Abstract—
The major substrates for methanogenesis were used for investigation of cultured methanogenic archaea from coastal methane seeps near the Tarkhankut Peninsula, Black Sea. Analysis of the 16S rRNA gene sequences revealed that growth of the classical methanogenic Euryarchaeota occurred in all enrichments but was absent in the controls without the substrates. Enrichments from the seep differed in microbial composition from those from the background point. The most numerous archaea belonged to the genera Methanolobus (medium with methanol and hydrogen), Methanosarcina (trimethylamine and hydrogen), Methanococcoides (trimethylamine), and Methanococcus (hydrogen and CO2). Syntrophic growth of hydrogenotrophic archaea of the genus Methanogenium with clostridia and members of the family Thermotogaceae probably occurred in enrichments with acetate. Relatively low similarity of the recovered 16S rRNA gene sequences with the closest cultured relatives (94% and lower) indicated that the Methanogenium phylotype belonged to a new species. The same was true for the Methanosarcina phylotype revealed in the culture with trimethylamine and hydrogen (97% and less similarity of the 16S rRNA gene sequences to those of the closest cultured relatives).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Release of hydrocarbon gases from marine bottom sediments (methane seeps) is an extremely interesting geological phenomenon that has been increasingly attracting researchers’ attention over the recent decades. Such gas seepage sites frequently represent sea bottom life oases that differ dramatically from the neighboring sediments in their physicochemical and microbiological characteristics and in the structure of benthic communities (Bernardino et al., 2012). High methane concentrations promote the growth of aerobic methanotrophic bacteria, which consume oxygen and produce additional organic matter that can enter the trophic chains (Ding and Valentine, 2008). The processes of extensive microbial methane oxidation and organic matter transformation are accompanied by a dramatic decrease in the concentrations of oxygen, which is present only in the uppermost sediment layers of methane seeps. Oxygen depletion and high activity of primary degraders lead to activation of sulfate-reducing and methanogenic microorganisms, which play a leading role in the terminal phase of organic matter degradation in marine bottom sediments (Kallistova et al., 2017).
Gas seeps in the Black Sea were discovered in the late 1980s (Polikarpov and Egorov, 1989). By present, researchers have been investigating methane seeps located mainly on the continental slope of the Northwestern Black Sea shelf, as well as deep-water mud volcanoes (Michaelis et al., 2002; Pimenov and Ivanova, 2005). A recent geoacoustic study of the coastal area of the Crimean Peninsula demonstrated that shallow-water methane seeps are commonly present at depths of up to 10 m (Egorov et al., 2011). The methane seeps discovered more than 30 years ago in the Kalamit Bay of the Black Sea near the coast of the Tarkhankut Peninsula are of particular interest. It was found that the maximal depth of oxygen penetration within bottom sediments at gas seep sites did not exceed several millimeters, while sulfide concentrations in subsurface sediments could reach 3 mM (Gulin et al., 2010). Previously, we showed that the local methanogenic communities were dominated by archaea of the families Methanosarcinaceae and Methanomicrobiaceae (Tarnovetskii et al., 2018). Since metagenomic studies using high-throughput sequencing cannot always provide an adequate estimate of the metabolic potential of microbial communities (Vigneron et al., 2015), we undertook a comprehensive investigation of methanogen diversity in coastal seeps of the Tarkhankut Peninsula by combining culturing techniques with molecular analysis.
The main goal of this work was to investigate the microbial potential of gas seeps of the Tarkhankut Peninsula and its development depending on the presence of different methanogenic substrates in the environment. For this purpose, we set up enrichment cultures with selective conditions promoting the growth of different methanogen groups.
MATERIALS AND METHODS
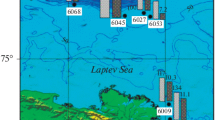
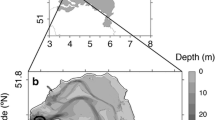
Sampling. The study analyzed bottom sediments of the Tarkhankut Peninsula gas seeps (45°35549 N, 32°73153 E) collected at the depth of 5 m. Samples were collected by a diver into plastic containers in May 2016. For enrichment cultures, the surface layer of bottom sediments (0‒10 cm) was used. On the shore, the sediment samples were thoroughly mixed and dispensed into 250-mL glass vials without leaving an air bubble, sealed with rubber stoppers, and transported into the laboratory.
Samples were collected at two sites: at the center of a methane seep and at a reference point located approximately 100 m away from the seep. The sediments were composed of aleuropelitic muds with an admixture of sand and detritus material in the center of the seep and of slightly silted sand at the reference site. Water for sample dilution was collected above the surface of the seep and the reference site using a bathometer.
Enrichment cultures. To start a culture, 10 mL of bottom sediments as inoculation material and 10 mL sea water were placed into a 60-mL vial. The vials were sealed with gas-tight rubber stoppers. Immediately after that, the air phase was removed from the vials and replaced with nitrogen. Selective growth conditions were created by adding different methanogenic substrates: trimethylamine (to the final concentration of 22.5 mM), acetate (15 mM), a mixture of carbon dioxide and hydrogen (85 and 15% of the gas phase, respectively), methanol (20 mM) and hydrogen (15%), or trimethylamine (22.5 mM) and hydrogen (15%). In the control cultures, no substrate was added. Samples from both sites were tested using the same assortment of substrates. Altogether, 12 enrichment cultures were started. The cultures were incubated in a thermostat at 20°C for 2 months. The growth of methanogens was assessed based on methane content in the vials, which was measured at the end of the incubation period using a Kristall gas chromatograph (Chromatec, Russia).
Phylogenetic composition of enrichment cultures determined using high-throughput sequencing of the 16S rRNA genes. DNA was isolated from the inocula (bottom sediment samples) and from enrichment cultures grown for 2 months. The samples or cultures were thoroughly mixed, and 5-mL aliquots were collected into clean tubes and centrifuged for 20 min at 14 000 g. The sedimented cells were homogenized mechanically and chemically (Lever et al., 2015). For this purpose, the cell pellet was mixed with guanidine hydrochloride and Triton X-100 (to the final concentrations of 800 mM and 0.5%, respectively) with a total volume of 500 µL and a mixture of glass beads 425–600 µm and 106 µm in diameter with a total volume of ~500 µL. The mixture was homogenized for 40 s at 6 m/s using a FastPrep®-24 Instrument (MP Biomedicals, United States) and incubated for 50 min at 50°C. Next, nucleic acids were extracted using the standard phenol‒chloroform procedure and precipitated with isopropanol. Libraries of DNA sequences representing the V3–V4 region of the 16S rRNA gene were obtained as described previously (Fadrosh et al., 2014). The fragments were amplified using the following primer system: the forward primer (5'-CAAGCAGAAGACGGCATACGAGATGTGACTGGAG-TTCAGACGTG-TGCTCTTCCGATCT XXXXXX-XXXXXX ZZZZ CCTAYGGGDBGCWSCAG-3') was composed of 5' Illumina Linker Sequence, Index 1, Heterogeneity Spacer (Fadrosh et al., 2014), and the Pro-mod-341F primer sequence (Merkel et al., 2017); the reverse primer (5'-AATGATACGGCGACCACCGAGATCTACACT-CTTTCCCTACACGACGC-TCTTCCGATCT XXXX-XXXXXXXX ZZZZ GACTACNVGGGTMTCTAATCC-3') included 3' Illumina Linker Sequence, Index 2, Heterogeneity Spacer, and Pro-mod-805R primer sequence (Merkel et al., 2017). The amplicons were separated by electrophoresis in 2% agarose gel, excised with a scalpel, and purified using the Cleanup Standard kit (Evrogen, Russia). Sequencing was performed in a MiSeq system (Illumina, United States) using the reagent kit that enabled the reading of 300 bases at either end of the amplicon. Demultiplexing was performed with the relevant software scripts of QIIME v. 1.9.0 (Caporaso et al., 2010). Subsequent sequence editing and analysis were also performed with QIIME v. 1.9.0. The data were filtered to select those with the minimal read quality score of 30 and the minimal read length of 350 bp. Chimeric reads were identified using the identify_chimeric_seqs.py script with the USEARCH v. 6.1544 algorithm (Edgar, 2010) and the Silva 123 database of reference 16S rRNA reads (Quast et al., 2013). The table of operational taxonomic units (OTUs) was constructed using the pick_open_reference_otus.py script. Sequences with an identity level of at least 97% were grouped into OTUs (Schloss and Handelsmann, 2006) using USEARCH v. 6.1544 (Edgar, 2010) and the Silva 123 database (Quast et al., 2013). Representative sequences were selected using UCLAST (Edgar, 2010). Alpha diversity analysis and construction of rarefaction curves were performed using the core_diversity_analyses.py script to obtain a normalized sample of 17 000 reads per specimen. The species of methanogenic archaea were identified using the BLAST online service. To calculate the alpha diversity parameters, the table of OTUs was normalized by cumulative sum scaling (CSS) technique using the QIIME script.
The obtained arrays of sequencing data were deposited into the NCBI database under the accession no. PRJNA549749.
RESULTS
Methane production. At the start of enrichment cultures, the gas phase was replaced with nitrogen, so it may be assumed that the initial methane content in the gas phase was zero. After 2 months of cultivation, the methane content in the gas phase ranged from 8.74 to 64.7%. In the control variants without substrate addition, methane was detected in trace amounts: 0.01 and 0.03% for the seep and the reference sample, respectively. The highest methane content was observed in enrichment cultures with methylated substrates: (1) trimethylamine, 33.3 and 14.8% for the seep and the reference sample, respectively; (2) trimethylamine and hydrogen, 64.7 and 51.8% for the seep and the reference sample, respectively; and (3) methanol and hydrogen, 30.4 and 20.2% for the seep and the reference sample, respectively. In the cultures grown on substrates of acetoclastic and hydrogenotrophic methanogenesis, methane levels were lower. The cultures from the seep and the reference site produced 8 and 8.8% methane, respectively, when grown on acetate and 14.7 and 8.7%, respectively, when grown on hydrogen and carbon dioxide.
Molecular analysis of the composition of microbial communities. Altogether, 427 501 16S rRNA gene sequences were obtained with an average length of 406 bp. The phylotype diversity coverage calculated using the Chao1 approach (Chao, 1984) ranged from 55.7 to 99.8% in enrichment cultures from the seep and from 51.7 to 92.9% in the reference cultures. The Shannon diversity index ranged from 3 to 9.5 (Table 1).
The obtained reads were grouped into OTUs with an identity level of 97%. Altogether, 10 437 unique OTUs were obtained. In the cultures from the reference site, most sequences were classified as archaeal. Their relative abundance was 49.6–85.5% (Table 1). In enrichment cultures from the seep, the share of archaea was lower: 76.5% in the culture grown on trimethylamine and 19.2–36% in other variants. In the inocula, archaea constituted only 2.7% in the seep sample and less than 1% in the reference sample. In the control cultures without added substrates, the share of archaea was 2.3% for the seep and 18.4% for the reference site.
In nearly all cultures, the dominant groups were known methanogens of the phylum Euryarchaeota. In enrichment cultures from seep bottom sediments, most reads represented the following microbial groups: uncultured clostridia (17.1 and 7.3%) and Methanococcus (7.3%) in variant 1 (hydrogen and CO2); Methanogenium (27.7%) in variant 2 (acetate); Methanococcoides (71.1%) in variant 3 (trimethylamine); Gracilibacteria (15.8%) and Methanosarcina (13.4%) in variant 4 (trimethylamine and hydrogen), and Methanolobus (19.5%) in variant 5 (methanol) (Table 2). In the inoculum from the seep, the dominant groups were bacteria of the taxa Gelria (8.2%), Nocardioides (4.1%), and uncultured Firmicutes (5.3%), and the control variant without added substrate was dominated by bacteria of the genus Sulfurimonas (70.5%).
All enrichment cultures from the reference site, except for variants 1 (methanol and hydrogen) and 2 (hydrogen and CO2), were dominated by archaea of the genus Methanococcoides (34.9–80.8%). In the culture grown on methanol and hydrogen, the most abundant were members of the genus Methanolobus (24.8%), and in the culture grown on hydrogen and CO2, it was archaea of the genus Methanogenium (49.4%). In the inoculum from the reference site, the largest groups were bacteria of the taxa Defluviicoccus (16.5%), Acidimicrobiales OM1 clade (7.2%), and Rhodospirillaceae (6.3%). In the control variant with no substrate added, the most abundant group were archaea of the phylum Woesearchaeota (10.5%), while 14.4% of reads could not be attributed to any known taxon.
DISCUSSION
The fact that methane production was observed in all enrichment cultures, but not in the control variants indicates that the microbial community studied is capable of methanogenesis using a broad range of substrates. Under natural conditions, methane production is limited by the amount of substrate available. The highest methane concentrations were reached on media that contained trimethylamine and methanol. Most probably, this was due to reaction stoichiometry and a larger energy gain achieved by methanogens during growth on methylated substrates. These results may also suggest that the community is dominated by microorganisms that employ the methylotrophic methanogenesis pathway, which could explain the low activity levels observed in our previous experiments with labeled carbon dioxide and acetate (unpublished data). Acetoclastic and hydrogenotrophic methanogenesis could be suppressed due to growth of sulfate-reducing bacteria. However, in all experiments the relative abundance of sulfate-reducing bacteria (2.3–9.3%) was lower than that of methanogenic archaea (15.9–82.5%). Therefore, it may be concluded that all substrates were added in large excess, and the competition between methanogens and sulfate-reducers need not be taken into account.
By analyzing the 16S rRNA gene sequences, we identified the microbial groups that predominated on each substrate. In most cases, these were one or two genera of classic methanogens. Importantly, the dominant microorganisms were different in the cultures obtained from the seep and the reference sample. The only exception were the cultures growing on methanol and hydrogen. These results indicate that the microbial communities of the seep and the reference site differ significantly in their potential.
In contrast to our previous work (Tarnovetskii et al., 2018), in this study we were able to obtain longer sequencing reads that spanned over two variable regions of the 16S rRNA gene. As a result, it was possible not only to analyze the composition of the communities at the genus level, but also to identify the species of microorganisms in the enrichment cultures.
The enrichment culture from the seep sample growing on acetate was dominated by sequences representing the genus Methanogenium. Based on the analyzed 16S rRNA gene fragment, the closest cultured species were M. frigidum and M. marinum classified as hydrogenotrophic psychrophilic methanogens (Franzman et al., 1997; Chong et al., 2002). It should be noted that the relationship to M. frigidum was remote (94% identity, Table 2). Most probably, the detected phylotype represents a novel species. This is an interesting result, because only two genera of methanogens are currently known to produce methane from acetate: Methanosaeta and Methanosarcina (Lyu and Liu, 2018). Acetate can also be oxidized to carbon dioxide and hydrogen by bacteria. This process as such does not provide an energy gain. However, if hydrogen is consumed by methanogens, the equilibrium is shifted and the reaction creates an energy gain (Hattori, 2008). Several group of bacteria are known to participate in syntrophic oxidation of acetate: members of the phyla Firmicutes (Thermacetogenium, Clostridium, Thermotoga, Candidatus Contubernalis, Candidatus Syntrophonatronum and Syntrophaceticus) and Proteobacteria (Desulfomicrobium, Geobacter) (Hattori, 2008; Westerholm et al., 2010; Kimura et al., 2013; Li et al., 2018; Timmers et al., 2018). In our enrichment cultures growing on acetate, none of bacterial species was clearly dominant. The highest relative abundances were observed for members of the taxa Thermotogaceae (3.5%), GoM-GC232-4463-Bac (order Clostridiales, 6.2%), livecontrolB21 (Clostridiales, 2.1%), Rhodobacteraceae (3.5%), and Desulfobacter (3.9%). Most probably, in the enrichment culture from the seep sample, typical hydrogenotrophic archaea produce methane in syntrophic association with acetate-oxidizing bacterial partners. Clostridia seem likely candidates for this role. Altogether, the order Clostridiales accounted for 14% of the reads. Bacteria of the family Thermotogaceae may also be involved in this process. Previously, it was found that Thermotogaceae and hydrogenotrophic methanogens dominated in enrichment cultures that utilized long-chain alkanes and acetate (Cheng et al., 2013; Li et al., 2018).
The observed active growth of Methanococcoides methylutens and M. alaskense in the reference cultures growing on acetate remains unexplained. Archaea of this genus are obligate methylotrophic methanogens. None of the known Methanococcoides species can utilize acetate for methane production. Members of this genus are widely present in marine ecosystems, including methane seeps (Lanoil et al., 2005; Omoregie et al., 2009; Roussel et al., 2009; Lazar et al., 2011; Lin et al., 2010; Vigneron et al., 2014). Usually, they can be detected even without preliminary cultivation under selective conditions (Orphan et al., 2001; Pachiadaki et al., 2011). Thanks to their ability to utilize methylated substrates, these methanogens can actively grow at high sulfate concentrations without competitive pressure from sulfate reducers. It is known that trimethylamine is an available substrate in marine ecosystems. In bottom sediments, it can be produced from lignin, pectin, betaine, choline, and trimethylamine-N-oxide (TMAO). Betaine and TMAO occur as osmolytes in many marine organisms. It was shown that some sulfate-reducing deltaproteobacteria of the genera Desulfovibrio, Desulfobacterium, and Desulfuromonas can transform glycine and betaine into trimethylamine (Hayward and Stadtman, 1959; Fiebig and Gottschalk, 1983; Heijthuijsen and Hansen, 1989). Some members of Methanococcoides can produce methane from choline and betaine without a bacterial partner (Watkins et al., 2014; L’Haridon et al., 2014).
The seep enrichment culture growing on trimethylamine and hydrogen produced large amounts of methane (33%). Some reads in this culture were classified into the genus Methanosarcina (13.2%), the members of which can produce methane via the hydrogenotrophic, acetoclastic, and methylotrophic pathways. However, the dominant group were bacteria of the phylum Gracilibacteria (15.8%). The first genome of Gracilibacteria was obtained from a thermal spring sample using single-cell sequencing (Rinke et al., 2013). Subsequently, two further genomes of Gracilibacteria were obtained from human mouth cavity. Members of this phylum can ferment glucose to acetate and are auxotrophic by many amino acids (Espinoza et al., 2018). The role of Gracilibacteria in methane production remains unknown. In our experiment, nearly all reads (886 of 908) belonged to the same OTU; at the same time, the abundance of Gracilibacteria in the inoculation material was below the detection threshold (0 reads). This suggests high levels of selective pressure and microbial specialization.
In comparison to methylotrophs, hydrogenotrophic methanogens were less diverse. In the cultures of samples from the seep and reference sites grown on CO2 and hydrogen, the dominant OTUs were the archaea Methanococcus maripaludis (seep) and Methanogenium marinum (reference site). Both these genera include typical hydrogenotrophic methanogens that produce methane by utilizing hydrogen or formate and carbon dioxide. M. maripaludis is a mesophilic microorganism first isolated from salt marsh sediments (Jones et al., 1983). In the bottom sediments of Tarkhankut rich in organic matter, a possible source of hydrogen are numerous heterotrophic microorganisms described in our previous study (Tarnovetskii et al., 2018). Probably, hydrogenotrophic methanogenesis takes place in the subsurface layers depleted in sulfate. The enrichment culture from the seep sample also contained abundant uncultured clostridia (livecontrolB21 and vadinBB60 group); however, nothing is currently known about their physiology.
In the control seep culture (without substrate addition), the highest relative abundance was observed for bacteria of the genus Sulfurimonas and archaea of the phylum Woesearchaeota. Initially, Sulfurimonas were considered human and animal pathogens. Since molecular techniques were introduced into microbial ecology, these microorganisms have been detected in thermal springs, marine, and land ecosystems. Currently, four cultured species of Sulfurimonas are known (Han and Perner, 2015). Based on the 16S rRNA gene sequence, the bacterium detected in our experiments exhibited little similarity to any of them, with only 96% homology to the nearest cultured relative. Members of the genus Sulfurimonas can utilize sulfur compounds as electron donors. This type of metabolism is quite plausible for microorganisms of seep bottom sediments.
Woesearchaeota is a phylum of uncultured archaea belonging to the superphylum DPANN. Metagenomic studies showed that this group of archaea has a high level of phylogenetic diversity. Presumably, they are heterotrophs growing in anoxic environments. It is possible that some members of Woesearchaeota can be engaged in syntrophic relationships with methanogens, providing them with acetate and hydrogen (Castelle et al., 2015; Liu et al., 2018).
Thus, our study demonstrated a high potential of the methanogenic community of coastal methane seeps of the Tarkhankut Peninsula. It was found that the microbial composition of bottom sediments differed significantly between the seep and the reference site. The enrichment cultures grown from the collected samples contained methanogens capable of producing methane via the hydrogenotrophic and methylotrophic pathways. Methane production from acetate occurred most probably via the hydrogenotrophic pathway thanks to syntrophy between methanogens and bacteria of the order Clostridiales and the family Thermotogaceae.
REFERENCES
Bernardino, A.F., Levin, L.A., Thurber, A.R., and Smith, C.R., Comparative composition, diversity and trophic ecology of sediment macrofauna at vents, seeps and organic falls, PLoS One, 2012, vol. 7, p. e33515.
Caporaso, J.G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F.D., Costello, E.K., Fierer, N., Peña, A.G., Goodrich, J.K., Gordon, J.I., Huttley, G.A., Kelley, S.T., Knights, D., Koenig, J.E., Ley, R.E., et al., QIIME allows analysis of high-throughput community sequencing data, Nat. Methods, 2010, vol. 7, pp. 335–336.
Castelle, C.J., Wrighton, K.C., Thomas, B.C., Hug, L.A., Brown, C.T., Wilkins, M.J., Frischkorn, K.R., Tringe, S.G., Singh, A., Markillie, L.M., Taylor, R.C., Williams, K.H., and Banfield, J.F., Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling, Curr. Biol., 2015, vol. 25, pp. 690–701.
Chao, A., Nonparametric estimation of the number of classes in a population, Scand. J. Statistics, 1984, vol. 11, pp. 265–270.
Cheng, L., Rui, J., Li, Q., Zhang, H., and Lu, Y., Enrichment and dynamics of novel syntrophs in a methanogenic hexadecane-degrading culture from a Chinese oilfield, FEMS Microbiol. Ecol., 2013, vol. 83, pp. 757–766.
Chong, S.C., Liu, Y., Cummins, M., Valentine, D.L., and Boone, D.R., Methanogenium marinum sp. nov., a H2-using methanogen from Skan Bay, Alaska, and kinetics of H2 utilization, Antonie Van Leeuwenhoek., 2002, vol. 81, pp. 263–270.
Ding, H. and Valentine, D., Methanotrophic bacteria occupy benthic microbial mats in shallow marine hydrocarbon seeps, Coal Oil Point, California, J. Geophys. Res., 2008, vol. 113, no. G1, p. G01015.
Edgar R.C., Search and clustering orders of magnitude faster than BLAST, Bioinform., 2010, vol. 26, pp. 2460–2461.
Egorov, V.N., Artemov, Yu.G., and Gilin, S.B., Metanovye sipy v Chernom more: sredoobrazuyushchaya i ekologichesk-aya rol’ (Black Sea Methane Seeps: Environment-Forming and Ecological Role), Sevastopol: EKOSI-Gidrofizika, 2011.
Espinoza, J.L., Harkins, D.M., Torralba, M., Gomez, A., Highlander, S.K., Jones, M.B., Leong, P., Saffery, R., Bockmann, M., Kuelbs, C., Inman, J.M., Hughes, T., Craig, J.M., Nelson, K.E., and Dupont, C.L., Supragingival plaque microbiome ecology and functional potential in the context of health and disease, MBio, 2018, vol. 9. e01631-18.
Fadrosh, D.W., Ma, B., Gajer, P., Sengamalay, N., Ott, S., Brotman, R.M., and Rave, l J., An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform, Microbiome, 2014, vol. 2, no. 1, p. 6.
Fiebig, K. and Gottschalk, G., Methanogenesis from choline by a coculture of Desulfovibrio sp. and Methanosarcina barkeri,Appl. Environ. Microbiol., 1983, vol. 45, pp. 161–168.
Franzmann, P.D., Liu, Y., Balkwill, D.L., Aldrich, H.C., Conway de Macario, E., and Boone, D.R., Methanogenium frigidum sp. nov., a psychrophilic, H2-using methanogen from Ace Lake, Antarctica, Int. J. Syst. Bacteriol., 1997, vol. 47, pp. 1068–1072.
Gulin, M.B., Timofeev, V.A., and Bondarenko, L.V., Zoobenthos in the microbiotopes of methane seeps in the shelf zone of the Crimean coast, Sist. Kontr. Okr. Sredy, 2010, no. 14, pp. 225–229.
Han, Y. and Perner, M., The globally widespread genus Sulfurimonas: versatile energy metabolisms and adaptations to redox clines, Front. Microbiol., 2015, vol. 6, article 989. https://doi.org/10.3389/fmicb.2015.00989
Hattori, S., Syntrophic acetate-oxidizing microbes in methanogenic environments, Microbes Environ., 2008, vol. 23, pp. 118–127.
Hayward, H.R. and Stadtman, T.C., Anaerobic degradation of choline. I. Fermentation of choline by an anaerobic, cytochrome-producing bacterium, Vibrio cholinicus n. sp., J. Bacteriol., 1959, vol. 78, pp. 557–561.
Hedlund, B.P., Dodsworth, J.A., Murugapiran, S.K., Rinke, C., and Woyke, T., Impact of single-cell genomics and metagenomics on the emerging view of extremophile “microbial dark matter,” Extremophiles, 2014, vol. 18, pp. 865–875.
Heijthuijsen, J.H.F.G. and Hansen, T.A., Betaine fermentation and oxidation by marine Desulfuromonas strains, Ap-pl. Environ. Microbiol., 1989, vol. 55, pp. 965–969.
Jones, W.J., Paynter, M.J.B., and Gupta, R., Characterization of Methanococcus maripaludis sp. nov., a new methanogen isolated from salt marsh sediment, Arch. Microbiol., 1983, vol. 135, pp. 91–97.
Kallistova, A.Yu., Merkel A.Yu., Tarnovetskii I.Yu., and Pimenov N.V., Methane formation and oxidation by prokaryotes, Microbiology (Moscow), 2017, vol. 86, pp. 671–691.
Kimura, Z. and Okabe, S., Acetate oxidation by syntrophic association between Geobacter sulfurreducens and a hydrogen-utilizing exoelectrogen, ISME J., 2013, vol. 7, pp. 1472–1482.
Lanoil, B.D., La Duc, M.T., Wright, M., Kastner, M., Nealson, K.H., and Bartlett, D., Archaeal diversity in ODP legacy borehole 892b and associated seawater and sediments of the Cascadia Margin, FEMS Microbiol. Ecol., 2005, vol. 54, pp. 167–177.
Lazar, C.S., Parkes, R.J., Cragg, B.A., L’Haridon, S., and Toffin, L., Methanogenic diversity and activity in hypersaline sediments of the centre of the Napoli mud volcano, Eastern Mediterranean Sea, Environ. Microbiol., 2011, vol. 13, pp. 2078–2091.
Lever, M.A., Torti, A., Eickenbusch, P., Michaud, A.B., Šantl-Temkiv, T., and Jørgensen, B.B., A modular method for the extraction of DNA and RNA, and the separation of DNA pools from diverse environmental sample types, Front. Microbiol., 2015, vol. 6, p. 476.
L’Haridon, S., Chalopin, M., Colombo, D., and Toffin, L., Methanococcoides vulcani sp. nov., a marine methylotrophic methanogen that uses betaine, choline and N,N-dimethylethanolamine for methanogenesis, isolated from a mud volcano, and emended description of the genus Methanococcoides,Int. J. Syst. Evol. Microbiol., 2014, vol. 64, pp. 1978–1983.
Li, D., Ran, Y., Chen, L., Cao, Q., Li, Z., and Liu, X., Instability diagnosis and syntrophic acetate oxidation during thermophilic digestion of vegetable waste, Water Res., 2018, vol. 139, pp. 263–271.
Lin, Y.-S., Biddle, J.F., Lipp, J.S., Orcutt, B.N., Holler, T., Teske, A., and Hinrichs, K.-U., Effect of storage conditions on archaeal and bacterial communities in subsurface marine sediments, Geomicrobiol. J., 2010, vol. 27, pp. 261–272.
Liu, X., Li, M., Castelle, C.J., Probst, A.J., Zhou, Z., Pan, J., Liu, Y., Banfield, J.F., and Gu, J.D., Insights into the ecology, evolution, and metabolism of the widespread Woesearchaeotal lineages, Microbiome, 2018, vol. 1. https://doi.org/10.1186/s40168-018-0488-2
Lyu, Z. and Liu, Y., Diversity and taxonomy of methanogens, in Biogenesis of Hydrocarbons, Stams, J.M. and Sousa, D., Eds., New York: Springer, 2018, pp. 1–59.
Merkel, A.Y., Pimenov, N.V., Rusanov, I.I., Slobodkin, A.I., Slobodkina, G.B., Tarnovetskii, I.Y., Frolov, E.N., Dubin, A.V., Perevalova, A.A., and Bonch-Osmolovskaya, E.A., Microbial diversity and autotrophic activity in Kamchatka hot springs, Extremophiles, 2017, vol. 21, pp. 307–317.
Michaelis, W., Seifert, R., Nauhaus, K., Treude, T., Thiel, V., Blumenberg, M., Knittel, K., Gieseke, A., Peterknecht, K., Pape, T., Boetius, A., Amann, R., Jørgensen, B.B., Widdel, F., Peckmann, J., Pimenov, N.V., and Gulin, M.B., Microbial reefs in the Black Sea fueled by anaerobic oxidation of methane, Science, 2002, vol. 297, pp. 1013–1015.
Omoregie, E.O., Niemann, H., Mastalerz, V., de Lange, G.J., Stadnitskaia, A., Mascle, J., Foucher, J.-P. and Boetius, A., Microbial methane oxidation and sulfate reduction at cold seeps of the deep Eastern Mediterranean Sea, Mar. Geol., 2009, vol. 261, pp. 114–127.
Orphan, V.J., Hinrichs, K.U., Ussler, W., Paull, C.K., Taylor, L.T., Sylva, S.P., Hayes, J.M., and Delong, E.F., Comparative analysis of methane-oxidizing archaea and sulfate-reducing bacteria in anoxic marine sediments, Appl. Environ Microbiol., 2001, vol. 67, pp. 1922–1934.
Pachiadaki, M.G., Kallionaki, A., Dahlmann, A., De Lange, G.J., and Kormas, K.A., Diversity and spatial distribution of prokaryotic communities along a sediment vertical profile of a deep-sea mud volcano, Microb. Ecol., 2011, vol. 62, pp. 655–668.
Pimenov, N.V. and Ivanova, A.E., Anaerobic methane oxidation and sulfate reduction in bacterial mats on coral-like carbonate structures in the Black Sea, Microbiology (Moscow), 2005, vol. 74, pp. 362–370.
Polikarpov, G.G., Egorov, V.N., Nezhdanov, A.I., et al., Active gas emission from elevations at the continental slope of the western Black Sea, Dokl. AN USSR, 1989, no. 12, pp. 13–15.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F.O., The SILVA ribosomal RNA gene database project: improved data processing and web-based tools, Nucl. Acids Res., 2013. database iss, pp. D590–D596.
Rinke, C., Schwientek, P., Sczyrba, A., Ivanova, N.N., Anderson, I.J., Cheng, J.F., Darling, A., Malfatti, S., Swan, B.K., Gies, E.A., Dodsworth, J.A., Hedlund, B.P., Tsiamis, G., Sievert, S.M., Liu, W.T., et al., Insights into the phylogeny and coding potential of microbial dark matter, Nature, 2013, vol. 499, pp. 431–437.
Roussel, E.G., Sauvadet, A.-L., Allard, J., Chaduteau, C., Richard, P., Bonavita, M.-A.C., and Chaumillon, E., Archaeal methane cycling communities associated with gassy subsurface sediments of Marennes-Oléron Bay (France), Geomicrobiol. J., 2009, vol. 26, pp. 31–43.
Schloss, P.D. and Handelsman, J., Toward a census of bacteria in soil, PLoS Comput. Biol., 2006, vol. 2, p. 92. https://doi.org/10.1371/journal.pcbi.0020092
Tarnovetskii, I.Y., Merkel, A.Y., Kanapatskiy, T.A., Ivanova, E.A., Gulin, M.B., Toshchakov, S., and Pimenov, N.V., Decoupling between sulfate reduction and the anaerobic oxidation of methane in the shallow methane seep of the Black Sea, FEMS Microbiol. Lett., 2018, vol. 365. https://doi.org/10.1093/femsle/fny235
Timmers, P.H.A., Vavourakis, C.D., Kleerebezem, R., Sinninghe Damsté, J.S., Muyzer, G., Stams, A.J.M., Sorokin, D.Y., and Plugge, C.M., Metabolism and occurrence of methanogenic and sulfate-reducing syntrophic acetate oxidizing communities in haloalkaline environments, Front. Microbiol., 2019, vol. 9, article 3039.
Vigneron, A., Cruaud, P., Roussel, E.G., Pignet, P., Caprais, J.C., Callac, N., Ciobanu, M.C., Godfroy, A., Cragg, B.A., Parkes, J.R., Van Nostrand, J.D., He, Z., Zhou, J., and Toffin, L., Phylogenetic and functional diversity of microbial communities associated with subsurface sediments of the Sonora Margin, Guaymas Basin, PLoS One, 2014, vol. 9. e104427.
Vigneron, A., L’Haridon, S., Godfroy, A., Roussel, E.G., Cragg, B.A., Parkes, R.J., and Toffin, L., Evidence of active methanogen communities in shallow sediments of the Sonora margin cold seeps, Appl. Environ. Microbiol., 2015, vol. 81, pp. 3451–3459.
Watkins, A.J., Roussel, E.G., Parkes, R.J., and Sass, H., Glycine betaine as a direct substrate for methanogens (Methanococcoides spp.), Appl. Environ. Microbiol., 2014, vol. 80, pp. 289–293.
Westerholm, M., Roos, S., and Schnürer, A., Syntrophaceticus schinkii gen. nov., sp. nov., an anaerobic, syntrophic acetate-oxidizing bacterium isolated from a mesophilic anaerobic filter, FEMS Microbiol. Lett., 2010, vol. 309, pp. 100–104.
ACKNOWLEDGMENTS
The authors are grateful to their colleagues from the Laboratory of Microbial Biotechnology, Faculty of Biology, Moscow State University for collaboration in growing anaerobic enrichment cultures.
Funding
This work was financially supported by the Russian Foundation for Basic Research (project no. 17-04-00023) and the Ministry of Science and Higher Education of the Russian Federation. The results of high-throughput sequencing were analyzed with support of the Russian Science Foundation grant 17-74-30025.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by D. Timchenko
Rights and permissions
About this article
Cite this article
Tarnovetskii, I.Y., Merkel, A.Y. & Pimenov, N.V. Analysis of Cultured Methanogenic Archaea from the Tarkhankut Peninsula Coastal Methane Seeps. Microbiology 88, 681–688 (2019). https://doi.org/10.1134/S0026261719060183
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261719060183