Abstract
MODY is a group of genetically and clinically heterogeneous forms of diabetes characterized by autosomal dominant inheritance and is subdivided in 13 subtypes dependent on the gene involved. The subtype MODY9 is a very rare form caused by mutations in the gene encoding the PAX4 transcription factor which is engaged in differentiation of pancreatic beta-cells. PAX4 contains two DNA-binding domains—Paired and Homeo. Expression of the human PAX4 gene is tissue-specific. The alternatively spliced mRNA variants encode for protein isoforms which differ within their N- and C-terminal regions. In this study, the transcriptional activities of the human PAX4 variants, both known and new ones, were determined. The full-length PAX4 containing intact DNA-binding domains was found to have maximal activity in transient expression system of the firefly luciferase reporter gene under control of the insulin promoter in HEK293 cells. The transcriptional activity is significantly reduced in the variants lacking eight N-terminal amino acid residues and/or variants whose Homeo domain is truncated from the C-terminus. Similar data were obtained with the glucagon promoter reporter system. The aberrant PAX4 variants were shown to retain stability and nuclear localization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
The PAX4 transcription factor plays important roles in organogenesis of the pancreas during embryonic development, these are, regulation of proliferation, migration of precursor cells and coordination of their differentiation programs, and resistance of insulin-producing cells to stress factors. In murine pancreatic α- and β-cell lines, PAX4 is bound to promoter regulatory elements of the main islet hormones: glucagon [1, 2], grehlin [3], and insulin [4]. Mutations in the human PAX4 gene are found in patients suffering from MODY9, one of the forms of MODY monogenic diabetes (Maturity Onset Diabetes of Young) [5, 6]. In the mature pancreas, PAX4 participates in adaptation to adverse factors. PAX4 includes two DNA-binding domains, Paired and Homeo, located in the N-terminal and central regions, respectively [7]. The important role of DNA-binding domains in PAX4 functioning is confirmed by the fact that the majority of missense mutations found in patients with different forms of diabetes substitute amino acids located in these domains [8–10]. The C-terminal region contains the transactivation domain whose activity depends on the cell type in which the protein is expressed, and is frequently associated with activity of E1A-like proteins [11]. Gene expression can give rise to alternatively spliced mRNAs encoding for protein isoforms which differ in the structure of their N- and C-terminal regions. This can substantially affect interactions with gene regulatory regions and counterpart proteins. Such isoforms were previously identified in rodent and human insulinomas [12, 13]. The goal of this study is to compare the transcriptional activity of the known and new natural alternative PAX4 isoforms in transfected human HEK293 cells.
EXPERIMENTAL
Reagents. The ImProm II Reverse Transcription System (Promega, United States) was used to synthesize cDNA. Genomic DNA fragments were amplified by PCR with High-fidelity DNA polymerase (Thermo Fisher Scientific, United States). The DNA fragments for subcloning were obtained by PCR with the Tersus Plus PCR kit (Evrogen, Russia).
The following enzymes were also used: EcoRI, BglII, SalI, XhoI, BamHI, SmaI, NotI, KpnI, T4 polynucleotide kinase, calf intestine alkaline phosphatase (CIAP), and T4 DNA ligase (Thermo Fisher Scientific, United States).
The PCR products were cloned using the pGEM-T Easy vector (Promega, United States). The following vectors were used for construction: pcDNA3.1(+) (Invitrogen, United States), pGL4.10[luc2] (Promega), and pEGFP-C3 (Clontech, United States). The pRL-tk (Promega) plasmid was used to normalize luciferase activity.
Plasmid DNA was purified with a Plasmid Miniprep kit. The DNA fragments were isolated from agarose gel with a Cleanup Miniprep kit (Evrogen, Russia), and the reaction mixtures were purified with the same kit.
Oligonucleotides. Primers synthesized by Syntol (Russia) were used for DNA amplification and sequencing (Table 1).
Sequencing. Sequencing was performed in the Genome Center of Shared Equipment (IMB, Russian Academy of Scienses) with an ABI PRISM Model 3100 sequencing system (Applied Biosystems, United States). The Chromas program was used for data treatment.
Human cell culture. Cells of the HEK293 line (epithelial-like cells from human embryonic kidney) were cultured in a DMEM medium containing 1 g/L glucose, L-glutamine, phenol red, sodium bicarbonate/sodium pyruvate, 10% inactivated (50°C, 30 min) FBS, and 10 mg/mL gentamycin. The cells were grown in a thermostat at 37°C and 5% CO2.
Cell transfection. Cells were placed in a 24-well plate (75 000 cells per well) without antibiotic. Plasmid DNA (500 ng) encoding one of the PAX4 variants, 100 ng pGL4 plasmid carrying the luciferase gene under control of the human insulin or glucagon promoters, and 30 ng pRL-tk plasmid carrying the luciferase Renilla reporter gene under control of the thymidine kinase promoter from simplex herpes virus (Promega, United States) were placed in each well. Transfection was performed with the TurboFect reagent according to the ThermoScientific instruction manual. The cells were grown in the thermostat at 37°C and 5% CO2 for 48 h.
Luciferase activity measurement. 48 h post-transfection, the medium was removed from the wells and the HEK293 cells were washed with 1 mL PBS. Next, 100 µL Passive Lysis buffer (Promega, United States) was added to the cells which were incubated at room temperature (RT) for 10 min. Lysates were transferred to 1.5 mL tubes and centrifuged for 10 min at RT. Next, 60 µL lysate was placed in a well of a 96-well plate. The samples were supplied with 40 µL luciferase substrate from a Dual-Glo Luciferase Assay System kit (Promega), incubated at RT for 10 min, and luminescence was measured with a ChameleonV Multilabel detection platform (HIDEX) reader. Finally, the wells were filled with 40 µL solution of Dual-Glo® Stop & Glo® Substrate in Dual-Glo® Stop & Glo® Buffer, incubated at RT for 10 min, and luminescence of Renilla luciferase was measured. The values of the firefly luciferase activity were normalized to the Renilla luciferase activity to eliminate differences in transfection efficacy and sample treatment.
Immunoblotting. After 48 h transfection of the HEK293 cells with the plasmids encoding for the fused EGFP-PAX4 proteins (1 µg per well), the culture medium was removed and the wells were washed with 1 mL PBS. Next, 100 µL lysis buffer of 1 M Tris-HCl (pH 8.0), 5 M NaCl, 1% Trton X100, and 100× PMSF) was added, and the cells were incubated at 4°C for 10 min. The lysates were transferred to 1.5 mL tubes, incubated for 10 min on ice with constant agitation, and centrifuged at 12 000 g and 4°C for 10 min. Protein concentration in the supernatants was determined using the Bradford method.
To resolve proteins in PAAG, 5× loading buffer was added to lysate aliquots, containing 20 mg protein. The loading buffer included 312 mM Tris-HCl (pH 6.8), 50% glycerin, 25% β-mercaptoethanol, 10% SDS, and 0.01% bromophenol blue. The samples were boiled for 5 min and loaded on PAAG (5% concentrating and 12% resolving gel). Electrophoresis was run in a buffer of 25 mM Tris (pH 8.3), 192 mM Glycine, and 1% SDS at 20 mA, 140 V, and 3 W. Gel was placed in a transfer buffer containing 25 mM Tris (pH 8.3), 192 mM glycine, 20% ethanol. Nitrocellulose membranes were incubated for several minutes in the same buffer. Protein transfer was carried out for 1 h at 50 V, 130 mA, and 8 W. Next, the membrane was blocked in a TBST buffer (140 mM NaCl, 1 M Tris-HCl, pH 7.5, 0.1% Tween, and 10% fat-free milk) and incubated overnight at 4°C.
The membrane was washed in TBST and incubated with agitation at RT for 1 h with primary antibodies against green fluorescent protein—GFP (Sigma, United States)—diluted in TBST with the blocking agent. Next, the membrane was washed in 75 mL TBST, and secondary antibodies (rabbit anti-IgG) conjugated with horseradish peroxidase (GE-Health Care, United States) diluted in TBST with the blocking agent were added. The membrane was washed in TBST once more and treated with the SuperSignal™ West Femto Maximum Sensitivity Substrate reagent (Thermo Fisher Scientific, United States). Visualization was performed with a Chemidoc MP unit (Bio-Rad, United States).
Statistics. The GraphPad Prism6 program and one-way ANOVA test were used for data treatment.
RESULTS AND DISCUSSION
Cloning of cDNA Coding for Alternative PAX4 Variants and Constructing Vectors for Their Expression in Human Cells
A diagram of the human PAX4 gene localized on chromosome 7 (https://www.ucsc.edu/) is shown in Fig. 1. The gene consists of 11 exons and 10 introns, occupies the chr7:127610292-127616027 region, and has restricted cellular and tissue specificity of expression. PAX4 mRNA is present in differentiating and mature pancreatic β-cells, placenta [14], intestine, and some other tissues [15]. The PAX4 gene contains at least two promoters located before the first and second coding exons, which initiate transcription in β-cells and placenta, respectively (Fig. 1).
Diagram of exon–intron structure of the human PAX4 gene (https://genome.ucsc.edu/). Exons are shown as rectangles. The exon and intron size (bp) is shown below and above, respectively. TISPan and TISPla designate transcription initiation sites from the pancreatic and placental promoters, respectively.
PAX4 cDNAs were obtained by reverse transcription of total RNA from human placenta and PCR with the PAX4-2f and PAX4-10r primers. The PCR products were cloned in the pGEM-T-Easy vector. Sequencing was used to identify PAX4 isoforms which differed in the long (70 bp) or short (53 bp) exon 7, as well as the C-terminal region. The next step was reconstruction of the complete sequences of the known PAX4 variants represented in the Genome Bank (NM_001366110 and NM_001366111). To do this the N-terminal sequence (encoded by exon 1) in all variants was completed with the use of the PAX4-5′ primer. Moreover, the PAX4e10-f and PAX4e11-r primers were used to amplify the fragment corresponding to the exon 11 C-terminus. All structural differences in the reconstructed coding sequences of the PAX4 variants are summarized in Fig. 2. Plasmids with the PAX4 variants based on the pcDNA3.1(+) vector and designated for expression in human cells were constructed at the next step. This resulted in eight recombinant plasmids (I–VIII) for constitutive expression of the PAX4 variants under control of the cytomegalovirus (CMV) promoter. The plasmid structure is represented in Table 2 and Fig. 2.
Difference in primary structure ofPAX4 alternative variants. The exon and intron sequences are shown in upper case and italic, respectively. (a) The N-terminal regions encoded by exons 1 and 2. ATG initiating codons are underlined. (b) The structural difference in the region encoded by complete and truncated exon 7. (c) The structural difference in the C-terminal region encoded by exons 10 and 11.
The PAX4 alternative variants are represented in Fig. 3. Plasmid II carries the known PAX4 variant (NM_001366111) consisting of 348 amino acid residues, including the intact Paired (5–124 aa) and Homeo (162–222 aa) domains. Pax4 from plasmid I lacks an eight N-terminal aa and has the truncated Paired domain (Figs. 2a and 3). Plasmid IV encodes one more previously described PAX4 variant of 351 aa (NM_001366110). It differs from plasmid I product in the structure of the C-terminal region due to alternative splicing: mature mRNA has exon 11 instead of exon 10 (Fig. 2c and 3). The PAX4 variants derived from plasmids III and I are truncated from the N-terminus. Finally, the PAX4 sequences located on plasmids V–VIII are characterized by a 17 bp deletion of exon 7 (Fig. 2b). This deletion originates from the use of the alternative splicing acceptor site in exon 7 and leads to a reading frame shift. For this reason, the protein products are shorter (278 or 286 aa) and differ from those encoded by plasmids I–IV since the Homeo domain is 7 aa truncated from the C-terminus followed by a nonsense sequence.
Diagram of the domain structure of PAX4 alternative variants. The intact (Paired and Homeo) and truncated (–Paired and Homeo–) DNA-binding domains are shown as grey boxes. The different C-terminal regions are differently dashed. The Roman numerals on the left designate the plasmids carrying the corresponding PAX4 variants (Table. 2).
Cloning of Human Insulin and Glucagon Promoters and Constructing Reporter Plasmids
To compare transcriptional activity of the PAX4 variants, reporter plasmids with the luciferase gene under control of the human insulin and glucagon promoters were constructed. PAX4 was found to bind with the 5'-gccagacctgtccctgctcacagct-3' sequence in the insulin promoter [16]. The promoter fragment of the insulin gene was amplified with the InsPr-f1and InsPr-r primers (Table 1), which resulted in a 518 bp PCR fragment (chr11:2161063-2161580) containing the PAX4 binding site. The promoter fragment of the glucagon gene was amplified with the GCG-Pr-Kp-f and GCG-Pr-Bg-r primers (Table 1), which resulted in a 1726 bp PCR fragment (chr2:162152135-162153860). The promoter fragments were cloned in the pGEM-T-Easy vector. The cloned fragments were sequenced and inserted in the pGL4.10[luc2] vector upstream of the luciferase gene.
Transcriptional Activity of PAX4 Alternative Variants in Transfected Human HEK293 Cells
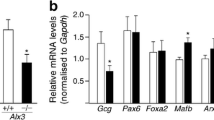
Transactivation properties of the PAX4 variants relative to the insulin promoter were determined in the transient expression system post-transfection. These results are represented in Fig. 4. Canonical PAX4 on the plasmid II (NM_001366111) with the DNA-binding Paired and Homeo domains exhibits the maximal transactivation. The luciferase activity level in the cells transfected with this plasmid exceeded that of the pcDNA3.1(+) vector by more than 3-fold. Transactivation capability of the other previously described variant (NM_001366110) located on plasmid IV, which has the significantly changed C-terminal region due to alternative splicing, is reduced 1.5-fold. The variants with the Paired domains truncated from the N-terminus (plasmids I and III) are about 2-fold less active compared with their full-length Paired counterparts. Finally, the short PAX4 variants (plasmids V–VIII) are almost totally inactive in the system used. This is the result of the C-terminal rearrangements and, possibly, the lack of a proposed activator domain [7]. Similar results were obtained with a reporter system based on the human glucagon promoter (Fig 5). The PAX4 variant with the intact DNA-binding domains (plasmid II) had the maximal activity compared to the glucagon promoter. The variant with the alternative structure of the C-terminal region is 2-fold less active, and the variants with the Homeo domain truncated from the C-terminus were also less active (Fig. 5).
Comparative transactivation of the human insulin gene by alternative isoforms of the PAX4 transcription factor. HEK293 cells were cotransfected with the pcDNA3.1(+) (C) plasmid or plasmids I–VIII carrying the different PAX4 isoforms (Table 2) and the plasmid containing the firefly luciferase gene under control of the human insulin promoter, as well as the pRL-tk plasmid containing the Renilla luciferase gene under control of the thymidine kinase promoter from the simplex herpes virus. Firefly luciferase activity is normalized to Renilla luciferase activity. The averages with the standard deviations from three independent experiments are shown (***P <0.001, ****P < 0.0001).
Transactivation characteristics of alternative PAX4 isoforms relative to the glucagon promoter. HEK293 cells were cotransfected with the pcDNA3.1(+) (C) plasmid or plasmids II, IV, VI or VIII carrying the different PAX4 isoforms (Table 2) and the plasmid containing the firefly luciferase gene under control of the human glucagon promoter, as well as the pRL-tk plasmid containing the Renilla luciferase gene under control of the thymidine kinase promoter from the simplex herpes virus. Firefly luciferase activity is normalized to Renilla luciferase activity. The averages with the standard deviations from three independent experiments are shown (*P <0.05, **P < 0.01).
Localization of Alternative Variants in Transfected HEK 293 Cells
The stability of aberrant proteins, or their different cellular localization may be reasons for the low activity of the PAX4 alternative variants. To confirm or deny this hypothesis, plasmids expressing proteins fused with EGFP were constructed. The DNA fragments encoding for the PAX4 variants were amplified from the plasmids I–VIII with the PAX4-Bg-f and PAX4-Sa-r primers (Table 1) and cloned in the pEGFP-C3 vector to give rise to the EGFP-PAX4 fusions. The coding frames were assessed by sequencing. The resulting plasmids were transfected in HEK293 cells and their lysates were analyzed for the production of the fused proteins by immunoblotting with the antibodies against GFP (Fig. 6). Immunoblotting revealed polypeptides of about 65 kD in the cells transfected with the pEGFP-PAX4-N1-10 and pEGFP-PAX4-N1-11 plasmids, which corresponds to the expected molecular weight of the fused proteins. The pEGFP-PAX4-N1-d7-10 and pEGFP-PAX4-N1-d7-11plasmids, as might be expected, produced shorter fusions of about 58 kD. There were no shorter products reacting with the antibodies, which indicates protein stability. Cellular protein localization was analyzed with fluorescent microscopy. Fluorescence in cell nuclei was found in all the cases (not shown). Thus, the PAX4 variants, are similar in their stability and intracellular localization.
Immunoblotting of fused EGFP-PAX4 proteins synthesized in HEK293 cells. M—Molecular weight markers. (1–4) The lysates from the cells transfected with the pEGFP-PAX4-N1-10, pEGFP-PAX4-N1-11, pEGFP-PAX4-N1-d7-10, and pEGFP-PAX4-N1-d7-11 plasmids, respectively. (5) The lysates from the cells transfected with the control pEGFP-С3 plasmid.
To summarize it can be concluded that the isoforms of the human PAX4 transcription factor with intact DNA-binding domains are transcription activators in the cotransfection system. This system includes HEK293 cells and plasmids based on the human insulin and glucagon promoters. The activity of the PAX4 transactivators with the Paired domain truncated from the N-terminus or the Homeo domain truncated from the C-terminus is significantly lower. The PAX4 variants with the completely changed C-terminal region, as the result of partial exon 7 deletion, are almost inactive with the insulin promoter, which may be associated with the absence of the transactivator domain. These variants can exhibit a dominant-negative effect similar to previously described isoforms identified in insulinomas [12, 13].
Recently, PAX4, as a factor of β-cell survival, became of more interest. PAX4 overexpression in the insulinoma line and islets of Langerhans is known to enhance expression of the antiapoptotic Bcl-xL factor [17]. Inhibition of PAX4 decreases the Bcl-xL expression level, which correlates with development of spontaneous apoptosis and higher sensitivity to cytokine induced apoptosis. Murine insulinoma cells are prevented from apoptosis induced with the TNF-α necrosis factor, which leads to higher Bcl-xL expression when the cells are treated with the recombinant PAX4 protein [18]. Overexpression of the PAX4 mutant does not elevate the Bcl-2 expression level and does not prevent murine islet β-cells from apoptosis caused by cytokines, while overexpression of wild type PAX4 in β-cells of mature murine pancreatic islets prevents stress-induced hyperglycemia [19]. It is also shown that PAX4 can participate in a compensatory response of β-cells to metabolic stress [20]. Therefore, possible effects on pathways regulated with PAX4 participation, in the case of diabetes, are discussed [21]. When studying these processes, possible heterogeneity of the PAX4 variants, including those with dominant-negative effects, must be taken into account.
REFERENCES
Petersen H.V., Jørgensen M.C., Andersen F.G., Jensen J., F-Nielsen T., Jørgensen R., Madsen O.D., Serup P. 2000. Pax4 represses pancreatic glucagon gene expression. Mol. Cell Biol. Res. Commun.3, 249–254.
Estreicher A., Gauthier B.R., Mamin A., Edlund H., Philippe J. 2002. The pancreatic beta-cell-specific transcription factor Pax4 inhibits glucagon gene expression through Pax 6. Diabetologia.45, 97–107.
Wang Q., Elghazi L., Martin S., Martins I., Srinivasan R.S., Geng X., Sleeman M., Houghton J.,Sosa-Pineda B. 2008. Ghrelin is a novel target of Pax4 in endocrine progenitors of the pancreas and duodenum. Dev. Dyn.237, 51–61.
Campbell S.C., Cragg H., Elrick L.J., Macfarlane W.M., Shennan K.I.J., Docherty K. 1999. Inhibitory effect of Pax4 on the human insulin and islet amyloid polypeptide (IAPP) promoters. FEBS Lett.463, 53–57.
Vaxillaire M., Froguel P. 2008. Monogenic diabetes in the young, pharmacogenetics and relevance to multifactorial form of type 2 diabetes. Endocr. Rev.29, 254–264.
Biason-Lauber A., Boehm B., Lang-Muritano M., Gauthier B.R., Brun T., Wollheim C.B., Schoenle E.J. 2005. Association of childhood type 1 diabetes mellitus with a variant of PAX4: Possible link to beta cell regenerative capacity. Diabetologia. 48, 900–905.
Smith S.B., Ee H.C., Conners J.R., German M.S. 1999. Paired-homeodomain transcriptional repressor in early pancreatic development. Mol. Cell. Biol.8, 8272–8280.
Plengvidhya N., Kooptiwut S., Songtawee N., Doi A., Furuta H., Nishi M., Banchuin N. 2017. PAX4 mutations in Thais with maturity onset diabetes of the young. J. Clin. Endocrinol. Metab.92, 2821–2826.
Martin-Montalvo A., Lorenzo P.I., Lopez-Noriega L., Gauthier B.R. 2016. Targeting pancreatic expressed PAX genes for the treatment of diabetes mellitus and pancreatic neuroendocrine tumors. Expert. Opin. Ther. Targets. 21, 77–89.
Sujjitjoon J., Kooptiwut S., Chongjaroen N., Tangjittipokin W., Plengvidhya N., Yenchitsomanus P.T. 2016. Aberrant mRNA splicing of paired box 4 (PAX4) IVS7–1G>A mutation causing maturity-onset diabetes of the young, type 9. Acta Diabetol. 53, 205–216.
Fujitani Y., Kajimoto Y., Yasuda T., Matsuoka T.A., Kaneto H., Umayahara Y., Fujita N., Watada H., Miyazaki J.I., Yamasaki Y., Hori M. 1999. Identification of a portable repression domain and an E1A-responsive activation domain in Pax4: A possible role of Pax4 as a transcriptional repressor in the pancreas. Mol. Cell. Bio-l.19, 8281–8291.
Tokuyama Y., Yagui K., Sakurai K., Hashimoto N., Saito Y., Kanatsuka A. 1998. Molecular cloning of rat Pax4: Identification of four isoforms in rat insulinoma cells. Biochem. Biophys. Res. Commun. 248, 153–156.
Miyamoto T., Kakizawa T., Ichikawa K., Nishio S., Kajikawa S., Hashizume K. 2001. Expression of dominant negative form of PAX4 in human insulinoma. Biochem. Biophys. Res. Commun. 282, 34–40.
Lorenzo P.I., Ju F., Cobo-Vuilleumier N., Garc M., Gauthier B.R. 2017. The diabetes-linked transcription factor PAX4 : From gene to functional consequences. Genes.8, 101. https://doi.org/10.3390/genes8030101
https://www.proteinatlas.org/
Brink C., Chowdhury K., Gruss P. 2001. Pax4 regulatory elements mediate beta cell specific expression in the pancreas. Mech. Dev.100, 37–43.
Brun T., Duhamel D.L., He K.H.H., Wollheim C.B., Gauthier B.R. 2007. The transcription factor PAX4 acts as a survival gene in INS-1E insulinoma cells. Oncogene.26, 4261–4271.
Lu J., Li G., Lan M. S., Zhang S., Fan W., Wang H., Lu D. 2007. Pax4 paired domain mediates direct protein transduction into mammalian cells. Endocrinology.148, 5558–5565.
Hui K., He H., Lorenzo P.I., Brun T., Moreno C.M.J., Aeberhard D., Gauthier B.R. 2011. In vivo conditional Pax4 overexpression in mature islet β-cells prevents stress-induced hyperglycemia in mice. Diabetes. 60, 1705–1715.
Lee G., Jang H., Kim Y.Y., Choe S.S., Kong J., Hwang I., Park J., Im S.-S., Kim J.B. 2019. SREBP1c-PAX4 axis mediates pancreatic β-cell compensatory responses upon metabolic stress. Diabetes.68, 81–94.
Lorenzo P.I., Cobo-Vuilleumier N., Gauthier B.R. 2018. Therapeutic potential of pancreatic PAX4-regulated pathways in treating diabetes mellitus. Curr. Opin. Pharmacol. 43, 1–10.
Funding
This study was supported by the Russian Foundation for Basic Research (project nos. 17-29-06028 and 18-04-01271).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare no conflict of interest.
Statement of the welfare of animals. No studies on animals or humans were carried out.
Additional information
Translated by A. Boutanaev
Rights and permissions
About this article
Cite this article
Melnikova, A.I., Krasnova, T.S., Zubkova, N.A. et al. Alternative Variants of Pax4 Human Transcription Factor: Comparative Transcriptional Activity. Mol Biol 54, 749–756 (2020). https://doi.org/10.1134/S0026893320050076
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893320050076