Abstract—
An alkaliphilic, sulfate-reducing, anaerobic bacterium (strain H1T) was isolated from a terrestrial mud volcano at the Taman Peninsula, Russia. The cells of the isolate were gram-negative motile vibrios, 1 µm in diameter and 2.0–2.5 µm in length. Strain H1T grew at 14–42°C (optimum at 37°C), pH 8.5–10.5 (optimum at pH 9.5), and NaCl concentrations of 0.5–6% (wt/vol) (optimum at 0.5–1.5%); pyruvate, lactate, butyrate, caproate, or pelargonate were used as electron donors, and elemental sulfur, sulfite, or sulfate were used as electron acceptors. Pyruvate and lactate were fermented. No growth occurred in the presence of oxygen. Thiosulfate, DMSO, fumarate, nitrate, nitrite, arsenate, selenite, and Fe(III) were not used as electron acceptors. Elemental sulfur, thiosulfate, and sulfite were not disproportionated. Glucose, fructose, sucrose, trehalose, galactose, xylose, fumarate, citrate, yeast extract, and peptone were not fermented. Predominant fatty acids were C20:0 (54.2%), C22:0 (24.6%), and C18:0 (11.1%). The genome of strain H1T was 3.66 Mb in size and had G + C DNA content of 51.1%. The genome contained the genes encoding the enzymes of dissimilatory sulfate reduction and β-oxidation of fatty acids. According to the results of analysis of the 16S rRNA gene sequence, Desulfobotulus mexicanus was the organism most closely related to strain Н1Т (98.3% similarity). Based on its phenotypic characteristics and the data of phylogenetic analysis, affiliation of the isolate as member of a novel Desulfobotulus species, Desulfobotulus pelophilus sp. nov., is proposed, with the type strain H1T (=DSM 112796T = VKM B-3697Т =UQM 41590T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Mud volcanism is a widespread geological phenomenon, which plays a significant part in the balance of atmospheric methane (Mazzini and Etiope, 2017). Terrestrial mud volcanoes (TMV) result from discharge of the particles of clay and silt, breccia, fluids, and gases from deep sediment layers. The Taman Peninsula is one of the regions with the most intensive mud volcanism. Over 100 active TMV are known in the Kerch-Taman mud volcano province (Gnatenko et al., 1986; Kholodov et al., 2012). Alkaline pH (over 8.5) of the fluids of these environments favor development of alkaliphilic microorganisms (Khomyakova et al., 2020, 2022; Frolova et al., 2021a, 2021b). In TMVs, anaerobic prokaryotes are involved in the biogeochemical cycles of carbon, sulfur, and other elements. Sulfate-reducing bacteria may participate in anaerobic methane oxidation as syntrophic partners of methanotrophic archaea (Knittel and Boetius, 2009). While the data on cultivation of sulfate-reducing bacteria from TMV are not abundant (Alain et al., 2006; Frolova et al., 2021a, 2021b), molecular analysis revealed in these communities the presence of microorganisms phylogenetically related to the known sulfate reducers (Tu et al., 2017; Ren et al., 2018; Merkel et al., 2021). The order Desulfobacterales, one of the largest and oldest taxonomic groups of sulfate reducers, is characterized by utilization of fatty acids as carbon and energy sources (Kuever et al., 2015). Members of Desulfobacterales inhabit anoxic sediments of freshwater, marine, and soda basins and may belong to various physiological groups (alkaliphiles, psychrophiles, halophilic, magnetotactic, and hydrocarbon-degrading). According to the List of Prokaryotic Names with Standing in Nomenclature, this order comprises 37 genera with validly published names (Parte et al., 2020).
The present work describes a strain of anaerobic, salt-tolerant, obligately alkaliphilic sulfate-reducing bacterium isolated from a mud volcano at the Taman Peninsula and identified as a new Desulfobotulus species.
MATERIALS AND METHODS
Source of isolation. The sample of the mud volcano fluid containing liquid and solid fractions was collected in May 2017 from an active gryphon of the Gnilaya Gora terrestrial mud volcano, Taman Peninsula, Krasnodar krai, Russia, GPS coordinates: 45.251° N, 37.436° E. The temperature at the sampling site was 21°C, pH 8.5, Cl– concentration, 15.7 mM, and SO\(_{4}^{{2 - }}\) concentration, 5.3 mM. The sample was collected anoxically into a plastic tube with a tight screw cap and was transported to the laboratory at ambient temperature for further investigation.
Media and cultivation. For the isolation and routine cultivation of strain H1T, an anoxic, reduced brackish medium was used containing the following (g/L distilled water): KH2PO4, 0.33; NH4Cl, 0.33; KCl, 0.33; CaCl2·6H2O, 0.33; NaHCO3, 2.00; MgCl2·6H2O, 0.33; NaCl, 20.0; resazurin, 0.001; vitamin solution (Wolin et al., 1963), 1 mL; and trace element solution (Slobodkin et al., 2012), 1 mL. The medium was prepared with boiling and cooling under a continuous N2 flow, then the reducing agent (Na2S·9H2O) was added. The medium was dispensed (10 mL) into 17‑mL Hungate tubes and autoclaved for 60 min at 121°C. After sterilization, pH was 9.0. Sodium pyruvate (10 mM) and sodium sulfate (14 mM) were added from sterile stock solutions prior to inoculation.
Phenotypic characteristics. Cell morphology and motility were examined in 48-h liquid cultures under a Zeiss Primo Star phase contrast microscope. Growth experiments were carried out in triplicate. For morphological, physiological, and metabolic characterization, strain H1T was grown in the medium used for its isolation, if not stated otherwise. The ranges of temperature, pH, and salinity were determined in reduced medium with pyruvate and sulfate. The salinity range studied was 0–10% NaCl (wt/vol). Different pH values were adjusted using the following buffers (Good’s buffers, Sigma-Aldrich, 30 g/L): MES (pH 6 and 6.5), HEPES (pH 7 and 7.5), Tricine (pH 8.0 and 8.5), CAPSO (pH 9.0 and 9.5), and CAPS (pH 10 and 11). The experiments with sulfur compounds and oxygen were carried out in unreduced medium.
Analytical procedures. Gaseous products of metabolism were analyzed by gas chromatography on a HayeSep N 80/100 mesh column at 40°C and flow rate of 20 mL/min. The carrier gas was argon. Sulfide was determined colorimetrically with dimethyl-p-phenylenediamine (Trüper and Schlegel, 1964).
Fatty acid composition. The composition of cellular fatty acids was determined as described previously (Slobodkina et al., 2020), using direct methylation of lyophilized biomass and chromato-mass spectral analysis; the concentrations were determined by internal normalization using the peak areas of the full ionic current of the fatty acid methyl esters.
DNA isolation, sequencing, and full genome analysis. DNA isolation for determination of the nucleotide sequence of the 16S rRNA gene and full-genome sequencing was carried out using the FastDNA Spin Kit (MP Bio), according to the manufacturer’s protocol. The 16S rRNA gene was amplified using the universal bacterial primers 27F, 357F, 530F, 1114F, 342R, 519R, and 1492R (Weisburg et al., 1991). PCR products were sequenced by the Sanger method. Preliminary phylogenetic screening of the 16S rRNA gene sequences was carried out using the GenBank database (Benson et al., 1999) with the BLAST software package (Altschul et al., 1990). For more accurate determination of the phylogenetic position of the isolate, the 16S rRNA gene sequence was aligned with those of the reference strains with Clustal W (Thompson, 1997). Phylogenetic analysis was carried out using MEGA 7.0 (Kumar et al., 2016). Statistical significance of the branching order was determined by bootstrap analysis of 1000 alternative trees (Felsenstein, 1985) constructed by the methods implemented in the MEGA 7 software package.
The genome of strain H1T was sequenced using MiSeq (Illumina, San Diego, California, United States). Gene search and annotation were carried out with the RAST server (Brettin et al., 2015). SEED was used to distribute the hypothetic genes according to the subsystem categories (Overbeek et al., 2014). The taxonomic position of strain H1T was verified using two methods: average nucleotide identity (ANI), as determined using the EzBioCloud ANI calculator (https://www.ezbiocloud.net/tools/ani) (Yoon et al., 2017) and in silico DNA-DNA hybridization, as determined using the Genome-to-Genome Distance Calculator (http://ggdc.dsmz.de) (Meier-Kolthoff et al., 2013).
The 16S rRNA gene sequence of strain H1T was deposited to GenBank/EMBL under accession no. MW872673.
The full genome sequence of strain H1T was depo-sited to GenBank/EMBL under accession no. JAPFPW010000000.
RESULTS
Isolation of the pure culture. Enrichment cultures were obtained by inoculation of the environmental sample ~10% vol/vol) into the sterile anoxic medium with pyruvate and sulfate. Abundant microbial growth was observed after 48-h incubation of the enrichments at 30°C. After three sequential transfers (5% vol/vol), tenfold serial dilutions of the culture were used to inoculate the same liquid medium. Only one morphological type of the cells was found in the highest dilution showing growth (10−8). Tenfold dilutions were repeated twice more, and the culture grown in the last dilution was designated strain H1T. Results of the 16S rRNA gene sequencing confirmed the purity of this culture. No colonies could be obtained under anoxic conditions either by the roll-tube method or in 1% Gelrite gellan gum or 1% agar.
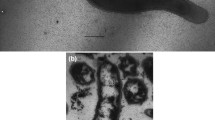
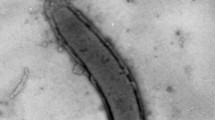
The cells of strain H1T were vibrios, 1 µm in diameter and 2.0–2.5 µm in length, growing singly and motile due to the single polar flagellum. No endospores were observed during 30 days of cultivation.
Growth physiology. Strain H1T grew at the temperatures from 14 to 42°C, with the optimum at 37°C. After 20-day incubation, no growth occurred at 50°C or higher and at 10°C or lower. The pH range for growth was 8.5 to 10.5, with the optimum at pH 9.5; no growth occurred at pH 8.0 and lower or at pH 11.0 and higher. Strain H1T grew at NaCl concentrations in the medium from 0.5 to 6.0% (wt/vol) with the optimum at 0.5–1.5%; no growth occurred at NaCl concentrations 7% NaCl or above.
Electron donors and acceptors. Strain H1T could use organic acids as electron donors and sulfur compounds as electron acceptors for growth. Addition of yeast extract (0.2 g/L) stimulated growth slightly, but was not necessary. The highest cell concentration (~5×107 cells/mL) was observed on pyruvate, both with sulfate or without electron acceptors. Pyruvate, lactate (10 mM), butyrate (10 mM), caproate (5 mM), caprylate (1 mM), and pelargonate (1 mM) could be used as electron donors in the presence of sulfate as an electron acceptor. In this case, sulfide and acetate were the reaction products. The compounds not used as electron donors (10 mM, if another concentration is not indicated) with sulfate as an electron acceptor were: acetate, formate, formate + acetate (2 mM) as a carbon source, propionate succinate, fumarate, malate, methanol, ethanol, butanol, isobutanol, glycerol, ribose, glucose, fructose, tributyrate, valerate, palmitate (1 mM), stearate (1 mM), oleate (1 mM), trioleate (1 mM), yeast extract (2 g/L), and molecular hydrogen (H2 + CO2, 80 : 20% in the gas phase). Elemental sulfur (5 g/L), sulfite (10 mM), and sulfate (14 mM) were used as electron acceptors in the presence of butyrate as an electron donor. Thiosulfate, dimethyl sulfoxide (DMSO), fumarate, nitrate, nitrite, arsenate, and selenite (all at 10 mM), as well as ferrihydrite (poorly crystalline Fe(III) oxide, 90 mM), were not used by strain H1T as electron acceptors with butyrate as an electron donor. The strain was incapable of growth under aerobic or microaerobic conditions (up to 3% oxygen).
Pyruvate and lactate (10 mM each) were fermented with formation of acetate and hydrogen as the end products. Strain H1T did not ferment formate, fumarate, citrate, succinate, fructose, sucrose, trehalose, galactose, xylose, peptone, or yeast extract after three weeks of incubation. The strain was incapable of disproportionation of elemental sulfur, thiosulfate, and sulfite both in the presence of ferrihydrite (acting as as a sulfide-scavenging agent.) and without it, in the bottles with large volumes of the gas phase.
Fatty acid composition. Cellular fatty acids of strain H1T were represented by a mixture of unbranched saturated and unsaturated acids: C20:0 (54.2%), C22:0 (24.6%) and C18:0 (11.1%). C16:0 (3.8%), C24:0 (2.4%), C18:1 ω7с (2.2%), and C18:1 ω9с (1.8%) were also pres-ent.
Phylogeny. The 16S rRNA gene sequences of strain H1T obtained by amplification with the universal bacterial primers and by full genome sequencing were identical. Comparison of 1539 nucleotides of the 16S rRNA gene sequence of strain H1T with the GenBank sequences (Benson et al., 1999) revealed that the isolate belonged to the genus Desulfobotulus, class Deltaproteobacteria with 98.31% similarity to the sequence of Desulfobotulus mexicanus (Pérez-Bernal et al., 2020). Reconstruction of the 16S rRNA-based phylogenetic tree showed that strain H1T represented a monophyletic lineage clearly separated from the most closely related species (Fig. 1).
Phylogenetic tree constructed using the 16S rRNA gene sequences and showing the position of strain H1T and related microorganisms. The tree was reconstructed using the maximum-likelihood method. The trees constructed using the neighbor-joining and minimum-evolution algorithms showed the same topology. Each numeral indicates the value of initial load of 1000 replicates. Scale bar, 0.020 replacements per nucleotide position. The GenBank accessionnumbers are indicated in parentheses.
The result of pairwise comparison of the average nucleotide identity of the genomes of strain H1T and its closest relative, D. mexicanus (DSM 105758T), was 88.7%. The value of in silico DNA-DNA hybridization between H1T and D. mexicanus (DSM 105758T) according to the recommended formula 2 was 26.10%. Both values were considerably lower than the thre-sholds recommended for delineation between prokaryotes at the species level, i.e., 95–96% (for ANI) and 70% (for DNA-DNA hybridization) (Meier-Kolthoff et al., 2013; Rodriguez and Konstantinidis, 2016).
General characteristics of the genome. The genome of strain H1T assembled out of 86 contigs had the overall length of 3 656 775 nt and N50 of 160 366 nt. The G + C content of genomic DNA was 51.1%. The genome contained 3783 nucleotide sequences encoding proteins and 55 RNA genes. Most of the annotated genes were responsible for the synthesis of amino acids and their derivatives (151), protein metabolism (147), carbohydrate metabolism (101), respiration (83), cofactors, vitamins, prosthetic groups, and pigment formation (55).
The genome of strain H1T contained the genes of the Embden−Meyerhof−Parnas pathway, including the NAD-dependent glyceraldhyde-3-phosphate dehydrogenase (WP_265423680), triose phosphate isomerase (WP_265423679), glucose-6-phosphate isomerase (WP_265426043), phosphoglycerate kinase (WP_265423283), enolase (WP_265423752), pyruvate kinase (WP_265423750), 6-phosphofructokinase (WP_265423438), fructose-bisphosphate aldolase (WP_265423362), and phosphoglycerate mutase (WP_265426113). The gene encoding hexokinase, which catalyzes glucose phosphorylation at the first stage of glycolysis, was, however, absent. The tricarboxylic acids cycle was encoded incompletely and did not contain the genes for malate dehydrogenase, succinate dehydrogenase, and succinyl-CoA synthetase.
The genome of strain H1T contained a complete set of genes required for dissimilatory sulfate reduction (Pereira et al., 2011), including sulfate adenylyl transferase Sat (WP_26542348), inorganic pyrophosphatase (WP_265426333), the AprA (WP_265426087) and AprB (WP_265426088) subunits of the APS reductase, the components of the dissimilatory sulfite reductase (WP_26542589−WP_265425892), and the electron-transporting complexes DsrMKJOP and QmoABC (WP_265425905−WP_265425909).
The genome of strain H1T contained two genes encoding the molybdopterin oxidoreductase (WP_265423973 and WP_265426274). The biochemical function of these enzymes is not clear. They are most probably not the catalytic subunit of the psrA polysulfide reductase, since the characteristic adjacent genes of such molybdopterin oxidoreductases were not revealed in the genome of strain H1T. The genome encoded the enzymes rhodanese (WP_265423671) and the hdr-like complex with the hdrA (WP_265423457), hdrB (WP_265425454), and hdrC (WP_265425453) subunits, which are involved in the redox reactions of sulfur compounds, although their biochemical mechanism is presently insufficiently studied (Zhang et al., 2021).
The genome of strain H1T contained all genes required for β-oxidation of fatty acids, including acyl-CoA dehydrogenase (WP_265423377, WP_ 265423535, WP_265424553, WP_265424646, WP_265425157, WP_265425230, WP_265425661, WP_265425744, WP_265426163), enoyl-CoA hydratase (WP_265423968, WP_265425159, WP_ 265425648, WP_265425692), 3-hydroxyacyl-CoA dehydrogenase (WP_265423389, WP_265425692), and 3-ketoacyl-CoA thiolase (WP_265423278, WP_265424057, WP_265423533, WP_265425693).
The genome of strain H1T contained all the genes encoding the components of the nitrogenase complex required for dinitrogen fixation, including the Mo-Fe- and Fe- proteins of nitrogenase nifHDK, (WP_265425310−WP_265425314), as well as the proteins required for its asembly and regulation: nifENB (WP_265425306–WP_265425309), nifU (WP_26542406), nifS (WP_265424064), NifA (WP_265424773), and NtrXY (WP_265425703–WP_265425704).
The genome of strain H1T contained the genes encoding catalase (WP_265425612), superoxide dismutase (WP_265425250), quinol oxidase CydAB (WP_265424027−WP_265424028), and several copies of the genes of the proteins involved in protection against oxidative stress, including the rubredoxin Rbo (WP_265424029, WP_265425910) and the rubrerythrin Rbr (WP_265425583), which probably carry out this function in Desulfovibrio vulgaris (Lumppio et al., 2001).
DISCUSSION
Strain H1T was isolated from a terrestrial mud volcano at the Taman Peninsula. It is an anaerobic alkaliphilic mesophilic sulfate-reducing bacterium.
Phylogenetic analysis based on the 16S rRNA gene sequences showed that strain H1T formed a separate lineage within the genus Desulfobotulus (Kuever et al., 2005), of the family Desulfobacteraceae, order Desulfobacterales, phylum Pseudomonadota. At the time of publishing, the genus Desulfobotulus was represented by three species with validly published names: D. sapovorans (Kuever et al., 2005), D. alkaliphilus (Sorokin et al., 2010), and D. mexicanus (Pérez-Bernal et al., 2020). Identity of the 16S rRNA nucleotide sequences between strain H1T and D. mexicanus (DSM 105758T) was 98.31%. Pairwise comparisons of the average nucleotide identity (ANI) and in silico DNA‒DNA hybridization between strain H1T and D. mexicanus (DSM 105758T) also supported identification of the isolate as a new Desulfobotulus species.
Members of the genus Desulfobotulus are geographically widespread. They have been detected in diverse ecosystems, including freshwater habitats and alkaline soda or crater lakes (GBIF Secretariat, 2021; GBIF Backbone Taxonomy. Checklist dataset https:// doi.org/10.15468/39omei). Strain H1T is the first member of this genus isolated from a terrestrial mud volcano. The new isolate had the growth ranges of pH, temperature, and salinity, which were close to the parameters of its habitat.
The metabolic potential encoded in the genome of strain H1T was in agreement with the phenotypic data. All Desulfobotulus species are characterized by utilization of aliphatic fatty acids in dissimilatory sulfate reduction. The differentiating characteristics of strain H1T and the known Desulfobotulus species are presented in Table 1. The most noticeable differences were its higher temperature optimum and ability to ferment lactate. Thus, based on our data, we propose to assign strain H1T to a new Desulfobotulus species, Desulfobotulus pelophilus sp. nov.
Description of Desulfobotulus pelophilus sp. nov.
Desulfobotulus pelophilus (pe.lo’phi.lus. Gr. m. n. pêlos, mud, N. Lat. adj. philus -a -um, loving; from Gr. adj. philos -ê -on, loving; N. Lat. adj. pelophilus, loving mud, since the species was isolated from a mud volcano).
The cells are motile vibrios, 1 µm in diameter and 2.0–2.5 µm in length. Growth occurs at 14–42°C (optimum at 37°C), pH 8.5–10.5 (optimum at 9.5), and NaCl concentrations (wt/vol) 0.5–6% (optimum at 0.5–1.5%). Pyruvate, lactate, butyrate, caproate, caprylate, and pelargonate are used as electron donors; elemental sulfur, sulfite, and sulfate are used as electron acceptors. Pyruvate and lactate are fermented. No growth occurs in the presence of oxygen. Thiosulfate, DMSO, fumarate, nitrate, nitrite, arsenate, selenite, and Fe(III) are not used as electron acceptors. No disproportionation of elemental sulfur, thiosulfate, or sulfite. Glucose, fructose, sucrose, trehalose, galactose, xylose, fumarate, citrate, yeast extract, and peptone are not fermented. Predominant cellular fatty acids are C20:0, C22:0, and C18:0.
The type strain H1T (=DSM 112796T = VKM B‑3697Т = UQM 41590T) was isolated from a terrestrial mud volcano at the Taman Peninsula. Genome size is 3.66 Mb. The G + C content of genomic DNA is 51.1% (full genome sequencing).
REFERENCES
Alain, K., Holler, T., Musat, F., Elvert, M., Treude, T., and Krüger, M., Microbiological investigation of methane- and hydrocarbon-discharging mud volcanoes in the Carpathian Mountains, Romania, Environ. Microbiol., 2006, vol. 8, pp. 574–590.
Altschul, S.F., Gish, W., Miller, W., Myers, E.W., and Lipman, D.J., Basic local alignment search tool, J. Mol. Biol., 1990, vol. 215, pp. 403–410.
Benson, D.A., Boguski, M.S., Lipman, D.J., Ostell, J., Ouellette, B.F.F., Rapp, B.A., and Wheeler, D.L., GenBank, Nucleic Acids Res., 1999, vol. 27, pp. 12–17.
Brettin, T., Davis, J.J., Disz, T., Edwards, R.A., Gerdes, S., Olsen, G.J., Olson, R., Overbeek, R., Parrello, B., Pusch, G.D., Shukla, M., Thomason, J.A., Stevens, R., Vonstein, V., Wattam, A.R., and Xia, F., RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes, Sci. Rep., 2015, vol. 5, p. 8365.
Felsenstein, J., Confidence limits on phylogenies: an approach using the bootstrap, Evolution (N.Y.), 1985, vol. 39, p. 783.
Frolova, A.A., Merkel, A.Y., Kuchierskaya, A.A., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., Pseudodesulfovibrio alkaliphilus, sp. nov., an alkaliphilic sulfate-reducing bacterium isolated from a terrestrial mud volcano, Antonie van Leeuwenhoek, 2021b, vol. 114, pp. 1387–1397.
Frolova, A., Merkel, A.Y., Novikov, A.A., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., Anaerotalea alkaliphila gen. nov., sp. nov., an alkaliphilic, anaerobic, fermentative bacterium isolated from a terrestrial mud volcano, Extremophiles, 2021a, vol. 25, pp. 301–309.
GBIF Secretariat, GBIF Backbone Taxonomy. Checklist dataset, 2021. https://doi.org/10.15468/39omei
Gnatenko, G.I., Kutnii, V.A., Naumenko, P.I., Sobolevskii, Yu.V., and Shnyukov, E.F., Gryazevye vulkany Kerchensko-Tamanskoi oblasti (atlas) (Mud Volcanoes of the Lerch-Taman Region: An Atlas), Kiev: Nauk. Dumka, 1986.
Kholodov, V.N., Mud volcanoes: distribution and genesis, Geologiya i Poleznye Iskopaemye Mirovogo Okeana, 2012, vol. 4, pp. 5–27.
Khomyakova, M.A., Merkel, A.Y., Kopitsyn, D.S., and Slobodkin, A.I., Pelovirga terrestris gen. nov., sp. nov., anaerobic, alkaliphilic, fumarate-, arsenate-, Fe(III)- and sulfur-reducing bacterium isolated from a terrestrial mud volcano, Syst. Appl. Microbiol., 2022, vol. 45, no. 2, p. 126304.
Khomyakova, M.A., Merkel, A.Y., Petrova, D.A., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., Alkalibaculum sporogenes sp. nov., isolated from a terrestrial mud volcano and emended description of the genus Alkalibaculum, Int. J. Syst. Evol. Microbiol., 2020, vol. 70, pp. 4914–4919
Knittel, K. and Boetius, A., Anaerobic oxidation of methane: progress with an unknown process, Annu. Rev. Microbiol., 2009, vol. 63, pp. 311–334. https://doi.org/10.1146/annurev.micro.61.080706.093130
Kuever, J., Rainey, F.A., and Widdel, F., Class IV. Deltaproteobacteria class nov., in Bergey’s Manual of Systematic Bacteriology, Brenner, D.J., Krieg, N.R., and Staley, J.T., Eds., Boston: Springer, pp. 970–971.
Kuever, J., Rainey, F.A., and Widdel, F., Desulfobacterales ord. nov., in Bergey’s Manual of Systematics of Archaea and Bacteria, Trujillo, M.E., Dedysh, S., DeVos, P., Hedlund, B., Kämpfer, P., Rainey, F.A., and Whitman, W.B., Eds., 2015. https://doi.org/10.1002/9781118960608.obm00084
Kumar, S., Stecher, G., and Tamura, K., MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets, Mol. Biol. Evol., 2016, vol. 33, pp. 1870–1874.
Lumppio, H.L., Shenvi, N.V., Summers, A.O., Voordouw, G., and Kurtz, D.M., Jr., Rubrerythrin and rubredoxin oxidoreductase in Desulfovibrio vulgaris: a novel oxidative stress protection system, J. Bacteriol., 2001, vol. 183, no. 1, pp. 101−108. Erratum in: J. Bacteriol., 2001, vol. 183, no. 9. PMID: 11114906; PMCID: PMC94855.https://doi.org/10.1128/JB.183.1.101-108.2001
Mazzini, A. and Etiope, G., Mud volcanism: an updated review, Earth-Sci. Rev., 2017, vol. 168, pp. 81–112.
Meier-Kolthoff, J.P., Auch, A.F., Klenk, H.P., and Göker, M., Genome sequence-based species delimitation with confidence intervals and improved distance functions, BMC Bioinformatics, 2013, vol. 14, p. 60.
Merkel, A.Y., Chernyh, N.A., Pimenov, N.V., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., Metabolic potential of the terrestrial mud volcano microbial community with a high abundance of archaea mediating the anaerobic oxidation of methane, Life, 2021, vol. 11, p. 953.
Overbeek, R., Olson, R., Push, G.D., Olsen, G.J., Davis, J.J., Disz, T., Edwards, R.A., Gerdes, S., Parrello, B., Shukla, M., Vonstein, V., Wattam, A.R., Xia, F., and Stevens, R., The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST), Nucl. Acids Res., 2014, vol. 42, no. D1, pp. D206–D214.
Parte, A.C., Sardà Carbasse, J., Meier-Kolthoff, J.P., Reimer, L.C., and Göker, M., List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ, Int. J. Syst. Evol. Microbiol., 2020, vol. 70, pp. 5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pereira, I.A.C., Ramos, A.R., Grein, F., Marques, M.C., da Silva, S.M., and Venceslau, S.S., A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea, Front Microbiol., 2011, vol. 2, p. 69. https://doi.org/10.3389/fmicb.2011.00069
Pérez-Bernal, M.F., Brito, E.M.S., Bartoli, M., Aubé, J., Ollivier, B., Guyoneaud, R., and Hirschler-Réa, A., Desulfobotulus mexicanus sp. nov., a novel sulfate-reducing bacterium isolated from the sediment of an alkaline crater lake in Mexico, Int. J. Syst. Evol. Microbiol., 2020, vol. 70, pp. 3219–3225. https://doi.org/10.1099/ijsem.0.004159
Ren, G., Ma, A., Zhang, Y., Deng, Y., Zheng, G., Zhuang, X., Zhuang, G., and Fortin, D., Electron acceptors for anaerobic oxidation of methane drive microbial community structure and diversity in mud volcanoes, Environ. Microbiol., 2018, vol. 20, pp. 2370–2385.
Rodriguez-R, L.M. and Konstantinidis, K.T., The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes, PeerJ Preprints, 2016, 4:e1900v1.
Slobodkina, G.B., Merkel, A.Y., Novikov, A.A., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., Pelomicrobium methylotrophicum gen. nov., sp. nov. a moderately thermophilic, facultatively anaerobic, lithoautotrophic and methylotrophic bacterium isolated from a terrestrial mud volcano., Extremophiles, 2020, vol. 24, pp. 177–185. https://doi.org/10.1007/s00792-019-01145-0
Slobodkin, A.I., Reysenbach, A.-L., Slobodkina, G.B., Baslerov, R.V., Kostrikina, N.A., Wagner, I.D., and Bonch-Osmolovskaya, E.A., Thermosulfurimonas dismutans gen. nov., sp. nov. a novel extremely thermophilic sulfur-disproportionating bacterium from a deep-sea hydrothermal vent, Int. J. Syst. Evol. Microbiol., 2012, vol. 62, pp. 2565–2571.
Sorokin, D.Y., Detkova, E.N., and Muyzer, G., Propionate and butyrate dependent bacterial sulfate reduction at extremely haloalkaline conditions and description of Desulfobotulus alkaliphilus sp. nov., Extremophiles, 2010, vol. 14, pp. 71–77.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G., The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools, Nucleic Acids Res., 1997, vol. 25, pp. 4876–4882.
Trüper, H.G. and Schlegel, H.G., Sulphur metabolism in Thiorhodaceae I. Quantitative measurements on growing cells of Chromatium okenii, Antonie van Leeuwenhoek, 1964, vol. 30, pp. 225–238. https://doi.org/10.1007/BF02046728
Tu, T.-H., Wu, L.-W., Lin, Y.-S., Imachi, H., Lin, L.-H., and Wang, P.-L., Microbial community composition and functional capacity in a terrestrial ferruginous, sulfate-depleted mud volcano, Front. Microbiol., 2017, vol. 8, p. 2137.
Weisburg, W.G., Barns, S.M., Pelletier, D.A., and Lane, D.J., 16S ribosomal DNA amplification for phylogenetic study, J. Bacteriol., 1991, vol. 173, pp. 697–703.
Wolin, E.A., Wolin, M., and Wolfe, R.S., Formation of methane, J. Franklin Inst., 1963, vol. 176, p. 737.
Yoon, S.H., Ha, S.-M., Lim, J., Kwon, S., and Chun, J., A large-scale evaluation of algorithms to calculate average nucleotide identity, Antonie van Leeuwenhoek, 2017, vol. 110, pp. 1281–1286
Zhang, L., Qiu, Y.Y., Zhou, Y., Chen, G.H., van Loosdrecht, M.C.M., and Jiang, F., Elemental sulfur as electron donor and/or acceptor: mechanisms, applications and perspectives for biological water and wastewater treatment, Water Research, 2021, vol. 202, р. 117373. https://doi.org/10.1016/j.watres.2021.117373
Funding
The work was supported by the Russian Science Foundation, project no. 22-14-00011 (isolation, determination of the physiological characteristics and taxonomic position of the isolate). Full genome sequencing and bioinformatic analysis were supported by the RF Ministry of Science and Higher Education. Fatty acid composition was determined at the Gubkin National University of Oil and Gas and was supported by a scholarship of the President of the Russian Federation (SP-4709.2022.1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of intere-st.
This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Frolova, A.A., Merkel, A.Y., Kuchierskaya, A.A. et al. Desulfobotulus pelophilus sp. nov., an Alkaliphilic Sulfate-Reducing Bacterium from a Terrestrial Mud Volcano. Microbiology 92, 493–499 (2023). https://doi.org/10.1134/S0026261723600878
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261723600878