Abstract—
Bacilli control behavioral reactions such as motility, biofilm formation, production of enzymes and metabolites, differentiation, and others by integrating a variety of environmental signals through a complex regulatory network. In the natural environment, Bacillus subtilis exists predominantly in the form of biofilms, which has made it an ideal model for studying the molecular strategy of biofilm formation. This paper systematizes information on the main regulatory systems responsible for the loss of mobility and the formation of B. subtilis biofilms, analyzes the behavior of bacteria within the biofilm population, leading to a state of bistability and differentiation into different types of subpopulations. It also evaluates the regulatory relationship between control systems responsible for the synthesis of structural components in the biofilm matrix. Particular emphasis is placed on data concerning signaling mechanisms that trigger the formation of a biofilm and its dispersion. In general, we summarize information about the latest discoveries in this area and their integration into the general idea of these complex microbial communities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
During the stationary growth phase, bacilli can display various adaptive responses associated with changes in the expression of survival-promoting genes. Under natural conditions, biofilms represent the predominant lifestyle; their formation under nutrient limitation is a complex and tightly regulated process. The biofilm-synthesizing capacity is a prerequisite for completion of the life-cycle of most microorganisms and it is a part of a successful strategy for protecting bacteria from detrimental environmental factors. Therefore, the molecular basis of biofilm formation has received increasing attention in microbiology. Presently, various bacterial models are used for elucidating the molecular mechanisms of biofilm formation and operation. The biofilms of the soil-dwelling bacterium Bacillus subtilis are envisaged as ideal models for this purpose. They are characterized by unique architectural features that result from carrying out complex programs of cell specialization and cell-cell communication within the community (Ryan-Payseur and Freitag, 2018; Kovacs and Dragos, 2019). This research is aimed at investigating the evolution, biological role, morphological traits, and structure of biofilms, as well as the molecular mechanisms underlying cell differentiation within a microbial community. The regulatory and metabolic relationship between biofilm formation in bacilli and other stationary phase-related processes in their cells, such as sporulation, motility, and secretion of secondary metabolites and proteins, including lipopeptides and extracellular enzymes, e.g., proteinases is of special interest (Aktuganov et al., 2019; Pisithkul et al., 2019). B. subtilis biofilms were shown to contain proteinase-producing cells, which increase in number during biofilm development (Kobayashi and Ikemoto, 2019). Moreover, the genes that encode biofilm formation are involved products are implicated, in regulatory terms, in the synthesis and secretion of various extracellular metabolites and signal molecules that enable cooperative interactions within a microbial community (Martin et al., 2020). Based on recent information, a platform for creating artificial biofilms by 3D printing for goal-directed practical use has been developed (Huang et al., 2018; Balasubramanian et al., 2019). We place special emphasis on summing up new data on the formation, development, and dispersion of B. subtilis biofilms.
The goal of this work was to systematically analyze data on the formation and dispersal of B. subtilis biofilms in terms of their interaction with other physiological processes during the stationary phase of this bacterium.
BIOFILM STRUCTURE
B. subtilis are motile gram-positive soil bacteria that can form stable biofilms, including those involved in symbiotic interaction with plant roots. Biofilm formation is one of the responses of B. subtilis cells to detrimental environmental factors; it results in the formation of a structured multicellullar bacterial community. In this community, cells adhere to one another and are embedded in the self-produced polymer matrix (Hobley et al., 2015). The matrix is made up of exopolysaccharides (EPSs), proteins, and nucleic acids; it contains channels used for transporting nutrients and oxygen and excreting bacterial metabolic products (Hobley et al., 2015). The matrix is responsible for biofilm stability and protection of matrix-embedded bacteria from bacteriophages, detrimental factors (osmotic shock, UV irradiation, dehydration, pH change, etc.) and antibacterial preparations (Strelkova et al., 2013; Dragos et al., 2017; Vidakovic et al., 2018; Pnomareva et al., 2018). A mechanism of “mutual aid” between matrix-forming and non-matrix-forming bacteria was described; non-matrix-forming bacteria gain a benefit from cooperative interaction with exopolysaccharide-synthesizing cells (Martin et al., 2020). Close contacts between bacterial cells via nanotubes enabling joint metabolite utilization and long-range signal transmission for orchestrating metabolic processes are aimed at creating a single integrated microbial community with rationally distributed extracellular “public goods.” Interaction among bacteria is based on the activation of complex intercellular communication processes. Of paramount importance for these processes are global metabolic regulatory facilities that involve quorum-sensing systems and cyclic di-GMP. As a result, the secretion of enzymes, secondary metabolites, siderophores, and antimicrobial compounds is activated (Bareia et al., 2018; Kalamara et al., 2018; Fu et al., 2019; Kovacs and Dragos, 2019). Biofilms enable metabolic cooperation between the cells in a community and promote symbiotic interactivity between bacteria and macroorganisms, e.g., between soil bacteria and plant roots (Townsley et al., 2018).
Natural biofilms exhibit a complex structure. They consist of a large number of microbial species that interact or compete for nutrients and habitats. Since natural biofilms represent mixtures of several microbial species, studies with them present difficulties. Therefore, research is often conducted with controllable single-species biofilms in a laboratory setting. The most widely used technique for obtaining biofilms under laboratory conditions is based on the formation of microbial communities on submerged solid surfaces, which are then visualized by staining with crystal violet. Another technique involves biofilm formation at the air-liquid interphase (Vlamakis et al., 2013; Hollenbeck et al., 2016). Instead of using plates, Plakunov et al. developed an alternative universal method of concomitant analysis of planktonic and biofilm cultures. It may be used to characterize biofilm growth, extracellular matrix synthesis, metabolic activity, and the minimum viable cell number per biofilm and is based on using polytetrafluorethylene cubes and glass fiber filters as surfaces for biofilm formation (Plakunov et al., 2016). The method applies to gram-positive and gram-negative bacteria. Moreover, the colonies growing on agar plate surfaces are currently considered as biofilm variants, since they represent stable communities of microbial cells embedded in the extracellular matrix (Branda et al., 2001; Gallegos-Monterrosa et al., 2016).
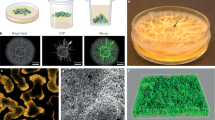
A distinctive property of B. subtilis biofilms is the fact that they display a large number of architectural features; their complexity is due to a high degree of cell specialization and the pattern of intercellular communication among subpopulations. They collectively form a community that is analogous to multicellular organisms in structural terms (Schafer and Turgay, 2019). The formation of a biofilm requires the transition of B. subtilis cells from the planktonic to the nonmotile state. This is associated with flagellar motility suppression and gene expression induction that is necessary for synthesizing the extracellular matrix in accord with environmental signals (surface tension, nutrient depletion, low oxygen levels, etc.) (Gingichashvili et al., 2019). Initially, bacillar cells are short motile rods. During biofilm development, they adhere to one another and the surface while forming long chains composed of nonmotile, extracellular matrix-producing cells. The B. subtilis biofilm matrix is characterized by hydrophobicity; it is responsible for the biofilm’s mechanical rigidity (Dragos and Kovacs, 2017). An innovative technique was suggested for spatially and temporarily monitoring the B. subtilis biofilm development dynamics under various conditions. Using triple fluorescence labeling makes it possible to detect biofilm shape irregularities, measure biofilm thickness, and estimate the distribution of cells with various phenotypes in the biofilm (in terms of motility, matrix production, and sporulation) (Wang et al., 2017).
Large creases or wrinkles on the bacillar biofilm surface are considered to result from the biofilm’s mechanistic instability, which is due to cell translocation inside the biofilm or localized cell death. Locally promoted crease formation brings about an increase in the biofilm’s surface to volume ratio that improves the cells’ access to oxygen (Asally et al., 2012; Trejo et al., 2013; Kolodkin-Gal et al., 2013). Wrinkling facilitates the formation of a complex channel network inside the biofilm, accelerating liquid circulation and nutrient distribution in diffusion-inaccessible biofilm parts. The channels inside biofilms are interconnected, and their morphology and number vary depending on biofilm age and the nutritive conditions of the bacteria (Gingichashvili et al., 2019). It was established that the channels perform an important function in transferring molecules across the biofilm structures, enabling the liquid to efficiently penetrate the deeper layers of the periphery of a mature biofilm (Gingichashvili et al., 2019). Aerial domains used for sporulation and bacterial spore spreading were identified on the surface of the B. subtilis biofilms (Branda et al., 2001). Such three-dimensional architecture secures the integrity of the B. subtilis biofilms and is of significant importance for the adaptability and viability of microbial cells.
REGULATION OF BIOFILM MATRIX SYNTHESIS
The genes that are responsible for matrix exopolysaccharide formation form a part of the eps operon (Nagorska et al., 2010; Marvasi et al., 2010; Roux et al., 2015). The exopolysaccharide-deficient eps mutants of bacilli form flat and highly fragile colonies; they can grow and give rise to cell chains that contain some extracellular material due to the synthesis of additional proteinaceous matrix components (Branda et al., 2006). Cells with mutations in the enzyme genes involved in matrix biosynthesis, pgcA (formerly denoted as yhxB) that encodes α-phosphoglucomutase, and gtaB that encodes UTP-glucose-1-phosphate-uridyltransferase, are characterized by disrupted biofilm formation (Lazarevic et al., 2005). Mutants deficient in the synthesis of uridine-diphosphate-galactose (UDP-Gal), a mandatory precursor for exopolysaccharide synthesis, do not form a biofilm (Chai et al., 2012). UDP-Gal is a toxic intermediate product in the galactose metabolic pathway; it is converted to nontoxic UDP-glucose by the UDP-glucose-4-epimerase (GalE); with a mutant galE gene, cell growth is arrested in the presence of galactose because of UDP-Gal accumulation. The most extensively studied product of the genes of the eps operon is the protein EpsE. This is a bifunctional protein. It is implicated in controlling EPS biosynthesis. The EpsE protein inhibits flagellar rotation via its interaction with the flagellar rotor protein FliG and exhibits glycosyl transferase activity that is necessary for EPS biosynthesis (Guttenplan and Kearns, 2013). Motility inhibition is independent of the glycosyl transferase activity of EpsE. This regulatory mechanism contributes to the active formation of the biofilm matrix upon cessation of cell motility. In some B. subtilis strains, γ‑polyglutamic acid (PGA), another extracellular polymer, is actively synthesized and promotes polymer matrix formation. (Morikawa et al., 2006; Wu et al., 2008). The spectrum of В. subtilis polysaccharides that are components of the biofilm matrix varies depending on growth conditions (Roux et al., 2015). When B. subtilis strain NCIB3610 was grown in a medium with glutamic acid and glycerol, the carbohydrate-containing biomass included such monosaccharides as galactose, glucose, and N-acetylglucose (GalNAc); the level of each of the sugars varied depending on the integrity of the eps operon genes (Chai et al., 2012). Based on these data, the genes involved in galactose metabolism are of significant importance for biofilm formation.
Analysis of the polysaccharides of В. subtilis strain NCIB3610 grown in TY broth (LB broth supplemented with manganese and magnesium sulfates) revealed a mannose-dominated, eps operon-dependent monosaccharide profile (Jones et al., 2014). Apparently, the type of the eps operon genes-produced polysaccharide varies depending on available substrates. Apart from exopolysaccharides synthesized under the control of the eps operon, matrix formation in B. subtilis strains involves the synthesis of an exogenous polysaccharide, levan (the fructose homopolymer). Its biosynthesis depends on levan saccharase encoded by the sacB gene (Benigar et al., 2014; Raga-Carbajal et al., 2016). If bacteria grow in the presence of sucrose, levan is incorporated in the biofilm matrix, which partially compensates for the absence of the eps operon products (Dogsa et al., 2013). With sucrose produced by plants, levan incorporation in the biofilm matrix by bacteria may be a prerequisite for B. subtilis biofilm formation in the rhizosphere under natural conditions.
The structural integrity of B. subtilis biofilms depend on two secreted proteins referred to as TasA and TapA. They are encoded by the gene cluster composed of three genes (tapA‒sipW‒tasA) and denoted as the tapA operon (Branda et al., 2006). TasA homologues are widely spread among bacilli (Dragos, 2017). The TasA protein was envisioned as a functional amyloid-like protein that is synthesized as the monomer. It assembles to form long filaments (fibrils) and is transferred across the membrane by means of a signal peptidase, SipW; the fibrils attach to the cell wall in a process involving the auxiliary protein TapA (Romero et al., 2014). The TasA protein is a functional homologue of E. coli curli amyloid protein. It polymerizes in vitro and can interact with the antibodies specific for amyloid polymers (Huang et al., 2018). In solution, the purified protein TasA is present as an oligomer. The formation of fibrils from TasA oligomers is stimulated in an acidic medium or the presence of a hydrophobic surface (Chai et al., 2013). In vitro research on the different oligomeric forms of secreted protein TasA using NMR, EM, X-ray diffraction, and analytic ultracentrifugation revealed conformational varieties of TasA ranging from the globular state to the proteinase-resistant fibrillar structure, which do not exist with other biofilm-forming proteins (Diehl et al., 2018). According to solid-state NMR data, the main amyloid component of TasA contains β-sheets and α-helices in the secondary structure, indicative of an atypical amyloid architecture in B. subtilis (Mammeri et al., 2019).
The tasA mutants form no biofilm, but this defect is not as serious as that of the eps gene cluster mutants (Branda et al., 2006). The tasA mutants produce cell chains that do not cohere; tasA inactivation also prevents wrinkly biofilm formation. The protein TasA is implicated in the orderly maturation of Bacillus biofilms. Its lack of results in a number of physiological alterations that are caused by changes in membrane stability and dynamics and in a decrease in interacting cells’ viability. This is due to the absence of a structured matrix, which, in turn, sets limits to stress adaptation options available for the cells (Camara-Almiron et al., 2020). TasA possesses antimicrobial properties; tasA expression in B. subtilis biofilms is stimulated in the presence of the pathogen Fusarium culmorum (Khezri et al., 2016). Interestingly, TasA also performs an important role in recognition of the kinship degree between closely related B. subtilis strains, which involves various inhibitor molecules (Lyons et al., 2016). Finally, tаsА-mutant strains fail to form root-associated biofilms (Dragos et al., 2017). Since the plant rhizosphere is the natural habitat of B. subtilis, TasA might be one of the key secreted compounds that impact the ecology and the evolution of this species.
The TapA protein is a minor component of TasA-based protein fibers, which binds to TasA and is involved in attaching TasA fibers to the cell wall. TapA plays an important part in TasA localization, as well as in its polymerization and TasA fiber assembly from monomers (Romero et al., 2014). In the absence of the tapA gene, no wild-type TasA fibers are formed, and the total TasA protein level in the exogenous matrix is decreased. These data indicate that the TapA protein provides for TasA stabilization. Therefore, the fibrillar TasA form is more resistant to proteinase degradation than the oligomeric form. This indicates that the tapA gene deletion critically affects biofilm formation in B. subtilis.
The signal peptidase SipW cleaves the TapA precursor, in which it recognizes the N-terminal signal sequence, enabling the protein’s release from the membrane and formation of the cell wall-attached fibrils (Stover and Driks, 1999). Mutations in the sipW gene in B. subtilis strains result in impeding their adherence to polyvinyl chloride or glass surfaces under laboratory conditions (Hamon et al., 2004). This is due to the fact that peptidase SipW is a bifunctional protein that operates as a signal peptidase during TasA and TapA processing. Its function is aimed at activating the expression of the eps operon, which involves the protein’s C-terminal domain. Such activation is a prerequisite for initiating biofilm matrix formation via cell adherence to the surface.
At the final stages of biofilm maturation, bacterial hydrophobin BslA is synthesized, which operates in tandem with the TasA protein (Hobley et al., 2015). The protein BslA (formerly denoted as YuaB) is necessary for the surface hydrophobicity of the B. subtilis biofilm and formation of its complex morphology and architecture (Arnaouteli et al., 2017). The biofilm of B. subtilis with the defective phenotype bslA− possesses amphiphilic properties and resembles those of the mutants with deletions in the eps and tasA genes (Ostrowski et al., 2011). The BslA protein is a surfactant that consists of an immunoglobulin-like domain and a unique highly hydrophobic domain (Hobley et al., 2015). It may assume two different structural conformations that correspond to two functional forms: monomeric BslA is necessary for the development of a complex biofilm structure, whereas the dimeric protein is required for forming the hydrophobic biofilm surface (Arnaouteli et al., 2017). BslA dimerization involves the formation of a disulfide bond and depends on two conserved cysteine residues that are located in the C-terminal domain of the protein molecule. The conformational transition is subject to regulation by the medium redox state. In a native biofilm, BslA oligomerization depends on electron acceptor availability, and the dimer is the predominant form in an oxygen-enriched area with an open surface.
Therefore, all biofilm matrix components, including exopolysaccharides (EPS and PGA) and proteins (TasA, ТасА, and BslA), are necessary for formation of the extracellular matrix of B. subtilis, and they influence the growth and the final size of the biofilm. Interval microscopy and light profilometry revealed the two matrix components, the exopolysaccharide produced by an epsA-O operon gene, and the surface-layer protein BslA, that control the biofilm surface area and demonstrated that the mature biofilm height depends on the protein BslA (Kesel et al., 2017). Interactions among the matrix components in biofilms are responsible for emergence of an adaptable structure during the biofilm development process. At the early stages of biofilm development, formation of the 3D structure involves interaction between matrix components and exogenous DNA (that also contributes to the maturation of B. subtilis biofilms) (Peng et al., 2020).
BIOFILM BISTABILITY
Biofilm formation and development and synthesis of the extracellular matrix components are stringently regulated at the level of transcription and post-transcriptional and post-translational modification (Reverdy et al., 2018). All cells in a planktonic population are motile. Gene expression that is necessary for motility proceeds when the translational regulators Spo0A, DegU, and ComA are in the dephosphorylated state. Their activation in response to the presence of various extracellular signals results in reducing the motile cell population (Vlamakis et al., 2013). During the course of biofilm maturation, bacillar cells differentiate into several types, forming subpopulations of genetically identical but phenotypically distinct cells that respond to different signals and perform different functions for the benefit of the whole community, in order to minimize the energy costs involved. For instance, the total transition of all cells to the energetically costly stage of sporulation is prevented in this fashion (Lopez and Kolter, 2010). A complex regulatory network is responsible for activating the specific gene cascades that are responsible for the differentiation of all coexisting cell types in B. subtilis biofilms.
Cell differentiation in biofilms was initially detected in a microscopic study which revealed that the aerial biofilm domain contains a sporulation-specific reporter (Branda et al., 2001). These findings were subsequently confirmed in experiments with fusion vectors containing fluorescent reporter genes (Veening et al., 2006). Within a biofilm, cells specializing in motility, polymer matrix formation, sporulation, and exoproteinase secretion were identified. Using flow cytometry and fluorescence microscopy, it was established that the number of exoproteinases-producing cells increases during biofilm maturation; this is due to elevated exoproteinase gene transcription and an increase in exoprotease activity in time (Marlow et al., 2014a). Proteinases are synthesized by matrix-producing and matrix-nonproducing cells, indicative of the important role of these enzymes at early stages of biofilm development. The data of these experiments demonstrated a large diversity of specialized biofilm cell types and testified to the regulatory plasticity of the community.
Activation of the Spo0A transcription factor via phosphorylation triggers spore formation only in specific B. subtilis cell subpopulations (Fujita et al., 2005). There are two different levels of Spo0A-P activation: the lower activity level (in Spo0A-ON cells) results in matrix formation and the higher activity level results in sporulation (Fig. 1b). Spo0A-ON cells become matrix producers, and they lose motility due to cohesion activation (protein EpsE) (Guttenplan and Kearns, 2013). The Spo0A-ON cells are also cannibals, and they are insensitive to the effects of two self-secreted peptide toxins (peptides Skf and Sdp) that kill their toxin-sensitive conspecifics (Höfler et al., 2016). This process is regarded as cannibalism: when the available nutrients are insufficient, dead cells are utilized as substrate by surviving cells in the population and impede sporulation: this behavior depends on secreted proteinases (Kobayashi, 2019).
Regulatory pathways controlling biofilm formation in B. subtilis: (a) mode of action of repressor SinR; (b) mechanism of biofilm formation in B. subtilis under the control of the Spo0A-P transcription regulator; (c) involvement of transcription factors DegU-P and ComA-P in controlling biofilm formation in B. subtilis. Т-shaped lines and arrows indicate inhibition and activation, respectively. See text for description.
Activation of the transcription factor ComA (ComA-P) results in cell differentiation in the community and formation of a subfraction of competent cells that are capable of taking up exogenous DNA in order to increase the genetic variability of the population. Activation of ComA-P is subject to regulation by a quorum sensing-dependent mechanism; it is induced once the threshold population density is achieved (Zeriouh et al., 2014). ComA-P cells also express the paracrine signal surfactin that triggers the expression of the genes and operons enabling matrix formation (Fig. 1c) (Aleti et al., 2016). The activity of the ComA/ComP response regulator pair is modulated by the ComX signal peptide. As an autoinducible signal, it controls competence, surfactin production, and, indirectly, extracellular proteinase synthesis in B. subtilis (Špacapan et al., 2018). These processes are regulated in a bacterial population in a size-dependent fashion, indicating that ComX is a quorum-sensing signal that controls exoproteinase production via the intermediate transcription regulator DegQ.
Activation of the DegU transcription factor (DegU-P) results in formation of an intra-community subpopulation of cells referred to as miners that specialize in secreting extracellular proteinases (Fig. 1c) (Veening et al., 2008). The products of these cells promote breaking down large biopolymers into smaller oligopeptides. It was revealed that the peptide ComX induces expression of the subtilisin gene (aprE) and stimulates that of the exoproteinase gene in B. subtilis biofilms. Exoproteases can degrade the autoinducible signal peptide ComX. The inhibitory effect on the biological activity of ComX can be removed by adding subtilisin. Suppression of metalloproteinases by EDTA lowers ComX activity, suggesting that serine proteinases and metalloproteinases are capable of degrading the ComX peptide (Špacapan et al., 2018). These data indicate that secreted proteolytic enzymes fulfill regulatory functions and break down the autoinducible peptide ComX that exerts an important influence on their expression in biofilms. The cell subpopulation with active exoproteinases differs from those of motile or matrix-producing cells, and it is located closer to the biofilm surface (Marlow et al., 2014b). This is consistent with the fact that DegU-ON cells actively express the water-repellent protein BslA that is necessary for maintaining the integrity of the B. subtilis biofilm (Hobley et al., 2015; Arnaouteli et al., 2017).
Analysis of the spatial arrangement of specific subpopulations in mature biofilms by means of fluorescent reporter proteins revealed that motile cells in the lower biofilm layer were responsible for its spatial expansion, whereas the cells located in the biofilm core specialized in producing the extracellular matrix and maintaining its rigidity (Vlamakis et al., 2013). New techniques were suggested to visualize the internal structure of the B. subtilis biofilm while retaining its architecture and ultrastructure (Fuchs et al., 2018). Spores are present in the upper biofilm part on aerial structures that facilitate their dispersal. Setting apart the spatial and temporal distribution of competent and surfactin-producing cells, the two separate subpopulations in the total microbial community, presents difficulties. The state of bistability is a strategy aimed at adapting the whole community to varying environmental conditions, enabling a genetically identical population to form subpopulations with distinct physiological properties and to provide the whole cell community with necessary metabolites. This strategy was called the growth-hedging strategy (Veening et al., 2008).
REGULATORY PATHWAY OF BIOFILM FORMATION
The genetic pattern of biofilm formation regulation in B. subtilis is understood better than the transfer of the signals that trigger this process (Mielich-Süss and Lopez, 2015). The factor Spo0A plays the role of the central transcription regulator in stationary-phase B. subtilis cells. It controls the expression of over one hundred genes required for sporulation and biofilm formation (Fig. 1b) (Chastanet et al., 2011). The activity of this protein is regulated via its phosphorylation: extracellular signals activate sensor histidine kinases that set the phosphorelay in action, resulting in Spo0A phosphorylation. The level of phosphorylated Spo0A in the cells impacts the gene expression profile (Schultz et al., 2016). Spo0A phosphorylation induces biofilm formation, while biofilm maturation is associated with accumulation of the Spo0A-P in some of the cells, which activates their sporulation. Once the threshold levels of Spo0A phosphorylation are attained, two antirepression pathways are induced, activating transcription of the operons that are of paramount importance for biofilm matrix synthesis.
The first pathway is aimed at suppressing the AbrB receptor that blocks transcription of the genes involved in a large number of processes, including biofilm formation (Chumsakul et al., 2011). It was established that the expression of the ahpA gene that codes for the AphA peroxidase depends on the transitional state regulator, AbrB, and transcription factor Spo0A. AhpA is specifically expressed during biofilm formation but not during the stationary phase or sporulation. This gave grounds for the suggestion that derepression of the peroxidase gene ahpA under the influence of the regulator AbrB requires a signal that is different from the signals associated with the cells’ transition to the stationary phase (Fig. 1b) (Zwick et al., 2017). The AbrB regulator is controlled by the factor Spo0A in two different ways: (1) Spo0A-P represses abrB gene transcription and (2) Spo0A-P facilitates the expression of the factor AbbA, the AbrB antirepressor that binds to AbrB and isolates it from the target DNA (Tucker et al., 2014).
The second pathway involves the small DNA-binding protein SinR that inhibits biofilm formation and plays a pivotal role in sealing the cells’ fate (Fig. 1a) (Kampf et al., 2018). The protein SinR forms tetramers and binds to the sites in the promoter regions of the epsA-O and tapA‒sipW‒tasA operons that are necessary for the synthesis of the extracellular polysaccharide and amyloid fibers in the extracellular matrix, respectively (Cairns et al., 2014; Stowe et al., 2014). SinR activity is controlled by antagonistic proteins, SinI and SlrR. SinI or SlrR binding to SinR inhibits the DNA-binding capacity of SinR and relieves the repression of the biofilm operons (Fig. 1a) (Newman et al., 2013). Moreover, insignificant changes in the SinR expression levels drastically influence expression of the matrix genes, predominantly because of the hypersensitivity of the SinI‒SinR and SlrR‒SinR regulatory modules, indicative of a stringent regulation of their formation, including the metabolic one (Chai et al., 2011; Greenwich et al., 2019).
One of the proteins involved, SinI, is activated upon Spo0A phosphorylation. It binds to SinR, forming a heterodimeric complex, and isolates SinR from DNA targets, resulting in expression of SinR-repressed genes (Fig. 1a) (Newman et al., 2013; Stowe et al., 2014). The phenotypic transition from the planktonic to the biofilm state is under the control of transcription repressor SinR and is, therefore, affected by its inactivation by its primary antagonist, SinI. The mechanism of degradation of the SinR tetramer was revealed; it involves the protein SinI that regulates the expression of SinR-repressed genes (Milton et al., 2020). If the levels of SinR exceed those of its antagonists, matrix gene expression is suppressed. It was established that SinR is expressed in the majority of cells, whereas SinI, only in a part of the population. The functions of proteins SinI and SinR are gene dosage-sensitive: duplicating sinI and sinR gene copies results in suppressing the synthesis of matrix polymers (Chai et al., 2011; Ogura et al., 2016). Overall, matrix gene expression is temporary because it depends on the affinity of Spo0A-P to the sinI gene promoter and the number of sinI and sinR gene copies; sporulating cells do not spend energy on producing the extracellular matrix.
The other protein SlrR antirepressor, SinR, is required for controlling biofilm formation in two different ways (Chai et al., 2010). First, it binds to the repressor SinR, forming the SinR‒SlrR complex and preventing SinR binding to the matrix gene promoters (Fig. 1а). Since the regulator SinR represses the slrR antirepressor gene that binds to the regulator SinR, this enables controlling the SinR‒SlrR complex according to the double feedback loop principle. The slrR gene remains derepressed as long as SlrR prevents SinR-dependent repression. When the SlrR level in the cells is high, expression of the matrix gene operons can proceed because the level of free receptor SinR is low under these conditions. When the SlrR level in the cells is low, the regulator SinR is uninhibited and it blocks the operons involved in biofilm matrix synthesis. Second, the SinR-SlrR complex is implicated in controlling the expression of (i) the hag gene encoding flagellin that is necessary for motility and (ii) autolysine genes that encode the proteins involved in cell detachment (the lytABC and lytF genes) (Fig. 1a). A recently identified regulatory mechanism involves interaction between SinR and the degU gene promoter; thereupon, the repressor SlrR forms a complex with SinR on the degU promoter. Hence, SlrR appears to play an active role in degU expression (Ogura et al., 2014).
Cell cohesion is of significant importance for the onset of biofilm formation in B. subtilis. The regulatory protein SlrR is unstable; it belongs to the LexA peptidase superfamily and is degraded in the cell, partly due to proteinase ClpCP. Instability of the SlrR protein enables the derepression of the autolysine-encoding genes; activation of these genes causes cell chains to separate into solitary cells (Chai et al., 2010). Therefore, the SlrR‒SinR switch can exist in two different states: (i) at a low SlrR level characteristic of solitary motile cells and (ii) at a high SlrR level typical of matrix-producing cell chains. The transition from the low- to the high-level state is made using the protein SinI that is synthesized under the control of factor Spo0A-P in response to the activation of the histidine kinases. Protein SinI accumulation inhibits repressor SinR, resulting in derepressing the gene and producing protein SlrR. This causes SlrR accumulation, switching SlrR‒SinR to the high SlrR level state and favoring biofilm formation.
Complex interactivity between SinR and its genetically related antagonists is responsible for the bistability state of the system. This interactivity is under the influence of phosphodiesterase YmdB involved in controlling activity of the repressor SinR: ymdB gene-deficient cells are characterized by SinR hyperactivity and do not form biofilms (Kampf et al., 2018). The majority of ymdB mutant cells express the genes that are necessary for motility and chemotaxis. Suppressor analysis revealed that ymdB mutants readily acquire SinR-targeting mutations that restore biofilm formation (Kampf et al., 2018). These data indicate that phosphodiesterase YmdB is required for the homeostasis of SinR and/or its antagonists. It was established that, at the post-transriptional level, sinl‒sinR and sinR transcripts are controllable via degradation by the RNase of Y-containing protein complexes (DeLoughery et al., 2016). An important function in regulating protein activity in Bacillus cells is performed by ATP-dependent proteinases. The results of global proteomic analysis and western immunoblotting revealed that SinR levels were decreased in a mutant with defective highly conserved ClpYQ proteinase and altered biofilm formation regulation (Yu et al., 2018).
Regulatory proteins Rem A and RemB are also mandatory for biofilm formation (Fig. 1a) (Cairns et al., 2014). The regulator RemA is a DNA-binding protein that activates the eps and tapA operon transcription and also controls slrR antirepressor gene transription. It was established that regulation of the slrR gene expression requires both regulatory proteins, RemA and RemB (Winkelman et al., 2013). In addition, genetic analysis revealed that regulators RemA and RemB control matrix operon expression in an antirepressor SlrR-independent fashion and operate in parallel with other regulators (DegU, AbrB, and SinR). RemA binds to the DNA upstream of the promoter of the eps operon, overlapping with the site used for the interaction with repressor SinR. In this system, the protein SinR prevents binding of the protein RemA to the promoter (Fig. 1а) (Winkelman et al., 2013). In an similar fashion, the RemA binding site is upstream of the promoter of the tapA operon but SinR and RemA can concomitantly bind to the DNA in this situation. These data testify to an additional pathway for controlling biofilm matrix gene expression. The remA gene is located on the chromosome DNA of B. subtilis close to the genes involved in the stringent response; this fact might suggest a regulatory relationship between remA gene expression and the cell’s metabolic status (Winkelman et al., 2013).
It was revealed that the components of the epigenetic switch can be subject to supplementary regulation involving regulatory proteins YwcC and SlrA. SlrA is one of the antirepressors of the regulator SinR, and protein YwcC is a TetR type transcription repressor. It represses the slrA gene (Fig. 1a) (Chai et al., 2009). If the regulator YwcC responds to an unidentified signal and its expression is suppressed, biofilm matrix genes are induced via SlrA-mediated inactivation of repressor SinR. In contrast to SinI, the protein SlrA is produced in almost all cells. Therefore, this YwcC–SlrA-involving regulation pathway enables B. subtilis cells to rapidly respond to environmental stress factors and to protect the bacterial community by forming a biofilm.
Expression regulation of the bslA and pgs genes in B. subtilis depends on the DegS‒DegU signal transduction system in which DegU is the transcription regulator. It is phosphorylated by a genetically related histidine kinase, DegS, and controls a number of cell processes: swimming and burrowing motility, biofilm formation, exoproteinase synthesis, gamma-polyglutamic acid (PGA) production, and sporulation (Gabdrakhmanova et al., 2005; Kayumov et al., 2006; Sharipova et al., 2008; Cheremin et al., 2014). It was established that the degU mutant forms no submerged biofilm that requires the PGA polymer, the product of the pgs operon (Stanley et al., 2005). Biofilm formation in the degU mutant is disrupted because the matrix loses its hydrophobicity, due to impaired expression of hydrophobin BslA gene (Kobayashi and Iwano, 2012). The phosphorylation degree of the transcription factor DegU (DegU-P) impacts bacterial cells’ behavior, initiation of biofilm formation requiring intermediate activation levels of transcription factor DegU. Indirect biofilm formation inhibition in the cells occurs at high factor DegU phosphorylation levels (Fig. 1c) (Marlow et al., 2014b). Biofilm formation activation starts when phosphorylated DegU-P triggers transcription of the bslA gene coding for the hydrophobic protein (Hobley et al., 2015). The DegS‒DegU pathway is activated upon flagellar rotation inhibition, which may result from cell adhesion to the surface and motility loss (Fig. 1c). A decrease in flagellar rotation causes an increase in phosphorylated DegU-P level, inducing transcription of the target genes, including the bslA gene (Chan et al., 2014; Cairns et al., 2014). Therefore, flagellar rotation arrest is an additional signal for polymer matrix formation.
Of paramount importance in biofilms with a high cell density is an autoinducible signaling system (Špacapan et al., 2018). Binding of signal molecules to specific receptors, results in transcription and synthesis of required secreted products referred to as public goods (exemplified by proteinases). In B. subtilis, the autoinducible signaling system is based on peptides, signal molecules that control many adaptive processes, including exoproteinase synthesis. The autoinducible signal peptide, ComX, is implicated in regulating bacterial competence and surfactant production in B. subtilis (Aleti et al., 2016; Pollak et al., 2016). Both traits are subject to regulation by the bacterial population size, which indicates that ComX is a quorum-sensing signal. In B. subtilis, the comQXPA gene cluster is the main autoinducible peptide-based system operating as a quorum-sensing system. ComX regulates expression of the srf operon that is responsible for surfactin synthesis and is involved in genetic competence development (Aleti et al., 2016). Within the ComQXPA system, the activity of the ComP‒ComA response regulator pair is modulated by the signal peptide ComX. ComX is modified by the isoprenyl transferase ComQ (Okada et al., 2005). Extracellular accumulation of modified ComX results in ComA phosphorylation and subsequent regulon ComA induction. The degQ gene also forms a part of this regulon. A high DegU-P level upregulates the synthesis of extracellular enzymes, involving the aprE gene that encodes the serine exoprotease of B. subtilis (Veening et al., 2008). Hence, ComX and the ComQXPA system positively control the transcription of the subtilisin gene aprE and exoproteinase production during biofilm growth. Exoproteinases can degrade the signal peptide ComX (Špacapan et al., 2018). In fact, two ComX-involving regulation types were described: genetic upregulation and biochemical downregulation. Thanks to the ComX signal molecule, this feedback loop maintains the bacterial demand–supply balance with respect to public goods.
Apart from complex regulation by transcription factors, the eps operon expression is controlled by the RNA cis-element that is encoded by the DNA region between the second and the third eps operon genes. The RNA cis-element was denoted as EAR (the element associated with RNA); it was identified in the overwhelming majority of the genomes of representatives of the family Bacillaceae. EAR operates as an antiterminator and increases the eps operon expression by interacting with RNA (Irnov et al., 2010).
While the main genetic system of regulating biofilm formation at the transcription level sufficiently is well characterized, there are scarce data on the role of metabolism (carbon metabolism, biosynthesis and fermentation pathways, and secondary metabolism) in this process (Pisithkul et al., 2019). The secondary messenger c-di-AMP performs an important function in B. subtilis biofilm formation and attachment to plant roots (Townsley et al., 2018). In contrast to most other amino acid biosynthesis genes, serine biosynthesis gene expression in B. subtilis cells decreases upon transition to the stationary phase, which represents a crucial intracellular signal for biofilm matrix biosynthesis activation (Greenwich et al., 2019). The relationship between carbon metabolism and biofilm formation was studied (Chen et al., 2015). Analysis of the lysine-acetylated proteins, or the acetylome, of B. subtilis revealed that such protein modification fulfills an important regulatory function in terms of biofilm development (Reverdy et al., 2018). A linkage between the cysteine biosynthesis pathway and biofilm formation was established (Kobayashi, 2019). Biofilm formation is triggered when the cells are depleted of glucose; the process is catabolite-controled (Chen et al., 2015). Removing the ccpA gene stimulates biofilm formation, indicating that the protein CcpA downregulates biofilm formation in B. subtilis. Another global metabolic regulator, CodY, is also involved in biofilm formation in B. subtilis (Brinsmade et al., 2014). The bacteria are capable of synthesizing polyamines, and transcriptomic analysis of spermidine-depleted B. subtilis mutants demonstrated that spermidine was necessary for biofilm formation. It activates the transcription of matrix exopolysaccharide synthesis genes and the tasA operon via regulator SlrR (Fig. 1a) (Hobley et al., 2017).
The aforementioned regulation pathways suggest the operation of a complex multilayer regulatory network that controls the differentiation of B. subtilis cells into subpopulations and biofilm formation. A majority of the regulators are specific for this organism only, their functional homologues have not been detected in other bacterial species. The evolution of the regulatory network was aimed at enabling the bacilli to adequately respond to metabolic alterations and environmental changes via formation of a single integrated community of genetically identical biofilm-embedded microorganisms that, nevertheless, differ in terms of physiological functions.
BIOFILM FORMATION SIGNALS
The data on the signals that induce biofilm formation and differentiation of microorganisms into subpopulations within a community are scarce. Nevertheless, numerous signal receptors for the regulatory systems have been described. This suggests that biofilm formation can be elicited by a variety of factors. The Spo0A-P concentration in the cell is determined by the activity of a family of histidine kinases (KinA, KinB, KinC, and KinD) that recognize signals and transmit them to the protein Spo0A via phosphotransferases (Jiang et al., 2000; Grau et al., 2015). The phosphorelay includes phosphotransferase Spo0F that is phosphorylated by a kinase and donates the phosphate group to the Spo0B phosphotransferase, which phosphorylates factor Spо0A. The fifth kinase, KinE, may be involved in this pathway; apparently, it influences the matrix gene expression (McLoon et al., 2011). None of the indentified kinases is fully responsible for expression of the polymer matrix genes; it is more likely to result from combined operation of several different kinases, and their activities vary depending on diverse input signals from the environment.
Histidine kinase autophosphorylation proceeds in compliance with the cis mechanism. One kinase subunit phosphorylates itself inside the multimer (in the trans mode), and one kinase subunit phosphorylates another subunit. It was established by in vivo and in vitro studies that autophosphorylation of the main histidine kinase, KinA, during sporulation can be based on the trans-mechanism and proceed upon nutrient depletion (Devi et al., 2015).
The first signal molecule to be identified as a biofilm gene inducer is the secreted antibiotic surfactin (Aleti et al., 2016). As a signal molecule, surfactin initiates factor Spo0A phosphorylation via the sensor kinase KinC and promotes the activation of expression of the polymer matrix genes (Mielich-Suss et al., 2015). The cells synthesizing and secreting surfactin do not belong to surfactin-recognizing cells (Aleti et al., 2016). This phenomenon is considered in terms of paracrine signal transfer: the signal producer does not respond to its own signal. This system is different from the quorum-sensing system in which each cell produces, and responds to, a signal molecule (Aleti et al., 2016). Gene expression activation in response to the surfactin signal does not comply with the traditional mechanism that involves sensor protein binding to the ligand. Surfactin is a lipopeptide; its molecule can be incorporated into the cytoplasmic membrane, resulting in potassium ion efflux that causes histidine kinase KinC activation, signal transfer to Spo0A, and induction of expression of the biofilm matrix genes (Grau et al., 2015).
Bacteria of the genus Bacillus have much agricultural potential because they produce lipopeptides with high antimicrobial activity (Velmourougane et al., 2017; Aktuganov et al., 2019). The three families of Bacillus lipopeptides considered in the literature are the surfactin, iturin, and fengycin family. Their molecules are amphiphilic and efficiently interact with biological membrane structures. Bacteria, fungi, oomycetes, and viruses are sensitive to their antimicrobial activities. It was revealed that these compounds stimulate biofilm formation, which is a key factor in terms of successful colonization activity of biological control agents (Penha et al., 2020). Other compounds that cause potassium efflux, such as the fungicide nystatin and the antibiotic valinomycin, also induce biofilm matrix gene expression via histidine kinase KinC. Surfactin, nystatin, and valinomycin are naturally produced by soil microorganisms, and B. subtilis encounters these compounds under natural conditions. Nystatin induces signal transfer in B. subtilis via binding to the membrane ergosterine. Taken together, the data obtained give grounds for the suggestion that B. subtilis membranes contain microdomains, similar to the lipid rafts in the plasma membranes of eukaryotic organisms (Lopez, 2015). Chlorine dioxide also induces biofilm matrix gene expression in a histidine kinase KinC-dependent fashion (Shemesh et al., 2010). Unlike surfactin, chlorine dioxide causes histidine kinase KinC activation by dissipating the cell’s membrane potential. An increase in matrix component synthesis in the presence of membrane structure-disrupting inhibitors is essential for the survival of B. subtilis, since the exopolysaccharide is responsible for protection against the lethal consequences of the impact of these compounds.
Bacillus sp.-produced cyclic lipopeptides, especially surfactins, launch biofilm formation and root colonization and are of paramount importance for the activity of biocontrol agents and systemic plant resistance (Aleti et al., 2016). An important function of surfactin is based on its capacity to destroy membrane vesicles in bacilli (Brown et al., 2014). The vesicles are formed by B. subtilis biofilm cells. Prophage-encoded endolysin can form openings in the cell wall, through which the cytoplasmic membrane material is released in the form of vesicles (Brown et al., 2015; Toyofuku et al., 2017). The vesicles can encapsulate the factors that are important for the whole community and contribute to its survival. In the case of gram-positive bacteria, surfactin lyses extracellular vesicles and releases their content.
In B. subtilis mutants with disrupted matrix synthesis, studies with transcription reporters revealed prolonged expression of eps and tasA operons and decelerated sporulation in the biofilm (Aguilar et al., 2010). This is due to the functioning of kinase KinD of B. subtilis, which exhibits kinase and phosphatase activities. In the capacity of a phosphatase, KinD is implicated in maintaining a low level of phosphorylated Spo0A-P, which is associated with biofilm matrix formation. In the capacity of a kinase, KinD facilitates spore formation. Transcription of the main operons responsible for matrix formation is markedly increased in response to the presence of glycerol and manganese in the medium; this involves the activation of histidine kinase KinD responsible for detecting these substances outside the cell (Shemesh et al., 2013). There is evidence that KinD is activated in response to compounds produced by soil microorganisms or contained in the plant root exudate (Chen et al., 2012).
All these genetic cascades are controlled by the Rap family of phosphatases that specifically interact with exogenous genetically related peptides (Phr peptides) (Veening et al., 2005). It is known that phosphatase RapGH dephosphorylates DegU-P, whereas phosphatase RapABEJ impacts phosphate transfer in the Spo0A phosphorelay system and phosphatase RapCFGHK targets ComA-P. Combined, they may be considered an alternative intercellular signal transfer system aimed at implementing the bistability program in B. subtilis; they are involved in regulating cell specialization and collective behavioral processes, enabling the cells’ interactivity and species survival (Verdugo-Fuentes et al., 2020).
We are still far from understanding the whole Phr‒Rap regulatory network of B. subtilis because of the complexity of the regulatory cascades. An additional complexity level in controlling the signaling networks in B. subtilis was revealed. The membranes of bacilli contain microdomains; their integrity is a prerequisite for histidine kinase KinC activation and, accordingly, for Spo0A-ON cell differentiation (Lopez, 2015). It was demonstrated that Bacillus membranes are characterized by compartmentation of signal transfer pathways enabled by functional membrane microdomains that play an important role in activating the signal transduction cascades (Wagner et al., 2017). A membrane-bound proteinase, FtsH, was established to form a part of the microdomains; it selectively hydrolyzes specific Rap phosphatases (Mielich-Suss et al., 2013). Therefore, integrity of the microdomains of the cytoplasmic membrane of B. subtilis is essential for inducing biofilm formation because it protects and stabilizes the proteins FtsH and KinC. Membrane microdomain disruption (caused by zaragozic acid) results in inhibiting the signal cascades that enable biofilm formation in B. subtillis (Lopez, 2015; Wagner et al., 2017). Likewise, proteinase activity of the protein FtsH is suppressed by the SpoVM peptide that can be used as a biofilm inhibitor (Yepes et al., 2012). Microdomains contain only a limited set of proteins that are involved in specific cell processes such as proteinase secretion and signal transduction (Lopez, Koch, 2017).
Nonsignal mechanisms can be used in B. subtilis for activating the genes encoding the matrix components. Apart from gene transcription activation mechanisms for matrix polymer synthesis at a low phosphorylated Spo0A-P level in the cells, two cannibalism gene clusters are expressed in B. subtilis. As a result, some of the cells in the population synthesize and secrete the toxic peptides SDP and SKF. Bacteria that synthesize these toxins are resistant to them (Höfler et al., 2016). A prerequisite for the SKF peptide transport into the extracellular space is the ABC transporter that removes the toxin from the cell (Gonzalez-Pastor, 2011). The resistance of bacteria to the other peptide, SDP, is due to availability of the specific protein SdpI in the membrane. Its synthesis is induced in the presence of toxic SDP in the cells with a high concentration of phosphorylated regulatory protein Spo0A-P (Ellermeier et al., 2006; Kobayashi and Ikemoto, 2019). Cannibal cells release toxic peptides that selectively kill B. subtilis cells that do not express the genes encoding them. Since the genes encoding the toxins and those encoding the matrix polymer are activated at a low level of the Spo0A-P phosphorylated regulatory protein, there is an overlap between the cell populations that form the biofilm and those that synthesize the toxic peptides (Lopez et al., 2009). As a result, the cells initiating matrix polymer synthesis also release toxic peptides aimed at reducing the cell population that does not produce the biofilm matrix. This results in forming a population containing an increased number of matrix-producing cells. Cannibalism toxins kill not only B. subtilis cells but also the cells of other species if B. subtilis grows in mixed cultures (Liu et al., 2010). Co-cultivation of B. subtilis (studies were conducted using fluorescent matrix gene reporters) with other soil microorganisms in the presence of biofilm formation inducers revealed that the overwhelming majority of microorganisms belonged to the genus Bacillus, despite the bacterial diversity of the soil samples.
To sum up, various signal molecules ranging from surfactin to cannibalism toxins can promote cell number growth in order to stimulate biofilm formation in the population. This involves transferring signals that cause differential gene expression or selective destruction of B. subtilis cells that do not synthesize the biofilm polymer matrix.
BIOFILM DISPERSAL AND DISSOCIATION
Bacterial biofilms are of significant importance for their natural habitats, as well as in terms of biotechnology and medicine. However, our understanding of the regulation of biofilm development and their maintenance in various niches is far from being complete. An important stage in a biofilm life cycle is its dispersal when motile cells exit a mature biofilm in order to spread and to colonize new niches (Plakunov et al., 2017). The process is triggered by various exogenous and endogenous factors, such as changes in available nutrient levels, bacterial autoinducers, fatty acids, peptide signals, nitric oxide, and stress signals (starvation, iron or phosphate excess, etc.). Such signals bring about changes in the concentrations of such secondary messengers as cyclic diguanylate monophosphate (c-di-GMP), cyclic diadenosine monophosphate (c-di-AMP), guanosine tetraphosphate (ppGpp), guanosine pentaphosphate (pppGpp), and small regulatory RNAs that change bacterial gene expression activity (Romling and Galperin, 2017; Townsley et al., 2018). Biofilm dispersal is subject to regulation at the level of transcription of operons that code for the biofilm matrix. It was established that a mechanism of inhibiting expression of the matrix genes is activated in certain B. subtilis cells, and they revert to the planktonic state (Vlamakis et al., 2013; Norman et al., 2013). At the gene expression level, this results from lowering the antirepressor SlrR level during biofilm maturation. This is due to SlrR instability caused by proteinase ClpCP-catalyzed degradation and self-destruction (Chai et al., 2010). Degradation of the SlrR regulator enables the SinR repressor to interact with the promoter regions of matrix operons, inhibiting the biosynthesis of the matrix components (Ogura, 2016; Milton et al., 2020).
The general stress-activated transcription factor SigB in B. subtilis is necessary for stopping the expansion of a mature biofilm and triggering the mechanism of dispersion when available nutrients become limited (Bartolini et al., 2018). Interestingly, biofilms with a defective sigB gene are larger than wild-type biofilms but display accelerated cell death, elevated stress sensitivity, and decreased cell dispersion. A regulatory relationship was revealed between the factor SigB and expression of the SinR repressor. This new SigB‒SinR regulatory system is important in terms of controlling biofilm adaptability to various media in which the regulatory status of the factors SigB and SinR determines whether the cells remain in the biofilm or leave it if the conditions become unfavorable.
Biofilm dissociation is based on controllable degradation of matrix macromolecules in the extracellular environment (Plakunov et al., 2017). This process may involve extracellular proteinases (Marlow et al., 2014a). There is evidence that nattokinase, a subtilisin-like B. subtilis proteinase (Dabbagh et al., 2014), and a subtilisin-like B. pumilus proteinase can degrade the amyloid peptide (Danilova et al., 2014). Presumably, the extracellular proteinases of bacilli are implicated in detaching the TasA fibers of the biofilm matrix (Mitrofanova et al., 2017). The quest for research strategies for investigating biofilm dissociation is of considerable social importance. These data have much potential in terms of eradicating chronic infections, preventing pipepline plugging, etc. (Hobley et al., 2015; Velmourougane et al., 2017; Vaccari et al., 2017). Adding D-amino acids to biofilm cultures causes B. subtilis biofilm dissociation because their incorporation into the peptidoglucan blocks the translocation of protein TapA into the cell wall, resulting in TasA fiber degradation (Romero et al., 2014; Yu et al., 2016). Incorporation of D-amino acids into proteins can be prevented by D-aminoacyl-tRNA-deacylase that removes D-amino acids from incorrectly loaded tRNAs. The B. subtilis strain with a mutation in the D-aminoacyl-tRNA-deacylase gene (the dtd gene) was characterized by suppressed biofilm formation; no biofilm inhibition was documented after eliminating the mutation (Leiman et al., 2013). In molecular terms, the toxicity of D-amino acids is based on their ability to replace respective L-isomers during protein synthesis and to disrupt protein functions.
Another compound that can be implicated in biofilm dissociation is norspermidine, a polyamine (Si et al., 2015; Wu et al., 2016). Norspermidine can specifically interact with the matrix exopolysaccharide and disrupt the biofilm structure. B. subtilis mutants that did not produce norspermidine formed long-lived biofilms (Kolodkin-Gal et al., 2012). Bioinformatic search for homologues of E. coli norspermidine biosynthesis pathway genes in the B. subtilis genome failed to detect homologous genes (Hobley et al., 2014). Nonetheless, this research demonstrated that specific biofilm formation triggers such as D-amino acids and norspermidine can destabilize the matrix polymer of the mature B. subtilis biofilm.
CONCLUSIONS
B. subtilis is an extremely convenient model organism for researching the diversity of the stationary-phase regulatory pathways because it is capable of surviving within a wide range of physical and chemical conditions, including extreme environments. Apart from the genetic information concerning spore formation, the genome of this bacterium contains instructions regarding the mechanisms of biofilm biosynthesis and dispersion. They are based on complex genetic programs that are responsible for creating a single integral association of the populations of genetically identical but phenotypically different cells, formal analogs of complex multicellular organisms (Kovacs and Dragos, 2019). In B. subtilis biofilms, a phenomenon was revealed that is similar to the kin selection effect (characteristic of higher organisms) and enhances the viability of filial cell generations. The bistability phenomenon that manifests itself in phenotypic alterations in the cells promotes the species’ adaptability because subpopulations interact within the community in a mutually beneficial fashion and a heterogeneous population can promptly respond to any environmental changes. In B. subtilis, an advanced network of regulatory systems involved in establishing a heterogeneous microbial community was identified. The signaling and regulatory pathways were elucidated that trigger biofilm formation. One of the pivotal events is the activation of transcription factor Spo0A that activates the expression of gene clusters encoding the biofilm matrix in response to a wide spectrum of extracellular signals. One of the peculiarities of biofilm-forming B. subtilis cells is the synthesis and secretion of toxic cannibalism peptides for eliminating the cells that do not synthesize the extracellular matrix polymers. Actual regulatory mechanisms may be more diverse, because, in nature, B. subtilis bacteria dwell in association with plant roots or otherwise in the presence of other organisms (Buzoleva et al., 2016; Yannarell et al., 2019; Hashem et al., 2019; Bóka et al., 2019). Therefore, there are still many unresolved issues regarding the development of B. subtilis biofilms. Integrated analysis of regulatory systems resulted in discovering new regulatory mechanisms in bacteria, including global protein acetylation that was described for the first time (Reverdy et al., 2018), or detecting specific areas called membrane functional microdomains that are equivalent to lipid rafts in the plasma membranes of eukaryotic cells (Lopez, 2015; Bramkamp et al., 2015; Lopez and Koch, 2017; Wagner et al., 2017). Signal transfer via regulatory cascades in bacilli involves microdomains that operate in conformity with the actual set of environmental factors. The structural organization of microdomains is characterized by an unexpected complexity level. This is an unprecedented fact in the bacterial realm, since bacteria have turned out to be more complex organisms than it was assumed earlier. Moreover, the data obtained indicate that membrane microdomains are widespread among bacteria. In contrast to the information on the regulation of biofilm formation mechanisms, our knowledge concerning the system of control over biofilm dispersal and dissociation in B. subtilis is still at the initial stage. The issues to be addressed are concerned with the synthesis of specific peptides and D-amino acids that are implicated in dispersing biofilms under various conditions. The external and internal stimuli that trigger this process are also still to be researched. This research is not only of theoretical but also of practical interest, and it is aimed, apart from other goals, at combating the biofilms formed by pathogenic organisms and developing efficient techniques for new combined therapeutic strategies.
REFERENCES
Aguilar, C., Vlamakis, H., Guzman, A., Losick, R., and Kolter, R., KinD is a checkpoint protein linking spore formation to extracellular-matrix production in Bacillus subtilis biofilms, MBio, 2010, vol. 1, p. e00035-10.
Aktuganov, G.E., Galimzyanova, N.F., Gil’vanova, E.A., Ryabova, A.S., Safina, V.R., Pudova, E.A., Kuz’mina, L.Yu., Boiko, T.F., and Melent’ev, A.I., Effect of carbon sources and mineral salts on biofilm formation and synthesis of cyclic lipopeptides by Bacillus and Paenibacillus PGPB strains, Ekobiotekh, 2019, vol. 2, pp. 468–481.
Aleti, G., Lehner, S., Bacher, M., Compant, S., Nikolic, B., Plesko, M., Schuhmacher, R., Sessitsch, A., and Brader, G., Surfactin variants mediate species-specific biofilm formation and root colonization in Bacillus, Environ. Microbiol., 2016, vol. 18, pp. 2634–2645.
Arnaouteli, S., Ferreira, A.S., Schor, M., Morris, R.J., Bromley, K.M., and Jo, J., Bifunctionality of a biofilm matrix protein controlled by redox state, Proc. Natl. Acad. Sci. U. S. A., 2017, vol. 114, pp. 6184–6191.
Asally, M., Kittisopikul, M., Rue, P., Du, Y., Hu, Z., and Cagatay, T., Localized cell death focuses mechanical forces during 3D patterning in a biofilm, Proc. Natl. Acad. Sci. U. S. A., 2012, vol. 109, pp. 18891–18896.
Balasubramanian, S., Aubin-Tam, M.E., and Meyer, A.S., 3D Printing for the fabrication of biofilm-based functional living materials, ACS Synth. Biol., 2019, vol. 8, pp. 1564–1567.
Bareia, T., Pollak, S., and Eldar, A., Self-sensing in Bacillus subtilis quorum-sensing systems, Nat. Microbiol., 2018, vol. 3, pp. 83–89.
Bartolini, M., Cogliati, S., Vileta, D., Bauman, C., Rateni, L., Leñini, C., Argañaraz, F., Francisco, M., Villalba, J.M., Steil, L., Völker, U., and Grau, R., Regulation of biofilm aging and dispersal in Bacillus subtilis by the alternative sigma factor SigB, J. Bacteriol., 2018, vol. 201, p. e00473-18.
Benigar, E., Dogsa, I., Stopar, D., Jamnik, A., Kralj Cigic, I., and Tomsic, M., Structure and dynamics of a polysaccharide matrix: aqueous solutions of bacterial levan, Langmuir, 2014, vol. 30, pp. 4172–4182.
Bóka, B., Manczinger, L., Kocsubé, S., Shine, K., Alharbi, N.S., Khaled, J.M., Münsterkötter, M., Vágvölgyi, C., and Kredics, L., Genome analysis of a Bacillus subtilis strain reveals genetic mutations determining biocontrol properties, World J. Microbiol. Biotechnol., 2019, vol. 35, p. 52.
Bramkamp, M. and Lopez, D., Exploring the existence of lipid rafts in bacteria. Version 2, Microbiol. Mol. Biol. Rev., 2015, vol. 79, pp. 81–100.
Branda, S.S., Chu, F., Kearns, D.B., Losick, R., and Kolter, R., A major protein component of the Bacillus subtilis biofilm matrix, Mol. Microbiol., 2006, vol. 59, pp. 1229–1238.
Branda S.S., Gonzalez-Pastor J.E., Ben-Yehuda S., Losick R., Kolter R. Fruiting body formation by Bacillus subtilis, Proc. Natl. Acad. Sci. U. S. A., 2001, vol. 98, pp. 11621–11626.
Brinsmade, S.R., Alexander, E.L., Livny, J., Stettner, A.I., Segre, D., Rhee, K.Y., and Sonenshein, A.L., Hierarchical expression of genes controlled by the Bacillus subtilis global regulatory protein CodY, Proc. Natl. Acad. Sci. U. S. A., 2014, vol. 111, pp. 8227–8232.
Brown, L., Kessler, A., and Casadevall, A., Vesicle isolation from Bacillus subtilis biofilm, Bio Protoc., 2015, vol. 5, p. e1409.
Brown, L., Kessler, A., Cabezas-Sanchez, P., Luque-Garcia, J.L., and Casadevall, A., Extracellular vesicles produced by the Gram-positive bacterium Bacillus subtilis are disrupted by the lipopeptide surfactin, Mol. Microbiol., 2014, vol. 93, pp. 183–198.
Buzoleva, L.S., Bogatyrenko, E.A., and Tsvetkova, N.B., Biofilm formation by different serological variants of Listeria monocytogenes in association with Bacillus pumilus, Microbiology (Moscow), 2016, vol. 85, pp. 311–316.
Cairns, L.S., Hobley, L., and Stanley-Wall, N.R., Biofilm formation by Bacillus subtilis: new insights into regulatory strategies and assembly mechanisms, Mol. Microbiol., 2014, vol. 93, pp. 587–598.
Cámara-Almirón, J., Navarro, Y., Díaz-Martínez, L., Magno-Pérez-Bryan, M.C., Molina-Santiago, C., Pearson, J.R., Vicente, A., Pérez-García, A., and Romero, D., Dual functionality of the amyloid protein TasA in Bacillus physiology and fitness on the phylloplane, Nat. Commun., 2020, vol. 11, p. 1859.
Chai, L., Romero, D., Kayatekin, C., Akabayov, B., Vlamakis, H., Losick, R., and Kolter, R., Isolation, characterization, and aggregation of a structured bacterial matrix precursor, J. Biol. Chem., 2013, vol. 288, pp. 17559–17568.
Chai, Y., Beauregard, P.B., Vlamakis, H., Losick, R., and Kolter, R., Galactose metabolism plays a crucial role in biofilm formation by Bacillus subtilis, MBio, 2012, vol. 3, p. e00184-12.
Chai, Y., Kolter, R., and Losick, R., Paralogous antirepressors acting on the master regulator for biofilm formation in Bacillus subtilis, Mol. Microbiol., 2009, vol. 74, pp. 876–887.
Chai, Y., Kolter, R., and Losick, R., Reversal of an epigenetic switch governing cell chaining in Bacillus subtilis by protein instability, Mol. Microbiol., 2010, vol. 78, pp. 218–229.
Chai, Y., Norman, T., Kolter, R., and Losick, R., Evidence that metabolism and chromosome copy number control mutually exclusive cell fates in Bacillus subtilis, EMBO J., 2011, vol. 30, pp. 1402–1413.
Chan, J.M., Guttenplan, S.B., and Kearns, D.B., Defects in the flagellar motor increase synthesis of polygamma-glutamate in Bacillus subtilis, J. Bacteriol., 2014, vol. 196, pp. 740–753.
Chastanet, A. and Losick, R., Just-in-time control of Spo0A synthesis in Bacillus subtilis by multiple regulatory mechanisms, J. Bacteriol., 2011, vol. 193, pp. 6366–6374.
Chen, Y., Cao, S., Chai, Y., Clardy, J., Kolter, R., Guo, J.H., and Losick, R., A Bacillus subtilis sensor kinase involved in triggering biofilm formation on the roots of tomato plants, Mol. Microbiol., 2012, vol. 85, pp. 418–430.
Chen, Y., Gozzi, K., Yan, F., and Chai, Y., Acetic acid acts as a volatile signal to stimulate bacterial biofilm formation, mBio, 2015, vol. 6, p. e00392.
Cheremin, M., Nyamsuren, Ch., Toimentseva, A.A., and Sharipova, M.R., Optimization of the expression of the subtilisin-like protease from Bacillus pumilus, Russ. J. Bioorg. Chem., 2014, vol. 40, pp. 694–698.
Chumsakul, O., Takahashi, H., Oshima, T., Hishimoto, T., Kanaya, S., Ogasawara, N., and Ishikawa S., Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation, Nucl. Acids. Res., 2011, vol. 39, pp. 414–428.
Dabbagh, F., Negahdaripour, M., Berenjian, A., Behfar, A., Mohammadi, F., Zamani, M., Irajie, C., and Ghasemi, Y., Nattokinase: production and application, Ap-pl. Microbiol. Biotechnol., 2014, vol. 98, pp. 9199–9206.
Danilova, Y.V., Shagimardanova, E.I., Margulis, A.B., Toymentseva, A.A., Balaban, N.P., Rudakova, N.L., Rizvanov, A.A., Sharipova, M.R., and Palotas, A., Bacterial enzymes effectively digest Alzheimer’s P-amyloid peptide, Brain Res. Bull., 2014, vol. 108, pp. 113–117.
DeLoughery, A., Dengler, V., Chai, Y., and Losick, R., Biofilm formation by Bacillus subtilis requires an endoribonuclease-containing multisubunit complex that controls mRNA levels for the matrix gene repressor SinR, Mol. Microbiol., 2016, vol. 99, pp. 425–437.
Devi, S.N., Kiehler, B., Haggett, L., and Fujita, M., Evidence that autophosphorylation of the major sporulation kinase in Bacillus subtilis is able to occur in trans, J. Bacterio-l., 2015, vol. 197, pp. 2675–2684.
Diehl, A., Roske, Y., Ball, L., Chowdhury, A., Hiller, M., Molière, N., Kramer, R., Stöppler, D., Worth, C.L., Schlegel, B., Leidert, M., Cremer, N., Erdmann, N., Lopez, D., Stephanowitz, H., et al., Structural changes of TasA in biofilm formation of Bacillus subtilis, Proc. Natl. Acad. Sci. U. S. A., 2018, vol. 115, pp. 3237–3242.
Dogsa, I., Brloznik, M., Stopar, D., and Mandic-Mulec, I., Exopolymer diversity and the role of levan in Bacillus subtilis biofilms, PLoS One, 2013, vol. 8, p. e62044.
Dragos, A., Kovacs, A.T., and Claessen, D., The role of functional amyloids in multicellular growth and development of Gram-positive bacteria, Biomolecules, 2017, vol. 7, p. 60.
Dragos, A., Kovacs, B.T., The peculiar functions of the bacterial extracellular matrix, Trends Microbiol., 2017, vol. 25, pp. 257–266.
Ellermeier, C.D., Hobbs, E.C., Gonzalez-Pastor, J.E., and Losick, R., A three-protein signaling pathway governing immunity to a bacterial cannibalism toxin, Cell, 2006, vol. 124, pp. 549–559.
Fuchs, F.M., Holland, G., Moeller, R., and Laue, M., Directed freeze-fracturing of Bacillus subtilis biofilms for conventional scanning electron microscopy, J. Microbiol. Methods, 2018, vol. 152, pp. 165–172.
Fu, H., Elena, R.C., and Marquez, P.H., The roles of small RNAs: insights from bacterial quorum sensing, ExRNA, 2019, vol. 1, p. 32.
Fujita, M., Gonzalez-Pastor, J.E., and Losick, R., High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis, J. Bacteriol., 2005, vol. 187, pp. 1357–1368.
Gabdrakhmanova, L., Vishniakov, I., Sharipova, M., Balaban, N., Kostrov, S., and Leshchinskaya, I., Salt stress induction of glutamyl endopeptidase biosynthesis in Bacillus intermedius, Microbiol. Res., 2005, vol. 160, pp. 233–242.
Gallegos-Monterrosa, R., Mhatre, E., and Kovács, A.T., Specific Bacillus Subtilis 168 variants form biofilms on nutrient-rich medium, Microbiology (SGM), 2016, vol. 162, pp. 1922–1932.
Gingichashvili, S., Duanis-Assaf, D., Shemesh, M., Featherstone, J.D.B., Feuerstein, O., and Steinberg, D., The adaptive morphology of Bacillus subtilis biofilms: a defense mechanism against bacterial starvation, Microorganisms, 2019, vol. 8, p. 62. https://doi.org/10.3390/microorganisms8010062
Gonzalez-Pastor, J.E., Cannibalism: a social behavior in sporulating Bacillus subtilis, FEMS Microbiol. Rev., 2011, vol. 35, pp. 415–424.
Grau, R.R., de Oña, P., Kunert, M., Leñini, C., Gallegos-Monterrosa, R., Mhatre, E., Vileta, D., Donato, V., Hölscher, T., Boland, W., Kuipers, O.P., and Kovács, Á.T., A duo of potassium-responsive histidine kinases govern the multicellular destiny of Bacillus subtilis, mBio, 2015, vol. 6, p. e00581.
Greenwich, J., Reverdy, A., Gozzi, K., Di Cecco, G., Tashjian, T., Godoy-Carter, V., and Chai, Y., A decrease in serine levels during growth transition triggers biofilm formation in Bacillus subtilis, J. Bacteriol., 2019, vol. 201, p. e00155-19.
Guttenplan, S.B. and Kearns, D.B., Regulation of flagellar motility during biofilm formation, FEMS Microbiol Rev., 2013, vol. 37, pp. 849–871.
Hamon, M.A., Stanley, N.R., Britton, R.A., Grossman, A.D., and Lazazzera, B.A., Identification of AbrB-regulated genes involved in biofilm formation by Bacillus subtilis, Mol. Microbiol., 2004, vol. 52, pp. 847–860.
Hashem, A., Tabassum, B., and Fathi Abd Allah, E., Bacillus subtilis: a plant-growth promoting rhizobacterium that also impacts biotic stress, Saudi J. Biol. Sci., 2019, vol. 26, pp. 1291–1297.
Hobley, L., Harkins, C., MacPhee, C.E., and Stanley-Wall, N.R., Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes, FEMS Microbiol. Rev., 2015, vol. 39, pp. 649–669.
Hobley, L., Kim, S.H., Maezato, Y., Wyllie, S., Fairlamb, A.H., Stanley-Wall, N.R., and Michael, A.J., Norspermidine is not a self-produced trigger for biofilm disassembly, Cell, 2014, vol. 156, pp. 844–854.
Hobley, L., Li, B., Wood, J.L., Kim, S.H., Naidoo, J., Ferreira, A.S., Khomutov, M., Khomutov, A., Stanley-Wall, N.R., and Michael, A.J., Spermidine promotes Bacillus subtilis biofilm formation by activating expression of the matrix regulator slrR, J. Biol. Chem., 2017, vol. 292, pp. 12041–12053.
Höfler, C., Heckmann, J., Fritsch, A., Popp, P., Gebhard, S., and Fritz, G., Cannibalism stress response in Bacillus subtilis, Microbiology (SGM), 2016, vol. 162, pp. 164–176.
Hollenbeck, E.C., Douarche, C., Allain, J.M., Roger, P., Regeard, C., Cegelski, L., Fuller, G.G., and Raspaud, E., Mechanical behavior of a Bacillus subtilis pellicle, J. Phys. Chem. B, 2016, vol. 120, pp. 6080–6088.
Huang, J., Liu, S., Zhang, C., Wang, X., Pu, J., Ba, F., Xue, S., Ye, H., Zhao, T., Li, K., Wang, Y., Zhang, J., Wang, L., Fan, C., Lu, T.K., and Zhong, C., Programmable and printable Bacillus subtilis biofilms as engineered living materials, Nat. Chem. Biol., 2018, vol. 15, pp. 34–41.
Irnov, I. and Winkler, W.C., A regulatory RNA required for antitermination of biofilm and capsular polysaccharide operons in Bacillales, Mol. Microbiol., 2010, vol. 76, pp. 559–575.
Jiang M., Shao, W., Perego, M., and Hoch, J.A., Multiple histidine kinases regulate entry into stationary phase and sporulation in Bacillus subtilis, Mol. Microbiol., 2000, vol. 38, pp. 535–542.
Jones, S.E., Paynich, M.L., Kearns, D.B., and Knight, K.L., Protection from intestinal inflammation by bacterial exopolysaccharides, J. Immunol., 2014, vol. 192, pp. 4813–4820.
Kalamara, M., Spacapan, M., Mandic-Mulec, I., and Stanley-Wall, N.R., Social behaviours by Bacillus subtilis: quorum sensing, kin discrimination and beyond, Mol. Microbiol., 2018, vol. 110, pp. 863–878.
Kampf, J., Gerwig, J., Kruse, K., Cleverley, R., Dormeyer, M., Grunberger, A., Kohlheyer, D., Commichau, F.M., Lewis, R.J., and Stulke, J., Selective pressure for biofilm formation in Bacillus subtilis: differential effect of mutations in the master regulator SinR on bistability, mBio, 2018, vol. 9, p. e01464-18.
Kayumov, A.R., Balaban, N.P., Mardanova, A.M., Sharipova, M.R., and Kostrov, S.V., Biosynthesis of the subtilisin-like serine proteinase of Bacillus intermedius under salt stress conditions, Microbiology (Moscow), 2006, vol. 75, pp. 557–562.
Kesel, S., Bronk, B., García, C.F., Götz, A., Lieleg, O., and Opitz, M., Matrix composition determines the dimensions of Bacillus subtilis NCIB 3610 biofilm colonies grown on LB agar, RSC Adv., 2017, vol. 7, pp. 31886–31898.
Khezri, M., Jouzani, G.S., and Ahmadzadeh, M., Fusarium culmorum affects expression of biofilm formation key genes in Bacillus subtilis, Braz. J. Microbiol., 2016, vol. 47, pp. 47–54.
Kobayashi, K. and Ikemoto, Y., Biofilm-associated toxin and extracellular protease cooperatively suppress competitors in Bacillus subtilis biofilms, PLoS Genet., 2019, vol. 15, p. e1008232.
Kobayashi, K. and Iwano, M., BslA (YuaB) forms a hydrophobic layer on the surface of Bacillus subtilis biofilms, Mol. Microbiol., 2012, vol. 85, pp. 51–66.
Kobayashi, K., Inactivation of cysL inhibits biofilm formation by activating the disulfide stress regulator Spx in Bacillus subtilis, J. Bacteriol., 2019, vol. 201. https://doi.org/10.1128/JB.00712-18
Kolodkin-Gal, I., Cao, S., Chai, L., Bottcher, T., Kolter, R., Clardy, J., and Losick, R., A self- produced trigger for biofilm disassembly that targets exopolysaccharide, Cell, 2012, vol. 149, pp. 684–692.
Kolodkin-Gal, I., Elsholz, A.K., Muth, C., Girguis, P.R., Kolter, R., and Losick, R., Respiration control of multicellularity in Bacillus subtilis by a complex of the cytochrome chain with a membrane-embedded histidine kinase, Genes Dev., 2013, vol. 27, pp. 887–899.
Kovacs, A.T. and Dragos, A., Evolved biofilm: review on the experimental evolution studies of Bacillus subtilis pellicles, J. Mol. Biol., 2019, vol. 431, pp. 4749–4759.
Lazarevic, V., Soldo, B., Medico, N., Pooley, H., Bron, S., and Karamata, D., Bacillus subtilis α-phosphoglucomutase is required for normal cell morphology and biofilm formation, Appl. Environ. Microbiol., 2005, vol. 71, pp. 39–45.
Leiman, S.A., May, J.M., Lebar, M.D., Kahne, D., Kolter, R., and Losick, R., D-amino acids indirectly inhibit biofilm formation in Bacillus subtilis by interfering with protein synthesis, J. Bacteriol., 2013, vol. 195, pp. 5391–5395.
Liu, W.T., Yang, Y.L., Xu, Y., Lamsa, A., Haste, N.M., Yang, J.Y., and Dorrestein, P.C., Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis, Proc. Natl. Acad. Sci. U. S. A., 2010, vol. 107, pp. 16286–16290.
Lopez, D. and Koch, G., Exploring functional membrane microdomains in bacteria: an overview, Curr. Opin. Microbiol., 2017, vol. 36, pp. 76–84.
Lopez, D. and Kolter, R., Extracellular signals that define distinct and coexisting cell fates in Bacillus subtilis, FEMS Microbiol. Rev., 2010, vol. 34, pp. 134–149.
Lopez, D., Molecular composition of functional microdomains in bacterial membranes, Chem. Phys. Lipids, 2015, vol. 192, pp. 3–11.
Lopez, D., Vlamakis, H., Losick, R., and Kolter, R., Cannibalism enhances biofilm development in Bacillus subtilis, Mol. Microbiol., 2009, vol. 74, pp. 609–618.
Lyons, N.A., Kraigher, B., Stefanic, P., Mandic-Mulec, I., and Kolter, R., A Combinatorial kin discrimination system in Bacillus subtilis, Curr. Biol., 2016, vol. 26, pp. 733–742.
Mammeri, N.E., Hierrezuelo, J., Tolchard, J., Cámara-Almirón, J., Caro-Astorga, J., Álvarez-Mena, A., Dutour, A., Berbon, M., Shenoy, J., Morvan, E., Grélard, A., Kauffmann, B., Lecomte, S., de Vicente, A., Habenstein, B., et al., Molecular architecture of bacterial amyloids in Bacillus biofilms, FASEB J., 2019, vol. 33, pp. 12146–12163.
Marlow, V.L., Cianfanelli, F.R., Porter, M., Cairns, L.S., Dale, J.K., and Stanley-Wall, N.R. The prevalence and origin of exoprotease-producing cells in the Bacillus subtilis biofilm, Microbiology (SGM), 2014a, vol. 160, pp. 56–66.
Marlow, V.L., Porter, M., Hobley, L., Kiley, T.B., Swedlow, J.R., Davidson, F.A., and Stanley-Wall, N.R., Phosphorylated DegU manipulates cell fate differentiation in the Bacillus subtilis biofilm, J. Bacteriol., 2014b, vol. 196, pp. 16–27.
Martin, M., Dragoš, A., Otto, S.B., Schäfer, D., Brix, S., Maróti, G., and Kovács, Á.T., Cheaters shape the evolution of phenotypic heterogeneity in Bacillus subtilis biofilms, ISME J., 2020. https://doi.org/10.1038/s41396-020-0685-4
Marvasi, M., Visscher, P.T., and Casillas Martinez, L., Exopolymeric substances (EPS) from Bacillus subtilis: polymers and genes encoding their synthesis, FEMS Microbiol. Lett., 2010, vol. 313, pp. 1–9.
McLoon, A.L., Kolodkin-Gal, I., Rubinstein, S.M., Kolter, R., and Losick, R., Spatial regulation of histidine kinases governing biofilm formation in Bacillus subtilis, J. Bacteriol., 2011, vol. 193, pp. 679–685.
Mielich-Suss, B. and Lopez, D., Molecular mechanisms involved in Bacillus subtilis biofilm formation, Environ. Microbiol., 2015, vol. 17, pp. 555–565.
Mielich-Suss, B., Schneider, J., and Lopez, D., Overproduction of flotillin influences cell differentiation and shape in Bacillus subtilis, MBio, 2013, vol. 4, p. e00719-00713.
Milton, M.E., Draughn, G.L., Bobay, B.G., Stowe, S.D., Olson, A.L., Feldmann, E.A., Thompson, R.J., Myers, K.H., Santoro, M.T., Kearns, D.B., and Cavanagh, J., The solution structures and interaction of SinR and SinI: elucidating the mechanism of action of the master regulator switch for biofilm formation in Bacillus subtilis, J. Mol. Biol., 2020, vol. 432, pp. 343–357.
Mitrofanova, O., Mardanova, A., Evtugyn, V., Bogomolnaya, L., and Sharipova, M., Effects of Bacillus serine proteases on the bacterial biofilms, Biomed. Res. Int., 2017, vol. 2017, p. 10.
Morikawa, M., Kagihiro, S., Haruki, M., Takano, K., Branda, S., Kolter, R., and Kanaya, S., Biofilm formation by a Bacillus subtilis strain that produces β-polyglutamate, Microbiology (SGM), 2006, vol. 152, pp. 2801–2807.
Nagorska, K., Ostrowski, A., Hinc, K., Holland, I.B., and Obuchowski, M., Importance of eps genes from Bacillus subtilis in biofilm formation and swarming, J. Appl. Genet., 2010, vol. 51, pp. 369–381.
Newman, J.A. and Lewis, R.J., Exploring the role of SlrR and SlrA in the SinR epigenetic switch, Commun. Integr. Bio-l., 2013, vol. 6, p. e25658.
Norman, T.M., Lord, N.D., Paulsson, J., and Losick, R., Memory and modularity in cell-fate decision making, Nature, 2013, vol. 503, pp. 481–486.
Ogura, M., Post-transcriptionally generated cell heterogeneity regulates biofilm formation in Bacillus subtilis, Genes Cells, 2016, vol. 21, pp. 335–349.
Ogura, M., Yoshikawa, H., and Chibazakura, T., Regulation of the response regulator gene degU through the binding of SinR/SlrR and exclusion of SinR/SlrR by DegU in Bacillus subtilis, J. Bacteriol., 2014, vol. 196, pp. 873–881.
Okada, M., Sato, I., Cho, S.J., Iwata, H., Nishio, T., Dubnau, D., and Sakagami, Y., Structure of the Bacillus subtilis quorum-sensing peptide pheromone ComX, Nat. Chem. Biol., 2005, vol. 1, pp. 23–24.
Ostrowski, A., Mehert, A., Prescott, A., Kiley, T.B., and Stanley-Wall, N.R., YuaB functions synergistically with the exopolysaccharide and TasA amyloid fibers to allow biofilm formation by Bacillus subtilis, J. Bacteriol., 2011, vol. 193, pp. 4821–4831.
Peng, N., Cai, P., Mortimer, M., Wu, Y., Gao, C., and Huang, Q., The exopolysaccharide–eDNA interaction modulates 3D architecture of Bacillus subtilis biofilm, BMC Microbiol., 2020, vol. 20, p. 115.
Penha, R.O., Vandenberghe, L., Faulds, C., Soccol, V.T., and Soccol, C.R., Bacillus lipopeptides as powerful pest control agents for a more sustainable and healthy agriculture: recent studies and innovations, Planta, 2020, vol. 251, p. 70.
Pisithkul, T., Schroeder, J.W., Trujillo, E.A., Yeesin, P., Stevenson, D.M., Chaiamarit, T., Coon, J.J., Wang, J.D., and Amador-Noguez, D., Metabolic remodeling during biofilm development of Bacillus subtilis, mBio, 2019, vol. 10, p. e00623-19.
Plakunov, V.K., Mart’yanov, S.V., Teteneva, N.A., and Zhurina, M.V., A universal method for quantitative characterization of growth and metabolic activity of microbial biofilms in static models, Microbiology (Moscow), 2016, vol. 85, pp. 509–513.
Plakunov, V.K., Mart’yanov, S.V., Teteneva, N.A., and Zhurina, M.V., Controlling of microbial biofilms formation: anti- and probiofilm agents, Microbiology (Moscow), 2017, vol. 86, pp. 423–438.
Pollak, S., Omer-Bendori, S., Even-Tov, E., Lipsman, V., Bareia, T., Ben-Zion, I., and Eldar, A., Facultative cheating supports the coexistence of diverse quorum-sensing alleles, Proc. Natl. Acad. Sci. U. S. A., 2016, vol. 113, pp. 2152–2157.
Ponomareva, A.L., Buzoleva, L.S., and Bogatyrenko, E.A., Abiotic environmental factors affecting the formation of microbial biofilms, Biol. Bull., 2018, vol. 45, pp. 490–496.
Raga-Carbajal, E., Carrillo-Nava, E., Costas, M., Porras-Dominguez, J., López-Munguía, A., and Olvera, C., Size product modulation by enzyme concentration reveals two distinct levan elongation mechanisms in Bacillus subtilis levansucrase, Glycobiology, 2016, vol. 26, pp. 377–385.
Reverdy, A., Chen, Y., Hunter, E., Gozzi, K., and Chai, Y., Protein lysine acetylation plays a regulatory role in Bacillus subtilis multicellularity, PLoS One, 2018, vol. 13, p. e0204687.
Romero, D., Vlamakis, H., Losick, R., and Kolter, R., Functional analysis of the accessory protein TapA in Bacillus subtilis amyloid fiber assembly, J. Bacteriol., 2014, vol. 196, pp. 1505–1513.
Römling, U. and Galperin, M.Y., Discovery of the second messenger cyclic di-GMP, Meth. Mol. Biol., 2017, vol. 1657, pp. 1–8.
Roux, D., Cywes-Bentley, C., Zhang, Y.-F., Pons, S., Konkol, M., and Kearns, D.B., Identification of poly-N-acetylglucosamine as a major polysaccharide component of the Bacillus subtilis biofilm matrix, J. Biol. Chem., 2015, vol. 290, pp. 19261–19272.
Ryan-Payseur, B.K. and Freitag, N.E., Bacillus subtilis biofilms: a matter of individual choice, mBio, 2018, vol. 9, p. e02339-18.
Schafer, H. and Turgay K., Spx, a versatile regulator of the Bacillus subtilis stress response, Curr. Genet., 2019, vol. 65, pp. 871–876.
Schultz, D., Coordination of cell decisions and promotion of phenotypic diversity in B. subtilis via pulsed behavior of the phosphorelay, Bioessays, 2016, vol. 38, pp. 440–445.
Sharipova, M., Balaban, N., Kayumov, A., Kirillova, Y., Mardanova, A., Gabdrakhmanova, L., Leshchinskaya, I., Rudenskaya, G., Akimkina, T., Safina, D., Demidyuk, I., and Kostrov S., The expression of the serine proteinase gene of Bacillus intermedius in Bacillus subtilis, Microbiol. Res., 2008, vol. 163, pp. 39–50.
Shemesh, M. and Chai, Y., A combination of glycerol and manganese promotes biofilm formation in Bacillus subtilis via histidine kinase KinD signaling, J. Bacteriol., 2013, vol. 195, pp. 2747–2754.
Shemesh, M., Kolter, R., and Losick, R., The biocide chlorine dioxide stimulates biofilm formation in Bacillus subtilis by activation of the histidine kinase KinC, J. Bacteriol., 2010, vol. 192, pp. 6352–6356.
Si, X., Quan, X., and Wu, Y., A small-molecule norspermidine and norspermidine-hosting polyelectrolyte coatings inhibit biofilm formation by multi-species wastewater culture, Appl. Microbiol. Biotechnol., 2015, vol. 99, pp. 10861–10870.
Spacapan, M., Danevčič, T., and Mandic-Mulec, I., ComX-induced exoproteases degrade ComX in Bacillus subtilis PS-216, Front. Microbiol., 2018, vol. 9, p. 105.
Stanley, N.R. and Lazazzera, B.A., Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-γ-DL-glutamic acid production and biofilm formation, Mol. Microbiol., 2005, vol. 57, pp. 1143–1158.
Stover, A.G. and Driks, A., Secretion, localization and antibacterial activity of tasA, a Bacillus subtilis spore-associated protein, J. Bacteriol., 1999, vol. 181, pp. 1664–1672.
Stowe, S.D., Olson, A.L., and Losick, R., Chemical shift assignments and secondary structure prediction of the master biofilm regulator, SinR, from Bacillus subtilis, J. Biomol. NMR Assign., 2014, vol. 8, pp. 155–158.
Strelkova, E.A., Pozdnyakova, N.V., Zhurina, M.V., Plakunov, V.K., and Belyaev, S.S., Role of the extracellular polymer matrix in resistance of bacterial biofilms to extreme environmental factors, Microbiology (Moscow), 2013, vol. 82, pp. 119–125.
Townsley, L., Yannarell, S.M., Huynh, T.N., Woodward, J.J., and Shank, E.A., Cyclic di-AMP acts as an extracellular signal that impacts Bacillus subtilis biofilm formation and plant attachment, mBio, 2018, vol. 9, p. e00341-18.
Toyofuku, M., Carcamo-Oyarce, G., Yamamoto, T., E-isenstein, F., Hsiao, C.C., Kurosawa, M., Gademann, K., Pilhofer, M., Nomura, N., and Eberl, L., Prophage-triggered membrane vesicle formation through peptidoglycan damage in Bacillus subtilis, Nat. Commun., 2017, vol. 8, p. 481.
Trejo, M., Douarche, C., Bailleux, V., Poulard, C., Mariot, S., Regeard, C., and Raspaud, E., Elasticity and wrinkled morphology of Bacillus subtilis pellicles, Proc. Natl. Acad. Sci. U. S. A., 2013, vol. 110, pp. 2011–2016.
Tucker, A.T., Bobay, B.G., Banse, A.V., Olson, A.L., Soderblom, E.J., and Moseley, M.A., A DNA mimic: the structure and mechanism of action for the antirepressor protein AbbA, J. Mol. Biol., 2014, vol. 426, pp. 1911–1924.
Vaccari, L., Molaei, M., Niepa, T.H.R., Lee, D., Leheny, R.L., and Stebe, K.J., Films of bacteria at interfaces, Adv. Colloid. Interface Sci., 2017, vol. 247, pp. 561–572.
Veening, J.W., Hamoen, L.W., and Kuipers, O.P., Phosphatases modulate the bistable sporulation gene expression pattern in Bacillus subtilis, Mol. Microbiol., 2005, vol. 56, pp. 1481–1494.
Veening, J.W., Igoshin, O.A., Eijlander, R.T., Nijland, R., Hamoen, L.W., and Kuipers, O.P., Transient heterogeneity in extracellular protease production by Bacillus subtilis, Mol. Syst. Biol., 2008, vol. 4, p. 184.
Veening, J.W., Kuipers, O.P., Brul, S., Hellingwerf, K.J., and Kort, R., Effects of phosphorelay perturbations on architecture, sporulation, and spore resistance in biofilms of Bacillus subtilis, J. Bacteriol., 2006, vol. 188, pp. 3099–3109.
Velmourougane, K., Prasanna, R., and Saxena, A.K., Agriculturally important microbial biofilms: present status and future prospects, J. Basic Microbiol., 2017, vol. 57, pp. 548–573.
Verdugo-Fuentes, A., Gastélum, G., Rocha, J., and Torre, M., Multiple and overlapping functions of quorum sensing proteins for cell specialization in Bacillus species, J. Bacteriol., 2020, vol. 202, p. e00721-19.
Vidakovic, L., Singh, P.K., Hartmann, R., Nadell, C.D., and Drescher, K., Dynamic biofilm architecture confers individual and collective mechanisms of viral protection, Nat. Microbiol., 2018, vol. 3, pp. 26–31.
Vlamakis, H., Chai, Y., Beauregard, P., Losick, R., and Kolter, R., Sticking together: building a biofilm the Bacillus subtilis way, Nat. Rev. Microbiol., 2013, vol. 11, pp. 157–168.
Wagner, R.M., Kricks, L., and Lopez, D., Functional membrane microdomains organize signaling networks in bacteria, J. Membr. Biol., 2017, vol. 250, pp. 367–378.
Wang, X., Meng, S., and Han, J., Morphologies and phenotypes in Bacillus subtilis biofilms, J. Microbiol., 2017, vol. 55, pp. 619–627.
Winkelman, J.T., Bree, A.C., Bate, A.R., Eichenberger, P., Gourse, R.L., and Kearns, D.B., RemA is a DNA-binding protein that activates biofilm matrix gene expression in Bacillus subtilis, Mol. Microbiol., 2013, vol. 88, pp. 984–997.
Wu, Q., Xu, H., Liang, J., and Yao, J., Contribution of glycerol on production of poly(γ-glutamic acid) in Bacillus subtilis NX-2, Appl. Biochem. Biotechnol., 2010, vol. 160, pp. 386–392.
Wu, Y., Quan, X., Si, X., and Wang, X., A small molecule norspermidine in combination with silver ion enhances dispersal and disinfection of multi-species wastewater biofilms, Appl. Microbiol. Biotechnol., 2016, vol. 100, pp. 5619–5629.
Yannarell, S.M., Grandchamp, G.M., Chen, S.Y., Daniels, K.E., and Shank, E.A., A dual-species biofilm with emergent mechanical and protective properties, J. Bacteriol., 2019, vol. 201. p. e00670-18.
Yepes, A., Schneider, J., Mielich, B., Koch, G., Garcia-Betancur, J.C., and Ramamurthi, K.S., The biofilm formation defect of a Bacillus subtilis flotillin-defective mutant involves the protease FtsH, Mol. Microbiol., 2012, vol. 86, pp. 457–471.
Yu, C., Li, X., Zhang, N., Wen, D., Liu, C., and Li, Q., Inhibition of biofilm formation by D-tyrosine: effect of bacterial type and D-tyrosine concentration, Water Res., 2016, vol. 92, pp. 173–179.
Yu, Y., Yan, F., He, Y., Qin, Y., Chen, Y., Chai, Y., and Guo, J.H., The ClpY-ClpQ protease regulates multicellular development in Bacillus subtilis, Microbiology (SGM), 2018, vol. 164, pp. 848–862.
Zeriouh, H., de Vicente, A., Perez-Garcia, A., and Romero, D., Surfactin triggers biofilm formation of Bacillus subtilis in melon phylloplane and contributes to the biocontrol activity, Environ. Microbiol., 2014, vol. 16, pp. 2196–2211.
Zwick, J.V., Noble, S., Ellaicy, Y.K., Coe, G.D., Hakey, D.J., King, A.N., Sadauskas, A.J., and Faulkner, M.J., AhpA is a peroxidase expressed during biofilm formation in Bacillus subtilis, MicrobiologyOpen, 2017, vol. 6, p. e00403.
Funding
This work was sponsored in terms of the state program aimed at increasing the competitiveness of Kazan’ (Volga Region) Federal University among the world leading research and education centers and also supported by the Russian Science Foundation (project no. 16-16-04062) and the Russian Foundation for Basic Research (project no. 19-08-00853).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This article does not contain any studies involving animals performed by any of the authors.
CONFLICT OF INTERESTS
The authors declare that they have no conflict of interests.
Additional information
Translated by A. Oleskin
Rights and permissions
About this article
Cite this article
Sharipova, M.R., Mardanova, A.M., Rudakova, N.L. et al. Bistability and Formation of the Biofilm Matrix as Adaptive Mechanisms during the Stationary Phase of Bacillus subtilis . Microbiology 90, 20–36 (2021). https://doi.org/10.1134/S002626172006017X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S002626172006017X