Abstract
A strain capable of efficient biodesulfurization of dibenzothiophene was isolated from oil-contaminated soil samples. The strain, designated BDS24, was a rod-shaped gram-negative bacterium. Molecular identification based on its 16S rDNA gene sequence showed that this strain belonged to the genus Klebsiella. Examining its ability to desulfurize dibenzothiophene by gas chromatography revealed that this isolate consumed 0.5 mM of dibenzothiophene within 72 h. Evaluation of growth characteristics by the tetrazolium chloride assay showed that BDS24 isolate reached its maximum growth in the exponential phase after 60 h of growth with DBT. Compared to the reported 4S metabolic pathway, an extended 4S pathway was proposed for the desulfurization of dibenzothiophene, which has not been formerly reported in the literature. A decrease of 88% of the total sulfur content of crude oil sample by a Klebsiella strain has not been achieved previously. These results indicate that Klebsiella sp. BDS24 is an adequate candidate for deep biodesulfurization of crude oil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Sulfur is the third most abundant element in crude oil, and more than 70% of the sulfur in crude oil is in the form of dibenzothiophene (DBT) and its derivatives (Aminsefat et al., 2012; Delegan et al., 2021). Combustion of fossil fuels releases large amounts of sulfur oxides into the atmosphere leading to dangerous environmental hazards such as acid rains (Olmo et al., 2005; Akhtar et al., 2009). Therefore, refineries must decrease the sulfur content of the fuels they produce. The high water content of crude oil makes biodesulfurization (BDS) of crude oil more feasible than that of gasoline and diesel oil. However, few results on biodesulfurization of crude oil have been published (Monticello and Finnerty, 1985; Gupta et al., 2005; Yu et al., 2006). While hydrogen desulfurization (HDS) is a technique ordinarily employed in oil refining companies, requires extreme pressure and temperature conditions. Some refractory heterocyclic sulfur compounds, such as dibenzothiophene, are not eliminated by this method (Park et al., 2011; Omar et al., 2020).
Biodesulfurization (BDS) has been proposed as an economically viable and environmentally safe alternative to desulfurization of difficult-to-decomposable sulfur compounds, which works in mild operating conditions and uses whole-cell biocatalysts instead of expensive enzymes (Canales et al., 2018; Nassar et al., 2020). In various researches, dibenzothiophene has been studied as a model compound for biodesulfurization (Li et al., 2019). Rhodococcus erythropolis IGTS8 is the first microorganism known to possess BDS activity and has been extensively studied (Gray et al., 1996, 2003; Parveen et al., 2020). Other desulfurizing bacteria belong to the genera Corynebacterium, Gordonia, Nocardia, Mycobacterium, Paenibacillus, Shewanella, Sphingomonas, Halothiobacillus, Bacillus, etc. (Akhtar et al., 2016a, 2016b, 2018; Nazari et al., 2017; Sadare et al., 2017). The discovery of the aerobic 4S pathway was a turning point for biodesulfurization, as it is a non-destructive pathway in which the calorific value of dibenzothiophene is preserved (Denome et al., 1993; Martínez et al., 2017). The 4S pathway contains four consecutive enzymatic reactions catalyzed by three enzymes encoded by the dszABC genes. DBT monooxygenase (DszC) catalyzes the oxidation of DBT to DBT-sulfoxide and then to DBT-sulfone. DBT-sulfone monooxygenase (DszA) catalyzes the oxidation of sulfone to 2-hydroxybiphenyl-2-sulfinate (HBPS), and HBPS sulfinolyase (DszB) catalyzes the conversion of HBPS to 2-hydroxybiphenyl (2-HBP). NADH-FMN oxidoreductase (DszD) also provides the FMNH2 cofactor regeneration required for reactions catalyzed by DszC and DszA (Kodama, 1977; Oldfield et al., 1997; Díaz and García, 2010; Sousa et al., 2020). Nowadays, biodegradation of polycyclic aromatic hydrocarbons (PAHs) are the focus of bioremediation of oil-contaminated soils and seawater, rather than polycyclic aromatic sulfur heterocycles (PASHs). Whereas, complete removal of PASHs without releasing secondary contamination to the environment is mandatory (Li et al., 2019).
In the present study, we report a new DBT-desulfurizing bacterial strain, Klebsiella sp. BDS24, isolated from soil at the Abadan Oil Refining Company, Iran. Its DBT desulfurization ability, morphological analysis, and growth rate under aerobic conditions were assessed. The metabolites produced via DBT desulfurization by the isolate and the extended metabolic desulfurization pathway were explicated via GC-MS. The decrease of sulfur content in crude oil was also studied.
MATERIALS AND METHODS
Chemicals. DBT (98% purity) and absolute ethanol were purchased from MERCK, Germany. A bacterial Genomic DNA extraction kit was purchased from FAVORGEN, Taiwan. Crude oil samples were procured from the Abadan oil refining company of Iran. All the other chemicals were of the analytical grade and commercially available, and were used without further purification.
Media. The basal salt medium (BSM) was prepared as recommended previously (Izumi et al., 1994; Davoodi-Dehaghani et al., 2010), with some modifications. This culture medium contained the following compounds (g/L distilled water): glycerol, 5; KH2PO4, 2; K2HPO4, 4; NH4Cl, 1; MgCl2·6H2O, 0.2; CaCl2, 0.02; and 10 mL of metal solution (pH 7.2). The metal solution contained (g/L distilled water): FeCl3·6H2O, 0.001; ZnCl2, 0.5; MnCl2·4H2O, 0.5; Na2MoO4·2H2O, 0.1; and CuCl2, 0.05. While containing the necessary growth elements and sources of carbon and nitrogen, this medium does not have a source of sulfur. By adding 0.54 mM of dibenzothiophene as the only source of sulfur, it becomes a selective culture medium for desulfurizing microorganisms. BSM-agar was prepared by adding 1.5% agar.
Enrichment and isolation of DBT-desulfurizing bacteria. From the designated sampling sites in the Abadan oil refining company (Khuzestan province, Iran), 18 samples of oil-contaminated soils were collected. The purpose of enrichment was to obtain microorganisms with high desulfurization capacity. In this method, 2 g of each sample was poured into 100 mL Erlenmeyer flasks containing 20 mL of sterile BSM medium supplemented with dibenzothiophene; then the flasks were incubated on a rotary shaker (150 rpm) at 30°C. After five days, for subcultivation, 2 mL of these primary cultures were added to 20 mL of fresh BSM medium supplemented with DBT and incubated under the same conditions for another 5 days. After five subcultivations, the cultures were prepared in 10-fold dilutions from 10–1 to 10–5. The suspensions (100 μL) were spread on BSM agar supplemented with DBT. These plates were incubated at 30°C for three days, and after the growth of microorganisms, each distinctive colony was purified separately onto a BSM-agar medium supplemented with DBT. A total of 25 isolates capable of desulfurization were obtained. Based on its growth which indicated great desulfurization performance, strain BDS24 was chosen for subsequent evaluation.
Identification of the DBT-desulfurizing bacterium. For molecular identification of the selected isolate, template DNA was extracted from fresh bacterial culture using a bacterial DNA extraction kit according to the manufacturer’s instructions. The 16S rDNA genes were amplified with the universal primers 27F and 1492R using a Mastercycler ep Gradient S (Eppendorf, Germany). PCR was performed according to the following procedure: denaturation at 94°C for 3 min, 35 cycles of 30 s at 94°C, 30 s at 54°C, and 2 min at 72°C, and a final extension step at 72°C for 5 min.
To confirm PCR by electrophoresis, PCR products and DNA Ladder were investigated on a 0.5% agarose gel in a 1× TAE buffer and stained with Safe stain for visualizing DNA bands using a UVITEC Cambridge Gel Documentation System. The PCR product with a size of about 1500 bp was sent to Microsynth Co. in Switzerland for sequencing. The resulting sequence was submitted into the Blast search of the NCBI database, and species whose 16S rDNA gene had a high degree of similarity with our sequence were introduced. The phylogenetic tree was constructed using MEGA 11 software.
Investigation of bacterial growth and desulfurization of DBT by BDS24 strain. To investigate the decrease of DBT and identify the metabolites produced during the desulfurization process by gas chromatography-mass spectrometry (GC-MS), a bacterial suspension with OD600 = 2.0 was prepared in the Ringer solution, and 200 μL of this suspension was inoculated to each Erlenmeyer flask with 20 mL of the medium. The flasks were placed on a rotary shaker (150 rpm) at 30°C. The cultures were removed from the shaker after 24, 44, 64, and 72 h, respectively.
Due to the nature of growth and biofilm formation by the isolate in the medium with DBT, the growth rate could not be measured by spectrophotometry. Owing to the presence of grains, the resulting culture was not homogeneous causing errors in the test results. Therefore, growth was measured by the triphenyl tetrazolium chloride (TTC) colorimetric assay (Hayashi et al., 2003; McCluskey et al., 2005; Sabaeifard et al., 2014). After optimizing the TTC assay conditions for this isolate using the Taguchi method (data not shown), two BSM media with DBT were prepared in 100-mL volumes. These media were inoculated from a 48-h culture of strain BDS24. The flasks’s were placed on a rotary shaker (150 rpm) at 30°C, and samples were collected every 6 h. For the TTC test, the samples (2.5 mL) were transferred to sterile test tubeswrapped in aluminum foil to prevent light access. The TTC stock solution (0.1%) was added (250 μL), and the tubes were placed on a rotary shaker for 5 h. After this time, the liquid was discarded, and the resulting red formazan precipitate was dissolved in 2.5 mL of dimethyl sulfoxide. The absorbance (OD480) of the samples was then measured spectrophotometrically (T60U, PG Instruments LTD., Leicestershire, United Kingdom).
GC-MS-based analysis.To determine the degree of DBT decrease and identify the compounds resulting from its desulfurization, a 7890B GC/5977A MS instrument (Agilent Company, United States) was used, equipped with the split/splitless injection system and electron impact ionization model; it had NIST and WILEY mass libraries. To analyze the desired compounds, an HP5-MS capillary column with a length of 60 m, an inner diameter of 0.25 mm, and a film thickness of 0.25 µm was used. Injection site temperature, interface temperature, and ionization site temperature were 270, 299, and 230°C, respectively. The temperature program of the column started with the initial temperature of 50°C. It was kept at this temperature for 7 min, then the temperature of the column reached 299°C with a slope of 13°C/min, and for 10 min remained at this temperature. The split ratio was set to 1 to 10, and the injection volume was 1 µL.
Broth cultures were acidified to pH 2−3, and the extraction process was performed for 45 min on a magnetic stirrer at 450 rpm. The extracting solvent was the hexane/dichloromethane mixture (0.3/0.7 vol/vol).
Investigation of sulfur consumption from crude oil by strain BDS24.Nutrient broth medium was prepared in a volume of 100 mL and inoculated with a single colony of strain BDS24. The flask was incubated on a rotary shaker (150 rpm) at 30°C until the OD600 nm reached to 0.8. Then, cells were harvested by centrifugation at 2870 g for 15 min and washed with 100 mM phosphate buffer (pH 7.0).
Two BSM media supplemented with 2% crude oil were prepared in volumes of 100 mL. One of these flasks was inoculated with the bacterial suspension, and the other flask was considered as a control. Both flasks were incubated for 72 h in a rotary shaker (150 rpm) at 30°C.
Crude oil was extracted from the aqueous phase using normal hexane. Then the total sulfur of the crude oil present in the control sample and the sample treated with BDS24 isolate was measured according to the ASTM D4294 Standard Test Method for Sulfur in Petroleum and Petroleum Products by Energy Dispersive X-ray Fluorescence Spectrometry.
RESULTS AND DISCUSSION
Isolation and characterization of DBT-desulfurizing bacteria. Oil-contaminated soil samples were collected from different parts of the Abadan refinery. After 30 days of enrichment, a total of 28 isolates with the ability to grow in a desulfurization medium were purified. After five transfers of the BRS13 sample, strain BDS24 was isolated. In the course of consecutive transfers, the dominance of some colonies and elimination of other colonies was observed. The BDS24 isolate was selected for subsequent analyzes based on its growth rate and high desulfurization activity when grown on DBT. The BDS24 colonies on BSM medium with DBT were colorless, opaque, spherical, medium in size, with a smooth margin, a mucoid surface and a sticky form (Fig 1b). After Gram staining, the cells of this isolate appeared as relatively short gram-negative rods. The colonies of this isolate on EMB agar medium were pink, spherical, large, and have a mucoid form (Fig. 1c). The BDS24 cells were rod-shaped and approximately 1.5 μm in length (Fig. 1d).
Based on the results of the 16S rDNA gene sequencing of the isolate BDS24 (1417 bp), this strain belonged to the Klebsiella sp. The sequence was deposited in GenBank under accession number OL823066 (https://www.ncbi. nlm.nih.gov/nucleotide/OL823066.1). The phylogenic tree of strain BDS24 and closely related bacteria constructed by the neighbor-joining method using the 16S rDNA gene sequences is shown on Fig. 2. The BDS24 strain was closely related to Klebsiella variicola CCFM8387 (KJ803944.1) (Long et al., 2022), with a 99% sequence identity. In the study of Bhatia and Sharma (2012), a thermophilic strain of Klebsiella sp. 13T was isolated by enrichment method from contaminated soil samples collected from an oil refinery. This Klebsiella sp. strain was isolated from an environmental sample and we expect it to be different from clinical isolates in terms of pathogenicity, so that it can be used industrially. Of course, this claim requires additional identification and investigation of pathogenic genes in this strain.
Evaluation of growth and efficiency of the BDS24 isolate in desulfurization of DBT. Growth and cell viability analysis of strain BDS24 was assessed by TTC assay, during which sampling of cultures continued for 120 h. The results are shown as the growth curve graph on Fig. 3. In the first stage of growth, there was a lag phase. The highest growth rate of the isolate observed at 60 h when OD480 was 0.538, and then its growth decreased. It is worth mentioning that in 108 to 120 h, there was an increase in OD, comparable to the values for 18–24 h.
Cultures of strain BDS24 were analyzed by GC-MS at 24, 44, 64, and 72 h. DBT in absolute ethanol solution with a final concentration of 0.5 mM was added to BSM culture media. Figures 4a and 4b show the chromatograms of the control sample and of the sample inoculated with the isolate after 72 h of incubation. The decrease of DBT content and the shortening of its peak can be seen compared to the control sample. DBT was recorded at retention times of 18−19 min. The DBT concentrations at different time intervals, determined based on the decrease of its sub-peak area in the chromatograms, are shown on Fig. 5. The percentage of DBT desulfurization 24, 44, 64, and 72 h after inoculation was calculated based on the area under its peak in the samples treated with the BDS24 isolate and the control sample, which were 37, 55, 62, and 97.5%, respectively.
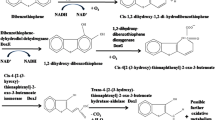
New proposed extended 4S pathway for DBT desulfurization. To determine the possible metabolic pathway for DBT desulfurization by the isolate BDS24, intermediates were extracted from different time intervals and analyzed by GC-MS.
Phthalic acid was found in this experiment at the retention time of 18.004 min. This metabolite was also detected during the desulfurization of DBT by Pseudomonas sp. LKY-5 (Li et al., 2019). 1,3-Dimethyl benzene was recorded at the retention time of 6.974 min, which had the highest abundance among metabolites. Phenol, 2-methyl-5-(1-methyl ethyl) was also detected at the retention time of 14.117 min.
Based on the reported metabolites and metabolic pathways of DBT, the metabolic pathway for desulfurizing of DBT by Klebsiella sp. BDS24 is proposed as shown in Fig. 6. In studies such as by El-Gendy et al., (2014) and Elmi et al. (2015), extended 4S pathways have been proposed, in which 2-hydroxy biphenyl, the end product of the 4S pathway, is broken down into other metabolites. During the metabolic pathway proposed in the present study, sulfur is released in the form of SO2 from dibenzothiophene. This is an advantage of biodesulfurization, as in hydrogen desulfurization sulfur is released from sulfur compounds in the form of H2S, which has higher toxicity. One of the advantages of this pathway for desulfurization is that it is not destructive; that is, the integrity of the carbon structure of DBT is not attacked by enzymes, and the end products such as toluene (RT 6.662 min), ethylbenzene (RT 8.125 min), and xylene (RT 7.077 min) are produced, which have a calorific value in crude oil.
Evaluation of crude oil sulfur content decrease by strain BDS24. Oil refineries commonly separate crude oil into several fractions and then desulfurize them separately. Refineries can make significant cost savings if most of the sulfur is removed from crude oil before it is fractionated (Yu et al., 2006). Therefore, the ability of strain BDS24 to remove sulfur compounds from crude oil was determined. After biphasic batch desulfurization of crude oil, the total sulfur content decreased from 0.265 to 0.030 mass % within the 72-h period. The percentage of crude oil sulfur content decrease compared to the control sample was calculated using the following formula:
In this formula, R is desulfurization efficiency, Si is total sulfur of crude oil present in the control sample, and Sf is total sulfur of the crude oil present in the desulfurized sample. According to this calculation, the BDS24 isolate can removed 88% of the sulfur content of crude oil present in the sample treated with this isolate for 72 h.
In the present study, a gram-negative bacterium, Klebsiella sp. BDS24, able to desulfurize DBT, was isolated. This bacterium is the first strain of Klebsiella with very high desulfurization efficiency, i.e., removal of 97.5% of DBT in 72 h, which was obtained for the first time in this study. An extended 4S pathway for desulfurization of DBT by this strain has been proposed, which has not ever been mentioned in the literature. The isolate was also able to decrease the sulfur content of crude oil by 88% without affecting its calorific value, which was one of its advantages. These results indicate that Klebsiella sp. BDS24 is an adequate candidate for biodesulfurization of crude oil.
REFERENCES
Akhtar, N., Ghauri, M. A., and Akhtar, K., Exploring coal biodesulfurization potential of a novel organic sulfur metabolizing Rhodococcus spp. (Eu-32)—a case study, Geomicrobiol. J., 2016a, vol. 33, no. 6, pp. 468−472. https://doi.org/10.1080/01490451.2015.1052119
Akhtar, N., Ghauri, M.A., and Akhtar, K., Dibenzothiophene desulfurization capability and evolutionary divergence of newly isolated bacteria, Arch. Microbiol., 2016b, vol. 198, no. 6, pp. 509–519. https://doi.org/10.1007/S00203-016-1209-5
Akhtar, N., Ghauri, M.A., Anwar, M.A., and Akhtar, K., Analysis of the dibenzothiophene metabolic pathway in a newly isolated Rhodococcus spp., FEMS Microbiol. Lett., 2009, vol. 301, no. 1, pp. 95–102. https://doi.org/10.1111/J.1574-6968.2009.01797.X
Akhtar, N., Akhtar, K., and Ghauri, M.A., Biodesulfurization of thiophenic compounds by a 2-hydroxybiphenyl-resistant Gordonia sp. HS126-4N carrying dszABC genes, Curr. Microbiol., 2018, vol. 75, pp. 597–603. https://doi.org/10.1007/s00284-017-1422-8
Aminsefat, A., Rasekh, B., and Ardakani, M.R., Biodesulfurization of dibenzothiophene by Gordonia sp. AHV-01 and optimization by using of response surface design procedure, Microbiology (Moscow), 2012, vol. 81, no. 2, pp. 154–159. https://doi.org/10.1134/S0026261712020026
Bhatia, S. and Sharma, D.K., Thermophilic desulfurization of dibenzothiophene and different petroleum oils by Klebsiella sp. 13T, Environ. Sci. Polluti. Res. Int., 2012, vol. 19, no. 8, pp. 3491–3497. https://doi.org/10.1007/S11356-012-0884-2
Canales, C., Eyzaguirre, J., Baeza, P., Aballay, P., and Ojeda, J., Kinetic analysis for biodesulfurization of dibenzothiophene using R. rhodochrous adsorbed on silica, Ecol. Chem. Eng. S, 2018, vol. 25, no. 4, pp. 549–556. https://doi.org/10.1515/ECES-2018-0036
Davoodi-Dehaghani, F., Vosoughi, M., and Ziaee, A.A., Biodesulfurization of dibenzothiophene by a newly isolated Rhodococcus erythropolis strain, Biores. Technol., 2010, vol. 101, no. 3, pp. 1102–1105. https://doi.org/10.1016/J.BIORTECH.2009.08.058
Delegan, Y., Kocharovskaya, Y., Frantsuzova, E., Streletskii, R., and Vetrova, A., Characterization and genomic analysis of Gordonia alkanivorans 135, a promising dibenzothiophene-degrading strain. Biotechnol. Rep., 2021, vol. 29. https://doi.org/10.1016/J.BTRE.2021.E00591
Denome, S.A., Olson, E.S., and Young, K.D. Identification and cloning of genes involved in specific desulfurization of dibenzothiophene by Rhodococcus sp. strain IGTS8, Appl. Environ. Microbiol., 1993, vol. 59, no. 9, pp. 2837–2843. https://doi.org/10.1128/AEM.59.9.2837-2843.1993
Díaz, E. and García, J.L., Genetics engineering for removal of sulfur and nitrogen from fuel heterocycles, in Handbook of Hydrocarbon and Lipid Microbiology, Berlin: Springer, 2010, pp. 2787–2801. https://doi.org/10.1007/978-3-540-77587-4_206
Long, D.-L., Wang, Y.-H., Wang, J.-L., Mu, S.-J., Chen, L., Shi, X.-Q., and Li, J.Q., Fatal community-acquired bloodstream infection caused by Klebsiella variicola: a case report, World J. Clin. Cases., 2022, vol. 10, no. 8, pp. 2474−2483. https://doi.org/10.12998/wjcc.v10.i8.2474
El-Gendy, N.S., Nassar, H.N., and Abu Amr, S.S., Factorial design and response surface optimization for enhancing a biodesulfurization process, Pet. Sci. Technol., 2014, vol. 32, no. 14, pp. 1669–1679. https://doi.org/10.1080/10916466.2014.892988
Elmi, F., Etemadifar, Z., and Emtiazi, G., A novel metabolite (1,3-benzenediol, 5-hexyl) production by Exophiala spinifera strain FM through dibenzothiophene desulfurization, World J. Microbiol. Biotechnol., 2015, vol. 31, no. 5, pp. 813–821. https://doi.org/10.1007/S11274-015-1835-0
Felsenstein, J., Confidence limits on phylogenies: an approach using the bootstrap, Evolution, 1985, vol. 39, no. 4, p. 783. https://doi.org/10.2307/2408678
Gray, K.A., Mrachkott, G.T., and Squires, C.H., Biodesulfurization of fossil fuels, Curr. Opin. Microbiol., 2003, vol. 6, no. 3, pp. 229–235. https://doi.org/10.1016/S1369-5274(03)00065-1
Gray, K.A., Pogrebinsky, O.S., Mrachko, G.T., Xi, L., Monticello, D.J., and Squires, C.H., Molecular mechanisms of biocatalytic desulfurization of fossil fuels, Nat. Biotechnol.,1996, vol. 14, no. 13, pp. 1705–1709. https://doi.org/10.1038/nbt1296-1705
Gupta, N., Roychoudhury, P.K., and Deb, J.K., Biotechnology of desulfurization of diesel: prospects and challenges, Appl. Microbiol. Biotechnol., 2005, vol. 66, no. 4, pp. 356–366. https://doi.org/10.1007/S00253-004-1755-7
Hayashi, S., Kobayashi, T., and Honda, H., Simple and rapid cell growth assay using tetrazolium violet coloring method for screening of organic solvent tolerant bacteria, J. Biosci. Bioeng., 2003, vol. 96, no. 4, pp. 360–363. https://doi.org/10.1016/S1389-1723(03)90137-X
Izumi, Y., Ohshiro, T., Ogino, H., Hine, Y., and Shimao, M., Selective desulfurization of dibenzothiophene by Rhodococcus erythropolis D-1, Appl. Environ. Microbiol., 1994, vol. 60, no. 1, pp. 223–226. https://doi.org/10.1128/AEM.60.1.223-226.1994
Kodama, K., Co-metabolism of dibenzothiophene by Pseudomonas jianii, Agric. Biol. Chem., 1977, vol. 41, no. 7, pp. 1305–1306.https://doi.org/10.1080/00021369.1977.10862668
Li, L., Shen, X., Zhao, C., Liu, Q., Liu, X., and Wu, Y., Biodegradation of dibenzothiophene by efficient Pseudomonas sp. LKY-5 with the production of a biosurfactant, Ecotoxicol. Environ. Saf., 2019, vol. 176, pp. 50–57. https://doi.org/10.1016/J.ECOENV.2019.03.070
Martínez, I., El-Said Mohamed, M., Santos, V.E., García, J.L., García-Ochoa, F., and Díaz, E., Metabolic and process engineering for biodesulfurization in Gram-negative bacteria, J. Biotechnol., 2017, vol. 262, pp. 47–55. https://doi.org/10.1016/J.JBIOTEC.2017.09.004
McCluskey, C., Quinn, J.P., and McGrath, J.W., An evaluation of three new-generation tetrazolium salts for the measurement of respiratory activity in activated sludge microorganisms, Microb. Ecol., 2005, vol. 49, no. 3, pp. 379–387. https://doi.org/10.1007/S00248-004-0012-Z
Monticello, D.J. and Finnerty, W.R., Microbial desulfurization of fossil fuels, Annu. Rev. Microbiol., 1985, vol. 39, pp. 371–389.https://doi.org/10.1146/ANNUREV.MI.39.100185.002103
Nassar, H.N., Abu Amr, S.S., and El-Gendy, N.S., Biodesulfurization of refractory sulfur compounds in petro-diesel by a novel hydrocarbon tolerable strain Paenibacillus glucanolyticus HN4, Environ. Sci. Pollut. Res., 2020, vol. 28, no. 7, pp. 8102–8116. https://doi.org/10.1007/S11356-020-11090-7
Nazari, F., Kefayati, M.E., and Raheb, J., Isolation, identification, and characterization of a novel chemolithoautotrophic bacterium with high potential in biodesulfurization of natural or industrial gasses and biogas, Energy Sources, 2017, vol. 39, no. 10, pp. 971–977. https://doi.org/10.1080/15567036.2016.1263255
Oldfield, C., Pogrebinsky, O., Simmonds, J., Olson, E.S., and Kulpa, C.F., Elucidation of the metabolic pathway for dibenzothiophene desulphurization by Rhodococcus sp. strain IGTS8, Microbiology, 1997, vol. 143, no. 9, pp. 2961–2973. https://doi.org/10.1099/00221287143-9-2961
Olmo, C.H.D., Santos, V.E., Alcon, A., and Garcia-Ochoa, F., Production of a Rhodococcus erythropolis IGTS8 biocatalyst for DBT biodesulfurization: influence of operational conditions, Biochem. Eng. J., 2005, vol. 3, no. 22, pp. 229–237. https://doi.org/10.1016/J.BEJ.2004.09.015
Omar, R.A., Verma, N., and Arora, P.K., Sequential desulfurization of thiol compounds containing liquid fuels: adsorption over Ni-doped carbon beads followed by biodegradation using environmentally isolated Bacillus zhangzhouensis, Fuel, 2020, vol. 277, pp. 118−208. https://doi.org/10.1016/J.FUEL.2020.118208
Park, T.H., Cychosz, K.A., Wong-Foy, A.G., Dailly, A., and Matzger, A.J., Gas and liquid phase adsorption in isostructural Cu3[biaryltricarboxylate]2 microporous coordination polymers, Chem. Commun., 2011, vol. 47, no. 5, pp. 1452–1454. https://doi.org/10.1039/C0CC03482G
Parveen, S., Akhtar, N., Ghauri, M.A., and Akhtar, K., Conventional genetic manipulation of desulfurizing bacteria and prospects of using CRISPR-Cas systems for enhanced desulfurization activity, Crit. Rev. Microbiol., 2020, vol. 46, no. 3, pp. 300–320. https://doi.org/10.1080/1040841X.2020.1772195
Sabaeifard, P., Abdi-Ali, A., Soudi, M.R., and Dinarvand, R., Optimization of tetrazolium salt assay for Pseudomonas aeruginosa biofilm using microtiter plate method, J. Microbiol. Methods, 2014, vol. 105, pp. 134–140. https://doi.org/10.1016/J.MIMET.2014.07.024
Sadare, O., Obazu, F., and Daramola, M.O., Biodesulfurization of petroleum distillates—current status, opportunities and future challenges, Environments, 2017, vol. 4, no. 4, p. 85. https://doi.org/10.3390/ENVIRONMENTS4040085
Sousa, J.P.M., Ferreira, P., Neves, R.P.P., Ramos, M.J., and Fernandes, P.A., The bacterial 4S pathway—an economical alternative for crude oil desulphurization that reduces CO2 emissions, Green Chem., 2020, vol. 22, no. 22, pp. 7604–7621. https://doi.org/10.1039/D0GC02055A
Yu, B., Xu, P., Shi, Q., and Ma, C., Deep desulfurization of diesel oil and crude oils by a newly isolated Rhodococcus erythropolis strain, Appl. Environ. Microbiol., 2006, vol. 72, no. 1, pp. 54–58. https://doi.org/10.1128/AEM.72.1.54-58.2006
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Emami, F., Zarrini, G. & Kadkhodaie, A. Utilization of Dibenzothiophene and Desulfurization of Crude Oil by a New Klebsiella sp. Strain BDS24 and the Proposed Metabolic Pathway. Microbiology 92, 920–928 (2023). https://doi.org/10.1134/S0026261723600167
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261723600167