repair-deficient cancers. Cancer Res. 71, 1836–1848. https://doi.org/10.1158/0008-5472.CAN-10-2836
Rashid S., Freitas M.O., Cucchi D., Bridge G., Yao Z., Gay L., Williams M., Wang J., Suraweera N., Silver A., McDonald S.A.C., Chelala C., Szabadkai G., Martin S.A. 201. MLH1 deficiency leads to deregulated mitochondrial metabolism. Cell Death Dis. 10, 795. https://doi.org/10.1038/s41419-019-2018-y
Storr S.J., Woolston C.M., Martin S.G. 2012. Base excision repair, the redox environment and therapeutic implications. Curr. Mol. Pharmacol. 5, 88–101. PMID: 22122466
Zinovkina L.A. 2018. Mechanisms of mitochondrial DNA repair in mammals. Biochemistry (Moscow). 83 (3), 233–249.
Valentin-Vega Y.A., Maclean K.H., Tait-Mulder J., Milasta S., Steeves M., Dorsey F.C., Cleveland J.L., Green D.R., Kastan M.B. 2012. Mitochondrial dysfunction in ataxia-telangiectasia. Blood. 119, 1490–1500. https://doi.org/10.1182/blood-2011-08-373639
Brown K.D., Rathi A., Kamath R., Beardsley D.I., Zhan Q., Mannino J.L., Baskaran R. 2003. The mismatch repair system is required for S-phase checkpoint activation. Nat. Genet. 33, 80–84. https://doi.org/10.1038/ng1052
Tadi S.K., Sebastian R., Dahal S., Babu R.K., Choudhary B., Raghavan S.C. 2016. Microhomology-mediated end joining is the principal mediator of double-strand break repair during mitochondrial DNA lesions. Mol. Biol. Cell. 27, 223–235. https://doi.org/10.1091/mbc.E15-05-0260
Seol J.H., Shim E.Y., Lee S.E. 2018. Microhomology-mediated end joining: Good, bad and ugly. Mutat. Res. 809, 81–87. https://doi.org/10.1016/j.mrfmmm.2017.07.002
Coene E.D., Hollinshead M.S., Waeytens A.A., Schelfhout V.R., Eechaute W.P., Shaw M.K., Van Oostveldt P.M., Vaux D.J. 2005. Phosphorylated BRCA1 is predominantly located in the nucleus and mitochondria. Mol. Biol. Cell. 16, 997–1010. https://doi.org/10.1091/mbc.e04-10-0895
Rotman G., Shiloh Y. 1997. Ataxia-telangiectasia: Is ATM a sensor of oxidative damage and stress? Bioessays. 19, 911–917.
Okuno Y., Nakamura-Ishizu A., Otsu K., Suda T., Kubota Y. 2012. Pathological neoangiogenesis dependson oxidative stress regulation by ATM. Nat. Med. 18, 1208–1216. https://doi.org/10.1038/nm.2846
Ambrose M., Gatti R.A. 2013. Pathogenesis of ataxia-telangiectasia: The next generation of ATM functions. Blood. 121, 4036–4045. https://doi.org/10.1182/blood-2012-09-456897
Bonilla F.A.1, Geha R.S. 2003. Primary immunodeficiency diseases. J. Allergy Clin. Immunol. 111, S571–581. https://doi.org/10.1067/mai.2003.86
Geha R.S., Notarangelo L.D., Casanova J.L., Chapel H., Conley M.E., Fischer A., Hammarstrom L., Nonoyama S., Ochs H.D., Puck J.M., Roifman C., Seger R., Wedgwood J.; International Union of Immunological Societies Primary Immunodeficiency Diseases Classification Committee. 2007. Primary immunodeficiency diseases: An update from the International Union of Immunological Societies Primary Immunodeficiency Diseases Classification Committee. J. Allergy Clin. Immunol. 120, 776–794. https://doi.org/10.1016/j.jaci.2007.08.053
Bonilla F.A., Barlan I., Chapel H., Costa-Carvalho B.T., Cunningham-Rundles C., de la Morena M.T., Espinosa-Rosales F.J., Hammarstrom L., Nonoyama S., Quinti I., Routes J.M., Tang M.L., Warnatz K. 2016. International Consensus Document (ICON): common variable immunodeficiency disorders. J. Allergy. Clin. Immunol. Pract. 4, 38–59. https://doi.org/10.1016/j.jaip.2015.07.025
Lim M.S., Elenitoba-Johnson K.S. 2004. The molecular pathology of primary immunodeficiencies. J. Mol. Diagn. 6, 59–83. https://doi.org/10.1016/S1525-1578(10)60493-X
Cirillo E., Giardino G., Gallo V., D’Assante R., Grasso F., Romano R., Di Lillo C., Galasso G., Pignata C. 2015. Severe combined immunodeficiency–an update. Ann. N.Y. Acad. Sci. 1356, 90–106. https://doi.org/10.1111/nyas.12849
Kumrah R., Vignesh P., Patra P., Singh A., Anjani G., Saini P., Sharma M., Kaur A., Rawat A. 2019. Genetics of severe combined immunodeficiency. Genes Dis. 24, 52–61. https://doi.org/10.1016/j.gendis.2019.07.004
Bredemeyer A.L., Sharma G.G., Huang C.Y., Helmink B.A., Walker L.M., Khor K.C., Nuskey B., Sullivan K.E., Pandita T.K., Bassing C.H., Sleckman B.P. 2006. ATM stabilizes DNA double-strand-break complexes during V(D)J recombination. Nature. 442, 466–470. https://doi.org/10.1038/nature04866
Zha S., Guo C., Boboila C., Oksenych V., Cheng H.-L., Zhang Y., Wesemann D.R., Yuen G., Patel H., Goff P.H., Dubois R.L., Alt F.W. 2011. ATM damage response and XLF repair factor are functionally redundant in joining DNA breaks. Nature. 469, 250–254. https://doi.org/10.1038/nature09604
Helmink B.A., Bredemeyer A.L., Lee B.-S., Huang C.-Y., Sharma G.G., Walker L.M., Bednarski J.J., Lee W.-L., Pandita T.K., Bassing C.H., Sleckman B.P. 2009. MRN complex function in the repair of chromosomal Rag-mediated DNA double-strand breaks. J. Exp. Med. 206, 669–679. https://doi.org/10.1084/jem.20081326
Chen H.T., Bhandoola A., Difilippantonio M.J., Zhu J., Brown M.J., Tai X., Rogakou E.P., Brotz T.M., Bonner W.M., Ried T., Nussenzweig A. 2000. Response to RAG-mediated VDJ cleavage by NBS1 and gamma-H2AX. Science. 290, 1962–1965. https://doi.org/10.1126/science.290.5498.1962
Saidi A., Li T., Weih F., Concannon P., Wang Z.Q. 2010. Dual functions of Nbs1 in the repair of DNA breaks and proliferation ensure proper V(D)J recombination and T-cell development. Mol. Cell. Biol. 30, 5572–5581. https://doi.org/10.1128/MCB.00917-10
Altshuler E.P., Serebryanaya D.V., Katrikha A.G. 2010. Production of monoclonal antibodies and methods to improve their affinity. Usp. Biol Khim. 50, 203–258.
Peron S., Pan-Hammarstrom Q., Imai K., Du L., Taubenheim N., Sanal O., Marodi L., Bergelin-Besançon A., Benkerrou M., de Villartay J.P., Fischer A., Revy P., Durandy A. 2007. A primary immunodeficiency characterized by defective immunoglobulin class switch recombination and impaired DNA repair. J. Exp. Med. 204, 1207–1216. https://doi.org/10.1084/jem.20070087
Peron S., Metin A., Gardes P., Alyanakian M.A., Sheridan E., Kratz C.P., Fischer A., Durandy A. 2008. Human PMS2 deficiency is associated with impaired immunoglobulin class switch recombination. J. Exp. Med. 205, 2465–2472. https://doi.org/10.1084/jem.20080789
Schrader C.E., Edelmann W., Kucherlapati R., Stavnezer J. 1999. Reduced isotype switching in splenic B cells from mice deficient in mismatch repair enzymes. J. Exp. Med. 190, 323–330. https://doi.org/10.1084/jem.190.3.323
Lahdesmaki A., Taylor A.M., Chrzanowska K.H., Pan-Hammarstrom Q. 2004. Delineation of the role of the Mre11 complex in class switch recombination. J. Biol. Chem. 279, 16479–16487. https://doi.org/10.1074/jbc.M312796200
Dinkelmann M., Spehalski E., Stoneham T., Buis J., Wu Y., Sekiguchi J.M., Ferguson D.O. 2009. Multiple functions of MRN in end-joining pathways during isotype class switching. Nat. Struct. Mol. Biol. 16, 808–813. https://doi.org/10.1038/nsmb.1639
Pan-Hammarstrom Q., Dai S., Zhao Y., van Dijk-Hard I.F., Gatti R.A., Borresen-Dale A.L., Hammarstrom L. 2003. ATM is not required in somatic hypermutation of VH, but is involved in the introduction of mutations in the switch mu region. J. Immunol. 170, 3707–3716. https://doi.org/10.4049/jimmunol.170.7.3707
Yun M.H., Hiom K. 2009. Understanding the functions of BRCA1 in the DNA-damage response. Biochem. Soc. Trans. 37, 597–604. https://doi.org/10.1042/BST0370597
Chahwan R., Edelmann W., Scharff M.D., Roa S. 2011. Mismatch-mediated error prone repair at the immunoglobulin genes. Biomed. Pharmacother. 65, 529–536. https://doi.org/10.1016/j.biopha.2011.09.001
Chahwan R., van Oers J.M., Avdievich E., Zhao C., Edelmann W., Scharff M.D., Roa S. 2012. The ATPase activity of MLH1 is required to orchestrate DNA double-strand breaks and end processing during class switch recombination. J. Exp. Med. 209, 671–678. https://doi.org/10.1084/jem.20111531
Babushkina N.P., Postrigan A.E., Kucher A.N. 2018. Involvement of DNA repair genes in the development of cardiovascular pathology. In Molekulyarno-biologi-cheskie tekhnologii v meditsinskoi praktike (Molecular Biological Technologies in Medical Practice), Maslennikov A.B., Ed., Novosibirsk: Akademizdat, pp. 48–62.
Babushkina N., Postrigan A., Kucher A. 2018. Association rs189037 in the ATM Gene with Bronchial Asthma Taking into Account Environmental Factors. BGRS\SB-2018: Systems Biology and Biomedicine (SBioMed-2018. Symp.), Abstracts Book. Novosibirsk, Russia, p. 16.
Babushkina N.P., Postrigan A.E., Kucher A.N. 2018. Involvemenr of MLH1 gene in the development of IHD clinical features. In: Novye tekhnologii – v praktiku zdravookhraneniya. Rossiiskii natsional’nyi kongress kardiologov, sbornik tezisov (Novel Technologies in Punic Healthcare Practice: Abstr. Russian Natural Congress of Cardiologists), Moscow, p. 1003.
Babushkina N.P., Postrigan A.E., Khitrinskaya E.Yu., Kucher A.N. 2019. Environmental effects of the association of genes encoding DNA repair proteins with bronchial asthma. In: VII S”ezd Vavilovskogo obshchestva genetikov i selektsionerov (VOGiS). Sbornik tezisov (Abstr. VII Congress of Vavilov Society of Geneticists and Breeders), St. Petersburg, p. 788
Zhou W., Nielsen J.B., Fritsche L.G., Dey R., Gabrielsen M.E., Wolford B.N., LeFaive J., VandeHaar P., Gagliano S.A., Gifford A., Bastarache L.A., Wei W.Q., Denny J.C., Lin M., Hveem K., et al. 2018. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341. https://doi.org/10.1038/s41588-018-0184-y
PheWeb v. 1.1.17. http://pheweb.sph.umich.edu/SAIGE-UKB/.
Funding
This work was partially supported by the State Assignment of the Ministry of Science and Higher Education (no. 075-00603-19-00).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. Ethical standards are met. The review was written using open source publications.
Additional information
Translated by A. Kashevarova
Abbrviation: GWAS, Genome-Wide Association Studies; BASC, BRCA1-associated genome surveillance complex.
Rights and permissions
About this article
Cite this article
Babushkina, N.P., Postrigan, A.E. & Kucher, A.N. Involvement of Variants in the Genes Encoding BRCA1-Associated Genome Surveillance Complex (BASC) in the Development of Human Common Diseases. Mol Biol 55, 278–296 (2021). https://doi.org/10.1134/S0026893321020047
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893321020047