Abstract—
The known approaches to sample preparation have been improved to achieve a more complete detection of microorganisms of the Black Sea bottom sediments using flow cytometry of SYBR Green I-stained cells. Total microbial abundance in the samples from the shelf and deep-sea sediments varied from 0.03 to 1.54 × 108 cells g–1 and from 0.002 to 1.24 × 108 cells g–1 dry weight, respectively. This is comparable to the data reported previously for various areas of the oceans, including the Black Sea. Application of sodium pyrophosphate was shown to be the most universal method for treating sediments of various types; along with this, using hydrofluoric acid is possible for the deep-sea reduced sediments, whereas treatment with methanol was preferable for the sediments of coastal waters with a normal degree of aeration of the bottom layer. For samples of various types, optimal sample preparation procedures were proposed (choice of chemical reagent, mode of ultrasonic processing and centrifugation, and additional washing procedures). These procedures resulted in significantly more efficient enumeration of bacterial cells, while application of flow cytometry ensured rapid determination of the total number of microorganisms in the bottom sediments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The Black Sea is the world largest meromictic basin, with the water column characterized by the presence of a stable halocline separating the brackish oxic layer from the anoxic one. Microbiological research dealt mainly with the shelf sediments (Mironov, 1979; Sorokin, 1982; Thamdrup et al., 2000; Lein et al., 2002; Pimenov et al., 2013; Tarnovetskii et al., 2018) and less often, with deep-water sites (Leloup et al., 2007; Schippers et al., 2012). The data on total microbial abundance in Black Sea bottom sediments are usually given in relation to research on specific microbial groups (methane oxidizers, sulfate reducers, etc.) and their role in the biogeochemical cycles (Thamdrup et al., 2000; Lein et al., 2002; Leloup et al., 2007; Egorov et al., 2011; Schippers et al., 2012; Pimenov et al., 2013; Malakhova et al., 2015; Tarnovetskii et al., 2018).
Difficulties in separation of bacterial cells from the abiotic fraction due to tight binding are the major problem in determination of total microbial abundance in the samples with high levels of suspended matter (e.g., from estuaries, tidal zones, or bottom sediments). Moreover, organic and inorganic particles of the samples may exhibit nonspecific binding to fluorochromes and/or autofluorescence, which hinders detection of bacterial cells by any technique used (Weinbauer et al., 1998; Danovaro et al., 2001; Kallmeyer et al., 2008; Morono et al., 2009).
Several methods have been recently proposed for separation of microbial cells from sediment particles. Nonionic (Tween 80) and ionic (sodium pyrophosphate) surfactants were most often used to treat bottom sediments (Frischer et al., 2000; Danovaro et al., 2001, 2010). Application of methanol as a reagent degrading the polysaccharide exopolymers responsible for microbial adherence to nonbiological particles was also reported (Lunau et al., 2005; Kallmeyer et al., 2008). A two-stage treatment first with hydrofluoric acid and then with a methanol-containing mixture to stop the reaction was proposed. This procedure caused dissolution of amorphous silicates and extracellular biomolecules, resulting in decreased nonbiological fluorescence after fluorochrome staining. Its effect on intracellular DNA was minimal (Morono et al., 2009). Apart from chemical treatment of bottom sediments, mechanical treatment (shaking, sonication, blending, and centrifugation) was also proposed (Lindahl and Bakken, 1995; Kallmeyer et al., 2008). However, the authors noted the probability of higher cell lysis accompanying more efficient cell removal from the substrate. Thus, every method had its advantages and shortcomings and was suitable only for specific basins of for oxidized or reduced sediments.
Most works on cell detection in the bottom sediments by direct microscopic techniques were based on fluorescence microscopy. Flow cytometry is more often used for investigation of soil microbial abundance (Porter et al., 1997), although it was recently also applied to lake and river bottom sediments (Duhamel and Jacquet, 2006; Amalfitano and Fazi, 2008). No publications reporting such works on marine sediments are available.
The goal of the present work was to enhance the efficiency of determination of total prokaryotic abundance in the sediments from different Black Sea regions by combination of chemical and physical approaches to sample preparation and utilization of flow cytometry.
MATERIALS AND METHODS
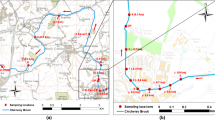
Subjects. Bottom sediments from the Bay of Sevastopol (44°37.53ʹ N, 33°31.32ʹ E; 14-m depth) and from a deep-water zone in the western part of the Black Sea (44°40.70ʹ N, 31°51.70ʹ E; 756-m depth) were used in the work. Bottom sediments were collected with 30 × 30 × 50 cm EPA box corer (United States) and/or an acrylic tube with a gate valve. Immediately after collection, the core was sliced vertically into layers 1–2 cm thick which were placed into petri dishes. The samples were stored at –20°C for not longer than 6 months and were thawed immediately before analysis.
Preparation of sample suspensions and fixation. Representative samples (in two replicates) were prepared by mixing three portions of sediments from relevant layers. Prior to preparation, the samples (0.2–1.0 g) were diluted with filtered seawater, mixed vigorously by hand, and fixed with 40% formaldehyde (50 µL mL–1 sample). The volume of the sediment suspension was 2–7 mL, depending on the method. Stock solutions and seawater were sterilized by filtration through 0.2-µm membranes (Sartorius, Germany) immediately before analysis.
Chemical treatment with sodium pyrophosphate. The sodium pyrophosphate (Na4P2O7) stock solution (50 mM) was added to the sample suspension to the final concentration of 5 mM. After 15-min incubation in the dark at room temperature, the samples were shaken by hand for 1 min and then sonicated 1 to 15 min (Unitra unima 01SZTYN UM-4, 140 VA, 220 V, 50 Hz). The samples were then centrifuged (1‒5 min, 700 g) for cell separation from the sediment particles and in order to increase the supernatant volume (Danovaro et al., 2001, 2010). To improve the signal quality of cytometric measurements, sonication time was increased to 30 min. In order to prevent sample overheating, sonication was carried out in an ice bath. The test tubes were shaken manually after 15 min and vortexed for 1 min after the treatment (Microspin FV-2400, Biosan, Latvia). Aliquots of the supernatant (1 mL) were used for microscopy and cytometric analysis.
Methanol treatment. The sample suspension (6 mL, unfixed) was supplemented with 0.6 mL 10% methanol (СН3OH) and shaken by hand. The samples were sonicated for 15 min at 35°C and then vortexes for 1 min. Detritus and inorganic particles were removed by centrifugation (1 min, 190 g) (Lunau et al., 2005; Kallmeyer et al., 2008). For subsequent analyses, 1 mL of the supernatant was collected.
Hydrofluoric acid treatment. The sediment suspension (0.5 mL, after dilution and fixation) was supplemented with 0.7 mL hydrofluoric acid (1% HF in 3% NaCl) in a plastic test tube and incubated for 20 min at room temperature. After incubation the samples were shaken by hand and treated with 4 mL of the mixture containing 1 M Tris-HCl, pH 8.0; 0.125 M CaCl2, and 25% СH3OH, sonicated for 1 min an ice bath, and centrifuged (1 min, 100 g) (Morono et al., 2009). The supernatant (1 mL) was then collected for analysis.
Washing procedures (similar for all methods). For more accurate detection of microbial cells remaining in the sediment after chemical and mechanical treatment, a series of washing procedures was carried out. After collecting aliquots for analysis, the supernatant was removed, filtered seawater was added to the sediment, and the sample was shaken and centrifuged as required for the method used. For each aliquot of the supernatant (1 mL), the number of microbial cells was determined in 3‒10 repeats.
Staining with SYBR Green I. Staining with SYBR Green I was carried out according to the protocols by Marie et al. (1997) and Noble and Fuhrman (1998). Stock solutions of the fluorochrome (10 µL mL–1 sterile Milli-Q water) were stored at –20°C. The samples (1 mL) were stained with 10 µL of the stock solution and incubated in the dark for 60 min.
Epifluorescence microscopy. Bacterial cells were concentrated using nuclear filters with 0.2 µm pore diameter (Joint Institute for Nuclear Research, Dubna, Russia) and a Sartorius filtration device (Germany). For uniform distribution of the cells, the filter was placed on a moist ashless paper filter with 1.5–2.5-µm pore diameter. During filtration, the vacuum did not exceed 150 mm Hg. Proper fluorescence of the filters was quenched by preliminary staining for 24 h with saturated Sudan black in 50% ethanol (Hobbie et al., 1977). The cells were enumerated in three repeats under a Jenalumar-a/d microscope (Carl Zeiss, Germany) equipped with an HBO-202 mercury lamp, at ×1000 magnification. For enumeration of SYBR Green I-stained cells, the following set of optical filters was used: 465 to 505 nm for excitation, 510-nm barrier filter, and 515 to 565 nm for emission (Marie et al., 1997; Noble and Fuhrman, 1998). The measurements were carried out in two repeats, with at least 200 cells counted on each filter in 10–20 fields, in order to obtain the data with an error not exceeding 20% at 95% significance level (Lebedeva et al., 1969).
Flow cytometry. The samples were analyzed on a CytomicsTM FC 500 flow cytometer (Beckman Coulter, United States) with a blue (488 nm) argon laser using the CXP software package. The data were analyzed using the Flowing Software v. 2.5.0 (Perttu Terho, Turku Centre for Biotechnology, University of Turku, Finland, www.flowingsoftware.com).
Microbial abundance was determined in fluorochrome-stained samples, isolating the fluorescent fraction corresponding to the size of bacterial cells, on two-parameter cytogramms of forward scattering (FS) and SYBR Green I green fluorescence (FL1 channel, 525 nm) using nondimensional logarithmic scales (Gasol et al., 2000). Cell concentrations were calculated taking into account the sample flow rate (15 µL min−1), counting time (60 s), and the number of cells detected during this time. Measurement quality was controlled using calibration fluorescent microspheres of different sizes (0.2–10 µm) (Beckman Coulter, Molecular Probes, United States). Suspensions were prepared so that the sample dilution after addition of filtered seawater and reagents was 10–30-fold. The sample volume used for analysis was 1 mL; measurement were carried out in two repeats. The number of particles counted during 1 s did not exceed 103–104.
The final versions of methodical protocols with our modifications are presented on Fig. 1. All methods involved the following common stages of sample treatment: (1) preparation of sediment suspensions in filtered seawater; (2) fixation (if necessary); (3) treatment with a chemical reagent; (4) mechanical treatment (sonication); and (5) cell enumeration by flow cytometry and/or under a microscope.
Generalized protocol showing the stages of separation of bacterial cells from sediment particles. Original protocols used were published by Danovaro et al. (2001, 2008, 2010) (for Na4P2O7); by Lunau et al. (2005) and Kallmeyer et al. (2008) (for СH3OH); and by Morono et al., 2009 (for combined method with HF and СH3OH).
Calculation of bacterial abundance. For calculation of total bacterial numbers, the volumes of solutions (fixing one, added reagents, and filtered seawater) used for sample preparation and washing were taken into consideration. A series of washings was carried out for each sample, apart from extraction, after chemical and mechanical treatment, and the data obtained after each stage were summarized. These data were used to construct cumulative curves, with the results obtained after 10 washings accepted as 100% cell desorption from the sediments. The total microbial abundance was expressed as cell numbers per 1 g of dry sample (cells g–1).
Statistical treatment was carried out using STATISTICA (data analysis software system) v. 10 (StatSoft, Inc., www.statsoft.com); the graphs were plotted using SigmaPlot 10.0 (SYSTAT Software, Inc.) and Grapher 8 (Golden Software, Inc.).
RESULTS
Cytometric analysis of the suspensions obtained after incubation of a sediment sample from the Bay of Sevastopol in 5-mM sodium pyrophosphate solution, sonication for 3–30 min, separation of large particles by short-term centrifugation, and staining with SYBR Green I revealed significant differences in cell abundance estimates depending on the sample preparation mode. Short-term sonication (3 min) and centrifugation (700 g, 1 min) resulted in emergence of a small fraction of particles with the size range and fluorescence intensity corresponding to those of bacterial cells (0.03–0.10 × 108 cells g–1 dry sample); most of the particles exhibited large size and low fluorescence (Fig. 2a, Table 1). The combination of longer sonication (15 min) with the same centrifugation mode resulted in detection of twice the cell number (0.25 ± 0.004 × 108 cells g–1 dry sample), although the fraction of noncellular particles remained abundant (Fig. 2b, Table 1). Relatively high numbers of microbial cells (0.50 ± 0.02 × 108 cells g–1 dry sample) and almost complete absence of the large-size fraction were observed after 30-min sonication in an ice bath and centrifugation for 3 min at 700 g (Fig. 2c, Table 1). This mode was used subsequently for determination of microbial abundance in the sediment samples from the Bay of Sevastopol and the deep-water Black Sea area (Table 1).
Cytograms obtained for the samples of the Bay of Sevastopol bottom sediments (2–3 cm layer) treated with sodium pyrophosphate with subsequent mechanical treatment and SYBR Green I staining: (a) sonication (3 min) and centrifugation (700 g, 1 min); (b) sonication (15 min) and centrifugation (700 g, 1 min); (c) sonication (30 min) and centrifugation (700 g, 3 min). Designations: FS, cell size; FL1, intensity of green fluorescence; N, bacterial abundance.
Epifluorescence microscopy was used to control the cytometric measurements. In both cases, the samples were treated with sodium pyrophosphate (15 min), sonicated (30 min at 0°C), centrifuged (700 g, 3 min), and stained with SYBR Green I. The data obtained for bacterial abundance in the Bay of Sevastopol sediments were within an order of magnitude and did not exhibit significant differences: (0.18 ± 0.12) and (0.16 ± 0.09) × 108 cells g–1 dry sample (cytometry and microscopy, respectively; the pairwise t-test, p > 0.05). We revealed a significant positive correlation between these values (r = 0.7); the regression equation and degree of determination are shown on Fig. 3.
Comparison of the data on bacterial abundance (N) in different layers of the Bay of Sevastopol sediments determined in the samples treated with sodium pyrophosphate (15 min), sonicated (30 min), centrifuged (700 g, 3 min), stained with SYBR Green I, and studied by flow cytometry and epifluorescence microscopy. Designations: FS, cell size; FL1, intensity of green fluorescence; N, bacterial abundance.
Cytometric analysis of the Bay of Sevastopol bottom sediments (the 3–4 cm layer), carried out after extraction with 10% methanol, sonication (15 min at 35°C), and centrifugation (190 g, 1 min), revealed high numbers of microbial cells (1.07 ± 0.04 × 108 cells g–1 dry sample). The fraction of noncellular large particles most probably consisted of nonbiological particles, as was indicated by the very low level of the FL1 signal (Fig. 4b). Comparison of two methods of chemical treatment (with Na4P2O7 and СH3OH) revealed similar values, although pyrophosphate treatment resulted in 1.5 times higher number of detected microorganisms (1.54 ± 0.04 × 108 cells g–1 dry sample) (Fig. 4a).
Cytometric analysis of the samples from bottom sediments of the deep-water anoxic Black Sea zone (western part of the sea, continental slope, 756 m) treated with methanol resulted in very low numbers of detected bacteria, from 0.002 to 0.16 × 108 cells g–1 dry sample (Table 2). After centrifugation, suspended particles remained, which contaminated the flow system of the cytometer. Microscopy revealed low cell numbers. Moreover, autofluorescence of suspended matter caused strong background fluorescence of the filter, which hindered cell enumeration. This method probably has certain limitations for investigation of deep-water Black Sea samples. The combined method with application of first hydrofluoric acid and then a methanol-containing mixture was therefore used (Morono et al., 2009). Microbial numbers revealed by this approach for different layers of the deep-water sediment sample were higher, up to 0.02–1.24 × 108 cells g–1 dry sample (Table 2). Pyrophosphate treatment, which was successfully used by other authors (Danovaro et al., 2001, 2008, 2010), was applied for control. In this case microbial numbers were 0.01–0.93 × 108 cells g–1 dry sample (Table 2). Thus, application of methanol resulted in microbial numbers (from the same layers of the deep-water Black Sea sediment) 4–20 times lower than those revealed after treatment with pyrophosphate or after combined treatment with hydrofluoric acid. This is in agreement with the data of Schippers et al. (2012), which were also obtained for deep-water Black Sea sediments.
In the case of the Bay of Sevastopol samples, efficiency of bacterial detachment after extraction from the sediment varied depending on the reagents used and the sonication mode: 62 ± 32 and 74 ± 17% for sodium pyrophosphate and methanol, respectively. These numbers were reliably lower than cumulative microbial numbers after three washings (pairwise t-test; p < 0.05). When the number of washings was increased to 4–10, the total number of bacteria continued to increase, although the difference between cumulative cell numbers in subsequent washings was not reliable (pairwise t-test; p > 0.05). Even after ten washings, some of the cells were certainly not removed from the sediment, but remained attached to particles. However, after chemical treatment, sonication, and three washings the number of bacterial cells in the supernatant was 92 ± 6% for pyrophosphate (Figs. 5a, 5b) and 95 ± 2% for methanol (Figs. 5c, 5d). Thus, the share of the cells attached to particles was low enough (below 5–8%) to be considered negligible.
DISCUSSION
Application of sodium pyrophosphate proved to be the universal method for treating all types of Black Sea sediments. This approach was most commonly used for determination of total bacterial numbers in bottom sediments from different depths of the World Ocean (Weinbauer et al., 1998; Danovaro et al., 2008, 2010; Schippers et al., 2012). However, the optimal sonication time used in the present work (30 min) was much longer than the short-term treatment (3 min) proposed in the original protocol (Danovaro et al., 2001, 2008, 2010). While Buesing and Gessner (2002) reported that increase in sonication duration from 3–10 to 90 min was possible, long-term treatment resulted in a small decrease of bacterial abundance (by 15%).
Methanol treatment proved to be most convenient when applied to the sediment of coastal areas with normal aeration of the near-bottom layer. Absence of the fixation stage was an advantage of this procedure, making it applicable for such analyses for which the presence of a fixing agent is unacceptable (e.g., ATP determination, molecular genetic techniques, etc.) (Lunau et al., 2005; Kallmeyer et al., 2008).
In the case of deep-water Black Sea sediments, apart from pyrophosphate treatment, a combined procedure involving treatment with hydrofluoric acid and then with a methanol-containing mixture proved useful. Unlike hydrochloric acid treatment, which was previously proposed for carbonate-rich soil (Kallmeyer et al., 2008), incubation with hydrofluoric acid was more efficient for deep-water samples containing high amounts of silicates (Morono et al., 2009).
In general, the values of total microbial abundance in the sediments determined in the present work (0.03–1.54 × 108 and 0.002–1.24 × 108 cells g–1 dry sample for the Bay of Sevastopol and the deep-water part of the Black Sea, respectively) were of the same order of magnitude that the results obtained by other researchers (Leloup et al., 2007; Schippers et al., 2012; Thamdrup et al., 2018; Tarnovetskii et al., 2018) for the Black Sea (Table 3) and for other areas of the World Ocean (Weinbauer et al., 1998; Danovaro et al., 2008). Some differences in absolute numbers (Table 3) are probably due to the differences in the methods of sample preparation, the stain used, and the ways of registering and calculation of the cell numbers (cells g−1 dry sample or cells mL−1 sediment suspension).
For exhaustive enumeration of bacterial cells in bottom sediments, sequential washing procedures became a necessary stage of sample preparation. While Danovaro et al. (2010) showed that 95-% removal of viruses was achieved by initial desorption and three sequential washings, they did not provide the data on bacteria. According to Siem-Jørgensen et al. (2008), the share of viruses and bacterial cells removal from the sediments was lower: 60 and 40% for viruses and bacteria, respectively. Our work showed that efficiency of bacterial detachment from the sediment varied from 62 to 74%, depending on the reagent applied. Although three subsequent washing procedures certainly increased the time of treatment for each sample, they provided for enumeration of 92–95% of the potential cumulative microbial abundance. At the stage of cell enumeration flow cytometry was a much more efficient technique than epifluorescence microscopy, saving much time when numerous samples are analyzed.
REFERENCES
Amalfitano, S. and Fazi, S., Recovery and quantification of bacterial cells associated with streambed sediments, J. Microbiol. Methods, 2008, vol. 75, pp. 237–243.
Buesing, N. and Gessner, M.O., Comparison of detachment procedures for direct counts of bacteria associated with sediment particles plant litter and epiphytic biofilm, Aquat. Microb. Ecol., 2002, vol. 27, pp. 29–36.
Danovaro, R. and Middelboe, M., Separation of free virus particles from sediments in aquatic systems, in Manual of Aquatic Viral Ecology, Wilhelm, S.W., Weinbauer, M.G., and Suttle C.A., Eds., Washington: ASLO, 2010, ch. 8, pp. 74–81.
Danovaro, R., Corinaldesi, C., Filippini, M., Fisher, U.R., Gessner, M.O., Jacquet, S., Magagnini, M., and Velimiriv, B., Viriobenthos in freshwater and marine sediments: a review, Freshwater Biol., 2008, vol. 53, pp. 1186–1213.
Danovaro, R., Dellánno, A., Trucco, A., Serresi, M., and Vanucci, S., Determination of virus abundance in marine sediments, Appl. Environ. Microbiol., 2001, vol. 67, pp. 1384–1387.
Duhamel, S. and Jacquet, S., Flow cytometric analysis of bacteria- and virus-like particles in lake sediments, J. Microbiol. Methods, 2006, vol. 64, pp. 316–332.
Egorov, B.V., Artemov, Y.G., Gulin, S.B., and Polikarpov, G.G., Methane seeps in the Black Sea: discovery, quantification and environmental assessment, J. Black Sea/Mediterranean Environ., 2011, vol. 17, pp. 171–185.
Frischer, M.E., Danforth, J.M., Newton Healy, M.A., and Saunders, F.M., Whole-cell versus total RNA extraction for analysis of microbial community structure with 16S rRNA-targeted oligonucleotide probes in salt marsh sediments, Appl. Environ. Microbiol., 2000, vol. 66, pp. 3037–3043.
Gasol, J.M. and Del Giorgio, P.A., Using flow cytometry for counting natural planktonic bacteria and understanding the structure of planktonic bacterial communities, Scientia Marina, 2000, vol. 64, pp. 197–224.
Hobbie, J.E., Daley, R.J., and Jasper, S., Use of Nuclepore filters for counting bacteria by fluorescence microscopy, Appl. Environ. Microbiol., 1977, vol. 33, pp. 1225–1228.
Kallmeyer, J., Smith, D.C., Spivack, A.J., and D’Hondt, S., New cell extraction procedure applied to deep subsurface sediments, Limnol. Oceanogr. Methods, 2008, vol. 6, pp. 236–245.
Lebedeva, M.N. and Shumakova, G.V., On rfeliability of the data obtained by direct microbial counts on filters, Mikrobiologiya, 1969, vol. 38, no. 2, pp. 351–357.
Lein, A., Pimenov, N., Guillouc, C., Martin, J.-M., Lancelot, C., Rusanov, I., Yusupov, S., Miller, Yu., and Ivanov, M., Seasonal dynamics of the sulphate reduction rate on the north-western Black Sea shelf, Estuarine, Coastal Shelf Sci., 2002, vol. 54, pp. 385–401.
Leloup, J., Loy, A., Knab, N.J., Borowski, C., Wagner, M., and Jørgensen, B.B., Diversity and abundance of sulfate-reducing microorganisms in the sulfate and methane zones of a marine sediment, Black Sea, Environ. Microbiol., 2007, vol. 9, pp. 131–142.
Lindahl, V. and Bakken, L.R., Evaluation of methods for extraction of bacteria from soil, FEMS Microbiol. Ecol., 1995, vol. 16, pp. 135‒142.
Lunau, M., Lemke, A., Walther, K., Martens-Habbena, W., and Simon, M., An improved method for counting bacteria from sediments and turbid environments by epifluorescence microscopy, Environ. Microbiol., 2005, vol. 7, pp. 961–968.
Malakhova, T.V., Kanapatskii, T.A., Egorov, V.N., Malakhova, L.V., Artemov, Yu.G., Evtushenko, D.B., Gulin, S.B., and Pimenov, N.V., Microbial processes and genesis of methane gas jets in the coastal areas of the Crimean Peninsula, Microbiology (Moscow), 2015, vol. 84, pp. 838–845.
Marie, D., Partensky, F., Jacquet, S., and Vaulot, D., Enumeration and cell cycle analysis of natural populations of marine picoplankton by flow cytometry using the nucleic acid stain SYBR Green I, Appl. Environ. Microbiol., 1997, vol. 63, pp. 186–193.
Mironov, O.G., Bacterial flora of the bottom sediments, in Osnovy biologicheskoi produktivnosti Chernogo morya (Basics of the Black Sea Biological Productivity), Kiev: Naukova Dumka, 1979, pp. 199–207.
Morono, Y., Takeshi, T., Noriaki, M., and Inagaki, F., Discriminative detection and enumeration of microbial life in marine subsurface sediments, ISME J., 2009, vol. 3, pp. 503–511.
Noble, R.T. and Fuhrman, J.A., Use of SYBR Green I for rapid epifluorescence counts of marine viruses and bacteria, Aquat. Microb. Ecol., 1998, vol. 14, pp. 113–118.
Pimenov, N.V., Egorov, V.N., Kanapatskii, T.A., Malakhova, T.V., Artemov, Y.G., Sigalevich, P.A., and Malakhova, L.V., Sulfate reduction and microbial processes of the methane cycle in the sediments of the Sevastopol Bay, Microbiology (Moscow), 2013, vol. 82, pp. 618–627.
Porter, J., Pickup, R., and Edwards, C., Evaluation of flow cytometric methods for the detection and viability assessment of bacteria from soil, Soil Biol. Biochem., 1997, vol. 29, pp. 91–100.
Schippers, A., Kock, D., Höft, C., Köweker, G., and Siegert, M., Quantification of microbial communities in subsurface marine sediments of the Black Sea and off Namibia, Front. Microbiol., 2012, vol. 3, p. 16. https://doi.org/10.3389/fmicb.2012.00016
Siem-Jørgensen, M., Glud, R.N., and Middelboe, M., Viral dynamics in a coastal sediment: Seasonal pattern, controlling factors and relations to the pelagic-benthic coupling, Mar. Biol. Res., 2008, vol. 4, pp. 165–179: suppl. material.
Sorokin, Yu.I., Chernoe more: priroda,resursy (Black Sea: Nature and Resources), Moscow: Nauka, 1982.
Tarnovetskii, I.Yu., Merkel, A.Y., Kanapatskiy, T.A., Ivanova, E.A., Gulin, M.B., Toshchakov, S., and Pimenov, N.V., Decoupling between sulfate reduction and the anaerobic oxidation of methane in the shallow methane seep of the Black Sea, FEMS Microbiol. Lett., 2018, vol. 365, iss. 21. fny235.https://doi.org/10.1093/femsle/fny235
Thamdrup, B., Rosselló-Mora, R., and Amann, R., Microbial manganese and sulfate reduction in Black Sea shelf sediments, Appl. Environ. Microbiol., 2000, vol. 66, pp. 2888–2897.
Weinbauer, M.G., Beckmann, C., and Höfle, M.G., Utility of green fluorescent nucleic acid dyes and aluminium oxide membrane filters for rapid epifluorescence enumeration of soil and sediment bacteria, Appl. Environ. Microbiol. 1998, vol. 64, pp. 5000–5003. Translated by P. Sigalevich
ACKNOWLEDGMENTS
The authors are grateful to V.S. Mukhanov for providing facilities for flow cytometric analysis, using the tables and protocols for analysis and interpretation of the results, to S.S. Svinin for assistance in cytometric analysis of the samples, to V.Yu. Proskurnin for collection of the bottom sediment samples and methodical recommendations, and to M.B. Gulin for valuable comments which improved the text of the article.
Funding
The work was carried out within the framework of the State Assignments Structural and Functional Organization, Productivity, and Stability of Marine Pelagic Ecosystems (state registration no. АААА-А18-118020790229-70828-2018-0005), Molismological and Biogeochemical Basis of Homeostasis in Marine Ecosystems (no. AAAA-A18-118020890090-2), and State assignment of the Research Center of Biotechnology, Russian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Rylkova, O.A., Gulin, S.B. & Pimenov, N.V. Determination of the Total Microbial Abundance in Black Sea Bottom Sediments Using Flow Cytometry. Microbiology 88, 700–708 (2019). https://doi.org/10.1134/S0026261719060158
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261719060158