Abstract—
Formation of a community of phosphate-accumulating microorganisms in a laboratory sequential batch reactor (SBR) ensuring alternated aerobic and anaerobic conditions during periodic removal and addition of the medium were investigated. The bioreactor removed 50% phosphorus from the incoming medium after 22 days from the start-up. Microscopy and X-ray microassay revealed the of cells of diverse morphology that contained phosphorus-enriched granules. High-throughput sequencing of the 16S rRNA gene fragments carried out on days 0, 8, 15, and 22 showed changes in the community composition and its decreasing diversity. On day 22, approximately twofold increase of the relative abundances of Bacteroidetes (up to 43% of the 16S rRNA gene sequences) and Proteobacteria of the classes alpha (up to 15%) and beta (up to 27%) was observed. While at the onset of the reactor operation, typical PAOs related to “Candidatus Accumulibacter” (class Betaproteobacteria) constituted 0.2% of the community, they were not detected on day 22. The most likely PAO candidates were beta-proteobacteria of the genus Dechloromonas, the share of which increased from 0.7 to 11% by the time of the highest phosphorus removal from the inflowing medium. The relative abundance of heterotrophs of the genus Zoogloea (family Rhodocyclaceae) increased from 0.1 to 11.5%.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Microorganisms of the group of phosphate-accumulating organisms (PAO) capable of intracellular accumulation of polyphosphates are responsible for phosphorus removal from wastewater. They develop in activated sludge under cyclic growth conditions, with alternated presence and absence of electron acceptors (primarily of oxygen) and therefore in the presence or absence of easily available organic compounds (Van Loosdrecht et al., 1997; Mino et al., 1998; Seviour et al., 2003; Wentzel et al., 2008). Periodic shifts of growth conditions determine the direction of most of the PAO metabolic pathways. Under anoxic conditions (without electron acceptors), PAO consume organic compounds, especially volatile fatty acids, and store them as intracellular polymers; this process is accompanied by degradation of intracellular polyphosphates and orthophosphate release from the cells. Under oxic conditions and/or in the presence of an alternative electron acceptor (nitrate or nitrite), PAO grow, consume orthophosphate, and synthesize intracellular polyphosphates using the energy produced by decomposition of intracellular carbon and energy sources accumulated under anoxic conditions. Thus, PAO are microorganisms with a cyclic type of metabolism (Dorofeev et al., 2019).
PAO do not form a monophyletic group, i.e., ability to accumulate phosphates has been revealed in diverse phylogenetic groups of microorganisms (both bacteria and archaea). Members of the candidate genus “Ca. Accumulibacter phosphatis” (family Rhodocyclacea, Betaproteobacteria) are presently considered as typical PAO (Hesselmann et al., 1999; Nguyen et al., 2012; Stokholm-Bjerregaard et al., 2017; Zeng et al., 2018; Qiu et al., 2019). These organisms use volatile fatty acids (acetate or propionate) as substrates. Depending on the electron acceptor, two phylogenetic groups are discerned within this phylotypes, “Ca. Accumulibacter phosphatis” types I and II, which in turn are subdivided into clades (clades IA to IE for type I and clades IIA to II I for type II) (Rubio-Rincon et al., 2017). PAO I use oxygen, nitrate and nitrite as electron acceptors, while PAO II use only oxygen (Figdore et al., 2018; Dasgupta et al., 2019). PAO include aerobic Tetrasphaera actionbacteria common in the wastewater treatment plant (Maszenan et al., 2000; Nielsen et al., 2019), as well as other prokaryotes, for which their role in biological phosphorus removal has not been reliably established: Thiothrix caldifontis (Rubio-Rincón et al., 2017), Microlunatus phosphovorus (Nakamura et al., 1995), Ca. “Accumulimonas” (Nguyen et al., 2012), a new group of organotrophic bacteria of the family Comamonadaceae (Ge et al., 2015), Dechloromonas (Ren et al., 2020), and Methanosarcina mazei (Paula et al., 2019).
Although PAO have been used for wastewater treatment for half a century and numerous publications dealt with this microbial group, the physiology of phosphate-accumulating bacteria has not been studied in detail, mainly because no one succeeded in isolation of pure cultures of PAO from the complex and diverse microbial communities of activated sludge, where bacteria are closely bound with the mucous matrix (Dorofeev et al., 2019). Our knowledge of PAO is based on metagenomic analysis of microbial consortia responsible for large-scale phosphate removal in water treatment facilities and of laboratory cultures of phosphate-accumulating organisms. Investigation of the metagenome of the “Ca. Accumulibacter phosphatis”-enriched sludge revealed PAO II to possess all genes required for dinitrogen and CO2 fixation (Flowers et al., 2013). Research on expression of the genes of the TCA cycle revealed considerable discrepancies in the aerobic and anaerobic metabolism of different “Ca. Accumulibacter phosphatis” populations (Wexler et al., 2009). PAO were shown to possess both the main pathways of carbon and phosphorus central metabolism and the specific ones associated with the spectrum of the used organic substrates and electron acceptors (Skennerton et al., 2015). Proteimic analysis revealed that in all studied “Ca. Accumulibacter phosphatis” anaerobic glycogen degradation was carried out via the Embden−Meyerhof−Parnas pathway (Wilmes et al., 2008).
PAO-enriched cultures are usually grown in SBR (Sequencing Batch Reactor) (Artan et al., 2005; Liu et al., 2020; Fan et al., 2020), which reproduce the thechnological conditions of large-scale wastewater treatment facilities. This technology has certain shortcomings. Thus, obtaining high-density cultures (several g/L) requires partial biomass retention in the bioreactor, which results in formation of floccules with the same complex multicomponent structure as the activated sludge. It is next to impossible to establish a PAO-dominated microbial community with the lowest number of components in such reactors. The authors consider development and improvement of alternative approaches to cyclic cultivation, which maintains the optimal conditions for PAO growth in a homogeneous culture and may be preferable to traditional SBR cultivation as an approach greatly facilitating PAO investigation. Homogeneous cultures are better adapted to physiological studies and isolation of pure cultures using traditional microbiological techniques.
In the present work, the classical feed-batch cultivation with stirring in a modified SBR bioreactor with alternating aerobic and anaerobic stages was used to minimize flocculation and to obtain a PAO-enriched homogeneous microbial community. The goal of the present work was to investigate the phosphate-accumulating community developing under these conditions and to describe its species composition.
MATERIALS AND METHODS
Bioreactor cultivation. The laboratory bioreactor for cyclic PAO cultivation was based on the BIOSTAT B bioreactor (Sartorius) with the working volume of 2 L, equipped with a stirrer and external cooling jacket. The reactor is schematically presented of Fig. 1.
Cyclic cultivation was achieved by alternating anaerobic conditions in the presence of acetate (an easily available source of carbon and energy) and aerobic conditions without acetate (which has been consumed during the anaerobic stage). Addition of fresh medium and removal of the culture were carried out using two peristaltic pumps. Aerobic and anaerobic conditions were established by bubbling the bioreactor with air or oxygen-free nitrogen using a gas supply system (Eltochpribor, Russia). Automatic monitoring of the pumps and the gas flows was carried out using a LOGО universal logical module (Siemens).
Each 6-h cycle of cultivation in the bioreactor consisted of several stages:
(1) establishing anaerobic conditions by bubbling nitrogen into the bioreactor for 5 min at 5 L/min (O2 concentration in the culture liquid decreased below 0.05 mg О2 /L), followed by addition of the medium (0.125 L). Total duration of this stage was 10 min;
(2) anaerobic stage, when the culture was mixed at 200 rpm. This stage continued for 170 min;
(3) aerobic stage, when the microbial community was aerated for 3 h.
After four cycles (24 h), 5 min before the end of the aerobic stage, air supply was terminated, and 0.5 L of the culture was removed from the bioreactor. The residual volume in the reactor was 1.5 L.
Thus, the average specific growth rate of the culture was 0.29 1/day, corresponding to the culture age of 3.4 days in the quasi-stationary state (generation time of 2.4 days).
To prevent microbial growth in the medium reservoir and the feeding pipes, they were washed daily with sterile tap water heated to 95°C.
The bioreactor was inoculated with activated sludge from an aeration basin of the Lyubertsy wastewater treatment plant (Moscow, Russia).
The medium composition adjusted according to the generalized experience of laboratory PAO cultivation (Onuki et al., 2002; Welles et al., 2017) contained the following (g/L tap water): CH3COONa (3H2O), 0.708; (NH4)2SO4, 0.046; КH2PO4, 0.109; yeast extract, 0.009; MgSO4 · 7H2O, 0.135. Acetate was used as the major carbon and energy source (Artan et al., 2005, Tchobanoglous et al., 2014).
Analytical techniques. The pH of the medium was measured using an Expert-001 pH meter-ion meter (Econix-Expert, Russia). During cultivation, pH was within the range of 8.5–8.7.
The cultivation temperature was maintained at 18–20°С using a Haake® WKL 26 thermostat (Thermo Fisher Scientific, United States).
Suspended matter was determined gravimetrically after filtration and drying of the sample (PNDF 14.1:2:4.254-09).
Phosphates were determined photometrically with ammonium molybdate both in the inflowing medium and in the effluent after the oxic and anoxic stages of cultivation (PNDF 14.1:2:4.248-07).
Dissolved oxygen was determined using an Oxi 197 meter (WTW, Germany).
Investigation of the cultures of phosphate-accumulating bacteria. Cell morphology was examined under an Olympus CX41 phase contrast microscope (Olympus, Japan).
Phosphate-accumulating bacteria were identified based on the presence of intracellular granules containing phosphorus compounds (as established by X-ray microanalysis).
The percentage of the cells containing various inclusions was determined using the average values for 80 microscope fields.
Electron microscopy of total cell preparations was carried out under a JEM 100 (JEOL, Japan) as described previously (Vasilyeva et al., 2006). The cells were fixed with 2.5% glutaraldehyde and postfixed with the OsO4 solution.
X-ray microanalysis was carried out on a JEM-1400 microscope (JEOL, Japan) equipped with an X-ray microanalyzer (Oxford Instruments, United Kingdom), at 80 keV and sample slope of 15°. The spectra were analyzed using AZtec (Oxford Instruments). This software package was also used for elemental charting of the samples.
The samples for X-ray microanalysis were prepared by applying native cells on Formvar-coated, carbon-covered copper grids.
Statistical processing was carried out using Microsoft Office Excel 2007.
Analyzed samples. Four activated sludge samples were collected in order to investigate the composition of the microbial consortium:
(1) activated sludge from an industrial bioreactor for OM, ammonium, and phosphorus removal, which implemented the Cape Town University technology (Tchobanoglous et al., 2014) based on alternation of one oxic and two anoxic zones rotary aeration tank. This sample was used to inoculate the laboratory bioreactor;
(2) activated sludge collected from a laboratory bioreactor on day 8 of its operation;
(3) activated sludge collected from a laboratory bioreactor on day 15 of its operation; and
(4) activated sludge collected from a laboratory bioreactor on day 22 of its operation.
DNA isolation for metagenomic analysis, amplification and sequencing of the 16S rRNA gene fragments. DNA was isolated using the DNeasy PowerSoil Kit (Qiagen, Germany) according to the manufacturer’s protocol. The variable V3−V4 region of the 16S rRNA genes was amplified using the universal primers 341F (CCTAYGGGDBGCWSCAG) and 806R (GGACTACNVGGGTHTCTAAT) (Frey et al., 2016). Obtained PCR fragments were used to prepare the sequencing libraries with the Nextera XT DNA Library Prep Kit (Illumina) according to the manufacturer’s protocol. Multiplexing was carried out with the Nextera XT Index Kit v2. The PCR fragments were sequences using Illumina MiSeq. At least 8000 16S rRNA gene fragments were obtained for each sample.
The reads from all samples were pooled together, and low-quality reads, singletons, and chimeras were excluded from analysis. The remaining reads were clustered into OTUs with identity of at least 97%. To determine the OTU shares in each sample, original reads (including low-quality and singleton ones) with at least 97% identity along the read length were superimposed over the representative OTU sequences using the usearch software package (Edgar, 2010). Taxonomic identification according th the 16S rRNA gene sequences was carried out using usearch and the Silva database.
RESULTS AND DISCUSSION
PAO cultivation. The method for PAO cultivation was developed, which was based on periodic (cyclic) changes of cultivation conditions. Suspended matter content in the bioreactor after inoculation with activated sludge was 3.43 g/L, pH was 8.5. During the first 15 days suspended matter content decreased to 0.2 g/L due to washout, and pH increased slightly (to 8.9). During this period the amount of coarse suspended particles decreased,and suspended matter became almost completely represented by microbial biomass. During days 15–22 both the concentration of microbial biomass and pH of the liquid phase did not change, indicating stable operation of the reactor. At the same time, the concentration of dissolved phosphates decreased gradually. The difference between phosphate concentrations in the medium and in the effluent was 4.9 mg/L on day 5, increased to 7.2 mg/L on day 8 and reached 11–12 mg/L on day 22, which corresponded to 50% phosphorus removal from the inflowing medium (Fig. 2).
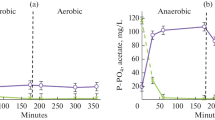
In order to determine the nature of phosphate removal, phosphate concentrations in the medium were measured during one cycle of operation (the oxic and anoxic phases). The data on phosphate concentrations during a cycle of cultivation on days 8, 15, and 22 are listed in Table 1. It should be noted that according to the biochemical reactions carried out by PAO, phosphate is released form the cells under anoxic conditions in the absence of oxidants and the presence of acetate. Under oxic conditions PAO oxidize the carbon substrates and consume phosphate from the medium (Terashgima et al., 2016).
Our results indicated that a microbial community containing physiologically active PAO developed in the laboratory bioreactor inoculated with activated sludge from a wastewater treatment plant. Importantly, phosphate concentration in the effluent was always lower than in the inflowing medium.
Investigation of PAO cells by microscopy and X-ray microanalysis. Microscopy of the bioreactor microbial community revealed morphologically diverse bacteria. Most of the cells (up to 85%) contained inclusions (Fig. 3). In general, elongated rounded cells (1–1.5) × (2–3.5) µm in size predominated, which contained refractive structures of diverse size located along the cell axis (Fig. 3a). Large rod-shaped cells, (2–3) × (3–5) µm in size, contained small dense dark inclusions (Fig. 3b). Apart from rod-shaped cells, coccoids 2.4 × 3.3 µm in size occurred, which contained bright round intracellular structures distributed uniformly throughout the cell volume (Fig. 3c). Electron microscopy confirmed occurrence of the cells with various inclusions (Figs. 3d, 3e).
Mapping of the microbial community using X-ray microanalysis, as well as pointwise analysis of the elemental composition of microbial cells, were used to determine the chemical composition of inclusions. X‑ray microanalysis provides for rapid determination of elemental composition of both individual cells and cell aggregates, making it possible to obtain information on the relative content of phosphorus and other elements of interest. The mapping mode implemented in the AZtec software package makes it possible to obtain a visual picture of elements distribution in the sample.
Mapping revealed different distribution of chemical elements, including phosphorus, among community members. Some elements were concentrated within the cells, while some were sorbed in the intercellular space on the cell surfaces. Carbon, potassium, sulfur, and calcium were uniformly distributed in the rod-shaped cells, (1−1.5) × (2−3.3) µm in size, while phosphorus, sodium, and magnesium occurred both in the cells and in the intercellular space (Fig. 4). Rod-shaped cells 0.5 × 2 µm in size were found to contain electron-dense structures, which were identified as phosphorus inclusions (Figs. 5a, 5b). To confirm the results of general mapping of the microbial community, pointwise elemental analysis of individual cells was carried out. The most characteristic spectra are shown on Fig. 5c. The percentage of chemical elements at the analyzed points is presented in Table 2. Pointwise analysis revealed higher phosphorus concentration in the sells (spectra 51–54) compared to the background level of this element. Phosphorus inclusions (spectra 51 and 52) in rod-shaped cells contained 10 times more phosphorus than the control. Inclusions in curved cells (spectrum 53) contained 8 times more of this element. The highest amount of phosphorus was accumulated in coccoid cells (spectrum 54).
Elemental composition of the microbial community determined by mapping with X-ray microanalysis. Location of individual chemical elements is marked by color. Electron microscopy (a) and distribution of carbon (b), phosphorus (c), sodium (d), potassium (e), magnesium (f), sulfur (g), and calcium (h). Accumulation of chemical elements in the intercellular space is indicated by arrows.
Pointwise analysis of the elemental composition of bacterial cells in the phosphate-accumulating community determined by mapping with X-ray microanalysis: electron micrograph of the cells (a); phosphorus distribution in the cells, with phosphor-rich granules indicated by arrows (b); and X-ray spectra of the control (background phosphorus distribution) and concentrations of the elements in the granules of rod-shaped bacteria (spectra 51 and 52), curved cells (spectrum 53), and coccoid cells (spectrum 54) (c). Phosphorus peaks are indicated by arrows.
Thus, phosphorus was shown to be present both on the cell surface and inside the cells. Intracellular distribution of this element varied. Some members of the microbial community contained phosphorus-enriched granules and probably belonged to PAO.
Changes in the taxonomic composition of the microbial consortium in the course of cultivation. Members of the Bacteria and Archaea domains were revealed in the microbial community at the initial stage of operation of the bioreactor. The share of archaea was low (1.17% of the total sequence number). Predominant archaeal sequences belonged to methanogens of the order Methanosarcinales and to the uncultured candidate phylum Wosearchaeota. The closest relatives of the revealed 16S rRNA gene sequences were found in the treatment facilities fermenting the sludge under anoxic conditions and in anaerobic digesters (Kirkegaard et al., 2017).
Bacterial sequences belonged to 19 phyla, with predominance of members of the phyla Bacteroidetes (22.4%), Proteobacteria (22.3%), Patescibacteria (8.8%), Chloroflexi (27.2%), and Nitrospirae (4.7%). They were responsible for 85.4% of the 16S rRNA gene sequences. The phyla Acidobacteria, Planctomycetes, Verrucomicrobia, etc. were the minor components of the community (Fig. 6).
About 4.8% of all the 16S rRNA gene sequences belonged to unknown deep lineages and were not identified even at the phylum level.
Most of identified phyla have been previously detected in wastewater treatment bioreactors (Wu et al., 2019).
In the course of cultivation of the community, its composition changed and its taxonomic diversity decreased. Thus, the share of archaea in the community decreased to 0.01% (Fig. 6). The shares of the phyla predominant at the bioreactor onset, Patescibacteria (8.8%), Chloroflexi (27.2%), and Nitrospirae (4.7%), decreased to 2.66, 2.63, and 0.15%, respectively. The phylum Patescibacteria has been described recently based on molecular data (Hug et al., 2016; Parks et al., 2018), and none of its members have been isolated in pure cultures. The metabolism of members of this phylum was predicted based on the metagenomic analysis of microbial communities. Patescibacteria usually have small genomes, with a number of metabolic pathways absent or incomplete. They are therefore considered possible parasites or symbionts.
Most of the minor groups were eliminated from the reactor during its operation,including members of the phyla Fusobacteria, Ca. Margulisbacteria, Ca. Latescibacteria, Spirochaetes, Elusimicrobia, Gemmatimonadetes, Armatimonadetes, and Omnitrophica. The shares of other minor groups decreased as well (Fig. 6).
The number of detected OTUs decreased from 445 to 184. At the final stage 61% of all reads belonged to 10 most abundant OTUs, which constituted only 1% of the community during the initial stage.
The shares of two phyla predominant in the inoculum, Bacteroidetes (22.4%) and Proteobacteria (22.3%), increased during cultivation under cyclic conditions. The percentage of Bacteroidetes, which are capable of degrading complex organic compounds, increased almost twofold, to 42.8% (Thomas et al., 2011; Fernández-Gómez et al., 2013; Hahnke et al., 2016). Members of the orders Chitinophagales, Cytophagales, and Sphingobacteriales were mostly responsible for this increase.
The share of proteobacteria increased from 22.3 to 43.2% due to increased abundance of members of two classes, Alpha- and Betaproteobacteria from 2.7 to 14.8% and from 10.7 to 27.3%, respectively, while the shares of Gamma-, Delta-, and Epsilonproteobacteria decreased from 9.5 to 1.4%. Among alphaproteobacteria, the share of microorganisms of the order Microvibrionales increased significantly (from 0.28 to 7.4%). Related 16S rRNA gene sequences have previously been detected in the biofilm of an anoxygenic hybrid reactor for nitrogen and phosphorus removal from wastewater (the sequence JN391746, with 98% identity to OTU1). Relative abundance of the genus Gemmobacter, comprising facultatively anaerobic heterotrophs (Rothe et al., 1987), also increased from 0.04 to 3.7%.
Betaproteobacteria, which predominated in most studied activated sludge microbial communities (Yu and Zhang, 2012; Wu et al., 2019), were represented in the bioreactor by the family Rhodocyclaceae. Relative abundance of the genera belonging to this family change significantly in the course of development of the community. Thus, the share of the genus Thauera decreased from 3.6 to 0.1% of the total reads number. These microorganisms are known to degrade a broad spectrum of organic acids and aromatic compounds (Mechichi et al., 2002; Mao et al., 2010). At the same time, abundance of the genus Zoogloea, which occurs in most of the activated sludge communities (Zhang et al., 2012; Wu et al., 2019), increased from 0.09 to 11.5%, as well as abundance of the genus Dechloromonas (family Rhodocyclaceae), from 0.74 to 11.1% of all 16S rRNA gene sequences. Related sequences have been detected in microbial communities of diverse plants, including an aerobic bioreactor in China (LT841763 and LT842295 with 99% identity) (Dasgupta et al., 2019). The family Rhodocyclaceae includes phosphate-accumulating organisms, including the species “Ca. Accumuliacter” phosphatis, which predominates in most microbial communities of industrial installations for phosphorus removal from wastewater (Bond et al., 1995; Mao et al., 2015; Barr et al., 2016; Dorofeev et al., 2019). As could be expected, Ca. Accumulibacter were identified in the bioreactor upon its inoculation with activated sludge. They were represented by two OTUs (0.01 and 0.2% of the total sequences number) with high similarity to the 16S rRNA gene sequences revealed in a phosphorus-removing SBR reactor (HM046420, 96% identity) and in wastewater treatment bioreactors (LR637422, 99% identity) (Bond et al., 1995; Mao et al., 2015). However, the share of these organisms decreased in the course of cultivation, so that they were not detected in the community by the end of the bioreactor operation. Thus, the PAO developing in the laboratory bioreactor during cyclic cultivation were not those typically found in large-scale facilities (Artan and Orhon, 2005).
In conclusion, it should be noted that sequential batch cultivation resulted in formation of a phosphate-accumulating community, differing from the community of flocculated activated sludge, which was used for inoculation in its higher homogeneity and low population density. Quantitative changes in phosphate concentration during the oxic and anoxic stages and the final decrease of phosphorus concentration in the medium indicated development of bacteria responsible for phosphorus removal in the processes similar to those occurring in the activated sludge of wastewater treatment facilities. The usual phosphorus concentration in wastewater is 4–8 mg/L (Qasim and Zhu, 2017) and is decreased to the concentrations below 1 mg/L after activated sludge treatment. The activated sludge concentration in industrial bioreactors is usually 2–4 g/L. In our experiments, the biomass concentration was an order of magnitude lower (0.2 g/L), while phosphorus removal was 12 mg/L, i.e., specific phosphorus removal per unit of activated sludge biomass was an order of magnitude higher. These results indicate the efficiency of sequential batch cultivation of activated sludge, which in a laboratory bioreactor resulted in development of a microbial community with high content of phosphate-accumulating bacteria.
Microscopic studies revealed phosphorus accumulation as granules in the cells of diverse morphology, which indicated the presence of diverse PAO in the community. Molecular biological techniques not only confirmed occurrence of phosphate-accumulating bacteria, but made it possible to monitor the dynamics of PAO community and to reveal the dominant members of this microbial group. The activated sludge used for inoculation was found to contain members of the family Rhodocyclaceae, Ca. Accumulibacter, which are considered typical inhabitants of waste treatment facilities. They were represented by two OTUs related to the 16S rRNA gene sequences previously revealed in an SBR reactor for phosphorus removal (HM046420, 96% identity) and in wastewater treatment bioreactors (LR637422, 99% identity) (Bond et al., 1995; Mao et al., 2015; Barr et al., 2016; Welles et al., 2017; Dorofeev et al., 2019). The relative abundance of these organisms decreased in the course of operation of the laboratory bioreactor. By the time of the most efficient phosphorus removal by the community, Candidatus “Accumulibacter phosphatis” was not detected. During this period the dominant groups were members of the family Rhodocyclaceae belonging to the genera Dechloromonas and Zoogloea; the shared of their 16S rRNA gene sequences increased to the greatest extent compared to those of other bacteria.
Members of the genus Zoogloea often occur in activated sludges (Barr et al., 2016; Wu et al., 2019). Some works reported accumulation of both Zoogloea and Dechloromonas in bioreactors for phosphorus removal under denitrifying conditions (Bond et al., 1995). Zoogloea were hypothesized to be potential PAO (Shao et al., 2009). Thus, intracellular volutin granules were found only in Zoogloea ramigera (Roinestad and Yall, 1970). Microorganisms related to Dechloromonas were detected in microbial communities of various waste treatment facilities. Thus, the 16S rRNA gene sequences most closely related to this genus were found in an aerobic bioreactor in China (LT841763 and LT842295, both 99% identity) (Bond et al., 1995; Mao et al., 2015; Welles et al., 2017; Dasgupta et al., 2019).
Thus, the operation mode with cyclic sequential batch cultivation of the activated sludge microbial community was unfavorable for development of Candidatus “Accumulibacter phosphatis” (family Rhodocyclaceae), which are common in waste treatment plants, and favored development of other members of this physiological group. Members of two genera of this family (Dechloromonas and Zoogloea) efficiently removed phosphorus from the medium.
REFERENCES
Artan, N. and Orhon, D., Mechanism and design of sequencing batch reactors for nutrient removal, London: IWA, 2005, vol. 19, p. 116.
Barr, J.J., Dutilh, B.E., Skennerton, C.T., Fukushima, T., Hastie, M.L., Gorman, J.J., Tyson, G.W., and Bond, P.L., Metagenomic and metaproteomic analyses of Accumulibacter phosphatis enriched floccular and granular biofilm, Environ. Microbiol., 2016, vol. 18, pp. 273–287.
Bond, P.L., Hugenholtz, P., Keller, J., and Blackall, L.L., Bacterial community structures of phosphate-removing and nonphosphate-removing activated sludges from sequencing batch reactors, Appl. Environ. Microbiol., 1995, vol. 61, pp. 1910–1916.
Dasgupta, S., De Clippeleir, H., and Goel, R., Short operational differences support granulation in a lab scale reactor in comparison to another conventional activated sludge reactor, Bioresource Technol., 2019, vol. 271, pp. 417–426.
Dorofeev A.G., Nikolaev, Yu.A., Mardanov, A.V., and Pimenov, N.V.,Cyclic metabolism as a mode of microbial existence, Microbiology (Moscow), 2019, vol. 88, no. 4, pp. 402–415.
Edgar R.C., Search and clustering orders of magnitude faster than BLAST, Bioinformatics, 2010, vol. 26, no. 19, pp. 2460–2461.
Fan, Z., Zeng, W., Wang, B., Guo, Y., Meng, Q., and Peng, Y., Transcriptional responses of “Candidatus Accumulibacter” clades to environmental dynamics in enhanced biological phosphorus removal, Bioresource Technol., 2020, p. 123108.
Fernández-Gómez, B., Richter, M., Schüler M., Pinhassi, J., Acinas, S.G., González, J.M., and Pedrós-Alió, C., Ecology of marine Bacteroidetes: a comparative genomics approach, ISME J., 2013, vol. 7, pp. 1026–1037.
Figdore, B.A., Stensel, H.D., and Winkler, M.K.H., Comparison of different aerobic granular sludge types for activated sludge nitrification bioaugmentation potential, Bioresource Technol., 2018, vol. 251, pp. 2189–2196.
Flowers, J.J., He, S., Malfatti, S., del Rio, T.G., Tringe, S.G., Hugenholtz, P., and McMahon, K.D., Comparative genomics of two ‘Candidatus Accumulibacter’ clades performing biological phosphorus removal, The ISME J., 2013, vol. 7, no. 12, pp. 2301–2314.
Frey, B., Rime, T., Phillips, M., Stierli, B., Hajdas, I., Widmer, F., and Hartmann, M., Microbial diversity in European alpine permafrost and active layers, FEMS Microbiol. Ecol., 2016, vol. 92, fiw018. https://doi.org/10.1093/femsec/fiw018
Hahnke, R.L., Meier-Kolthoff, J.P., García-López, M., Mukherjee, S., Huntemann, M., Ivanova, N.N., Woyke, T., Kyrpides, N.C., Klenk, H.P., and Göker, M., Genome-based taxonomic classification of Bacteroidetes, Front. Microbiol., 2016, vol. 20, p. 2003. https://doi.org/10.3389/fmicb.2016.02003
Hesselmann, R.P., Werlen, C., Hahn, D., van der Meer, J.R., and Zehnder, A.J.B., Enrichment, phylogenetic analysis and detection of a bacterium that performs enhanced biological phosphate removal in activated sludge, Syst. Appl. Microbiol., 1999, vol. 22, pp. 454–465.
Hug, L.A., Baker, B.J., Anantharaman, K., Brown, C.T., Probst, A.J., Castelle, C.J., Butterfield, C.N., Hernsdorf, A.W., Amano, Y., Ise, K., Suzuki, Y., Dudek, N., Relman, D.A., Finstad, K.M., Amundson, R., et al., A new view of the tree of life, Nat. Microbiol., 2016, vol. 1, p. 16048. https://doi.org/10.1038/nmicrobiol.2016.48
Kirkegaard, R.H., McIlroy, S.J., Kristensen, J.M., Nierychlo, M., Karst, S.M., Dueholm, M.S., Albertsen, M., and Nielsen, P.H., The impact of immigration on microbial community composition in full-scale anaerobic digesters, Sci. Rep., 2017, vol. 7, p. 9343. https://doi.org/10.1038/s41598-017-09303-0
Liu, S., Daigger, G.T., Liu, B., Zhao, W., and Liu, J., Enhanced performance of simultaneous carbon, nitrogen and phosphorus removal from municipal wastewater in an anaerobic-aerobic-anoxic sequencing batch reactor (AOA-SBR) system by alternating the cycle times, Bioresource Technol., 2020, vol. 301, p. 122750. https://doi.org/10.1016/j.biortech.2020.122750
Mao, Y., Graham, D.W., Tamaki, H., and Zhang, T., Dominant and novel clades of Candidatus “Accumulibacter phosphatis” in 18 globally distributed full-scale wastewater treatment plants, Sci. Rep., 2015, vol. 5, article 11857.
Mao, Y., Zhang, X., Xia, X., Zhong, H., and Zhao, L., Versatile aromatic compound-degrading capacity and microdiversity of Thauera strains isolated from a coking wastewater treatment bioreactor, J. Ind. Microbiol. Biotechnol., 2010, vol. 37, pp. 927–934.
Maszenan, A.M., Seviour, R.J., Patel, B.K.C., Schu-mann, P., Burghardt, J., Tokiwa, Y., and Stratton, H.M., Three isolates of novel polyphosphate-accumulating Gram-positive cocci, obtained from activated sludge, belong to a new genus, Tetrasphaera gen. nov., and description of two new species, Tetrasphaera japonica sp. nov. and Tetrasphaera australiensis sp. nov. Int. J. Syst. Evol. Microbiol., 2000, vol. 50, pp. 593–603.
Mechichi, T., Stackebrandt, E., Gad’on, N., and Fuchs, G., Phylogenetic and metabolic diversity of bacteria degrading aromatic compounds under denitrifying conditions, and description of Thauera phenylacetica sp. nov., Thauera aminoaromatica sp. nov., and Azoarcus buckelii sp. nov., Arch. Microbiol., 2002, vol. 178, pp. 26–35.
Mino, T., Van Loosdrecht, M.C.V, and Heijnen, J.J., Microbiology and biochemistry of the enhanced biological phosphate removal process, Water Res., 1998, vol. 32, no. 11, pp. 3193–3207.
Nakamura, K., Hiraishi, A., Yoshimi, Y., Kawaharasa-ki, M., Masuda, K., and Kamagata, Y., Microlunatus phosphovorus gen. nov., sp. nov., a new gram-positive polyphosphate-accumulating bacterium isolated from activated sludge, Int. J. Syst. Bacteriol., 1995, vol. 45, pp. 17–22.
Nguyen, H.T., Nielsen, J.L., and Nielsen, P.H., “Candidatus Halomonas phosphatis,” a novel polyphosphate-accumulating organism in full-scale enhanced biological phosphorus removal plants, Environ. Microbiol., 2012, vol. 14, no. 10, pp. 2826–2837.
Nielsen, P.H., McIlroy, S.J., Albertsen, M., and Nierychlo, M., Re-evaluating the microbiology of the enhanced biological phosphorus removal process, Curr. Opin. Biotechnol., 2019, vol. 57, pp. 111–118.
Onuki, M., Satoh, H., and Mino, T., Analysis of microbial community that performs enhanced biological phosphorus removal in activated sludge fed with acetate, Water Sci. Technol., 2002, vol. 6, nos. 1–2, pp. 145–153.
Parks, D.H., Chuvochina, M., Waite, D.W., Rinke, C., Skarshewksi, A., Chaumeil, P.-A., and Hugenholtz, P., A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life, Nat. Biotechnol., 2018, vol. 36, pp. 996–1004.
Paula, F.S., Chin, J.P., Schnürer, A., Müller, B., Manesiotis, P., Waters, N., Macintosh, K.A., Quinn, J.P., Co-nnolly, J., Abram, F., McGrath, J.W., and O’Flaherty, V., The potential for polyphosphate metabolism in Archaea and anaerobic polyphosphate formation in Methanosarcina mazei, Sci. Rep., 2019, vol. 9, no. 1, pp. 1–12.
PND F (Nature Conservation Standard Documents) 14.1:2:4.248-07, Method for measuring mass concentrations of orthophosphates, polyphosphates, and total phosphorusin samples of drinking, natural, and wastewater by the photometric method, Moscow, 2016. https://files.stroyinf.ru/Data2/1/4293773/4293773265.htm
PND F (Nature Conservation Standard Documents) 14.1:2:4.254-09, Quantitative chemical analysis of water. Methods for measuring mass concentrations of suspended and calcined weighed materials in samples of drinking, natural, and wastewater by the gravimetric method, Moscow, 2017. http://docs.cntd.ru/document/556339176
Qasim, S.R. and Zhu, G., Wastewater Treatment and Reuse, Theory and Design Examples, vol. 1: Principles and Basic Treatment, CRC, 2017.
Qiu, G., Zuniga-Montanez, R., Law, Y., Thi, S.S., Nguyen, T.Q.N., Eganathan, K., Liu, X., Nielsen, P.H., Williams, R.B.H., and Wuertz, S., Polyphosphate-accumulating organisms in full-scale tropical wastewater treatment plants use diverse carbon sources, Water Res., 2019, vol. 149, pp. 496–510.
Ren, S., Li, X., Yin, X., Luo, C., and Liu, F., Characteristics of intracellular polyphosphate granules and phosphorus-absorption of a marine polyphosphate-accumulating bacterium, Halomonas sp. YSR-3, J. Oceanol. Limnol., 2020, vol. 38, no. 1, pp. 195–203.
Roinestad, F.A. and Yall, I., Volutin granules in Zoogloea ramigera, Appl. Microbiol., 1970, vol. 19, no. 6, pp. 973–979.
Rothe, B., Fischer, A., Hirsch, P., Sittig, M., and Stackebrandt, E., The phylogenetic position of the budding bacteria Blastobacter aggregatus and Gemmobacter aquatilis gen. nov., sp. nov., Arch, Microbiol., 1987, vol. 147, pp. 92–99.
Rubio-Rincon, F.J., Lopez-Vazquez, C.M., Welles, L., van Loosdrecht, M.C.M., and Brdjanovic, D., Cooperation between Candidatus Competibacter and Candidatus Accumulibacter clade I, in denitrification and phosphate removal processes, Water Res., 2017, vol. 120, pp. 156–164.
Rubio-Rincón, F.J., Welles, L., Lopez-Vazquez, C.M., Nierychlo, M., Abbas, B., Geleijnse, M., Nielsen, P.H., van Loosdrecht, M.C.M., and Brdjanovic, D., Long-term effects of sulphide on the enhanced biological removal of phosphorus: the symbiotic role of Thiothrix caldifontis, Water Res., 2017, vol. 116, pp. 53–64.
Seviour, R.J., Mino, T., and Onuki M., The microbiology of biological phosphorus removal in activated sludge systems, FEMS Microbiol. Rev., 2003, vol. 27, pp. 99–127.
Shao, Y., Chung, B.S., Lee, S.S., Park, W., Lee, S.S., and Jeon, C.O., Zoogloea caeni sp. nov., a floc-forming bacterium isolated from activated sludge, Int. J. Syst. Evol. Microbiol., 2009, vol. 59, no. 3, pp. 526–530.
Skennerton, C.T., Barr, J.J., Slater, F.R., Bond, P.L., and Tyson G.W., Expanding our view of genomic diversity in Candidatus Accumulibacter clades, Environ. Microbiol., 2015, vol. 17, no. 5, pp. 1574–1585.
Stokholm-Bjerregaard, M., McIlroy, S.J., Nierychlo, M., Karst, S.M., Albertsen, M., and Nielsen, P.H., A critical assessment of the microorganisms proposed to be important to enhanced biological phosphorus removal in full-scale wastewater treatment systems, Front. Microbiol., 2017, vol. 8, p. 718. https://doi.org/10.3389/fmicb.2017.00718
Tchobanoglous, G., Burton F.L., and Stensel, H.D., Wastewater Engineering: Treatment and Reuse, McGraw Hill, N.Y.: Metcalf and Eddy, 2014.
Terashima, M., Yama, A., Sato, M., Yumoto, I., Kamagata, Y., and Kato, S., Culture-dependent and -independent identification of polyphosphate-accumulating Dechloromonas spp. predominating in a full-scale oxidation ditch wastewater treatment plant, Microbes Environ., 2016, vol. 31, no. 4, pp. 449–455.
Thomas, F., Hehemann, J.-H., Rebuffet, E., Czjzek, M., and Michel, G., Environmental and gut Bacteroidetes: the food connection, Front. Microbiol., 2011, vol. 2, article 93.
Van Loosdrecht, M.C.M., Hooijmans, C.M., Brdjanovic, D., and Heijnen, J.J., Biological phosphorus removal processes, Appl. Microbiol. Biotechnol., 1997, vol. 48, pp. 289–296.
Vasilyeva, L.V., Omelchenko, M.V., Berestovskaya, Y.Y., Lysenko, A.M., Abraham, W.R., Dedysh, S.N., and Zavarzin, G A., Asticcacaulis benevestitus sp. nov., a psychrotolerant, dimorphic, prosthecate bacterium from tundra wetland soil, Int. J. Syst. Evol. Microbiol., 2006, vol. 56, no. 9, pp. 2083–2088.
Welles, L., Abbas, B., Sorokin, D.Y., Lopez-Vazquez, C.M., Hooijmans, C.M., van Loosdrecht, M., and Brdjanovic, D., Metabolic response of “Candidatus Accumulibacter Phosphatis” clade II C to changes in influent P/C ratio, Front. Microbiol., 2017, vol. 7, p. 2121.
Wentzel, M.C., Comeau, Y., Ekama, G.A., van Loosdrecht, M.C.M., and Brdjanovic, D., Enhanced biological nutrient removal, in Biological Wastewater Treatment. Principles, Modelling and Design, Henze, M., van Loosdrecht, M.C.M., Ekama, G.A., and Brdjanovic, D., Eds., London: IWA, 2008, pp. 155–220.
Wexler, M., Richardson, D.J., and Bond, P.L., Radiolabelled proteomics to determine differential functioning of Accumulibacter during the anaerobic and aerobic phases of a bioreactor operating for enhanced biological phosphorus removal, Environ. Microbiol., 2009, vol. 11, no. 12, pp. 3029–3044.
Wilmes, P., Andersson, A.F., Lefsrud, M.G., Wexler, M., Shah, M., Zhang, B., Hettich, R.L., Bond, P.L., VerBerkmoes, N.C., and Banfield, J.F., Community proteogenomics highlights microbial strain-variant protein expression within activated sludge performing enhanced biological phosphorus removal, ISME J., 2008, vol. 2, pp. 853–864.
Wu, L., Ning, D., Zhang, B., Li, Y., Zhang, P., Shan, X., and Ling, F., Global diversity and biogeography of bacterial communities in wastewater treatment plants, Nature Microbiol., 2019, vol. 4, no. 7, pp. 1183–1195.
Yu, K. and Zhang, T., Metagenomic and metatranscriptomic analysis of microbial community structure and gene expression of activated sludge, PLoS One, 2012, vol. 7, no. 5, e38183.
Zeng, W., Zhang, L., Fan, P., Guo, J., and Peng, Y., Community structures and population dynamics of “Candidatus Accumulibacter” in activated sludges of wastewater treatment plants using ppk1 as phylogenetic marker, J. Environ. Sci., 2018, vol. 67, pp. 237–248.
Zhang, T., Shao, M.-F., and Ye, L., 454 pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants, ISME J., 2012, vol. 6, pp. 1137–1147.
Funding
The work was supported by the Russian Foundation for Basic Research, projects 18-29-25016 (setup development and assembly, morphological analysis of the microbial community, microscopy, chemical analysis, and molecular analysis of ther microbial community during the bioreactor operation) and 18-34-00627 (analysis of the composition of the activated sludge microbial community) and was also partially supported by the Russian Federation Ministry of Science and Higher Education.
Author information
Authors and Affiliations
Contributions
VAG and AGD assembled and maintained the laboratory setup for PAO cultivation; VVS and IKD carried out electron microscopy; YYB, AVP, YAN, and NVP carried out chemical analysis, microscopy, data analysis, and writing of the article; RYK, AVB, NVR, and AVB isolated metagenomic DNA, sequenced and analyzed the 16S rRNA gene sequences, and participated in the preparation of the manuscript.
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by P. Sigalevich
Rights and permissions
About this article
Cite this article
Pelevina, A.V., Berestovskaya, Y.Y., Grachev, V.A. et al. A Microbial Consortium Removing Phosphates under Conditions of Cyclic Aerobic-Anaerobic Cultivation. Microbiology 90, 66–77 (2021). https://doi.org/10.1134/S0026261721010082
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261721010082