Abstract
Drug assays with patient-derived cells such as circulating tumor cells requires manipulating small sample volumes without loss of rare disease-causing cells. Here, we report an effective technology for isolating and analyzing individual tumor cells and their clusters from minute sample volumes using an optimized microfluidic device integrated with pipettes. The method involves using hand pipetting to create an array of cell-laden nanoliter-sized droplets immobilized in a microfluidic device without loss of tumor cells during the pipetting process. Using this technology, we demonstrate single-cell analysis of tumor cell response to the chemotherapy drug doxorubicin. We find that even though individual tumor cells display diverse uptake profiles of the drug, the onset of apoptosis is determined by accumulation of a critical intracellular concentration of doxorubicin. Experiments with clusters of tumor cells compartmentalized in microfluidic drops reveal that cells within a cluster have higher viability than their single-cell counterparts when exposed to doxorubicin. This result suggests that circulating tumor cell clusters might be able to better survive chemotherapy drug treatment. Our technology is a promising tool for understanding tumor cell-drug interactions in patient-derived samples including rare cells.
Similar content being viewed by others
Introduction
Understanding interactions between tumor cells and drugs is important for discovery of new oncogenic targets1,2,3, development of cancer drug candidates4 and generating insights into the mechanisms of chemotherapy drug resistance5,6. Despite significant advances in understanding mechanisms of tumor development and progression7,8, the current clinical success rate of lead cancer drug candidates remains below 5%, significantly lower than that of cardiovascular (~20%) and infectious diseases (~17%) therapies2. Likewise, chemotherapy drug resistance is believed to be responsible for treatment failure in more than 90% patients with metastatic disease9, motivating the need to better understand in a patient-specific manner how chemotherapy drugs interact with cancer cells so that personalized treatments can be designed.
Identifying new drug targets or compounds and the molecular mechanisms of chemotherapy resistance requires preclinical models that adequately capture the complexities of cancer. Established tissue culture cell lines are often used as an in vitro model of cancer10,11,12, but these cell lines display amplified proliferation, transformed sensitivity to chemotherapy, and reduced cellular heterogeneity13,14,15. As a result, there has been a growing interest in conducting drug studies with patient-derived cells including human tissues and biofluids as a superior model of the in vivo situation10,13,16. Patient-derived cells are expected to better predict patient outcomes as they have been found to be more heterogeneous, with reduced proliferation rates and enhanced resistance to chemotherapy compared to established cell culture lines17.
Among the patient-derived cells, circulating tumor cells (CTCs) isolated from the blood of cancer patients offer a rich test bed for drug development and chemoresistance assays because (i) CTCs and their clusters (of typically 2–50 cells18,19,20) provide a compelling mechanism for metastasis19, with clusters having significantly more metastatic potential19, (ii) molecular profiling of CTCs shows they are very heterogeneous, similar to cells in a primary tumor, and share some common genetic mutations21,22, (iii) blood samples are less invasive compared to tissue biopsies and are easier to procure, and (iv) they can be sampled longitudinally for identifying drug resistance. Thus, CTCs are an attractive candidate for drug discovery and probing mechanisms of chemoresistance.
The promise of CTCs for drug investigations has been complemented by an explosion in the number of available microfluidic technologies available for isolating CTCs, even though they are present in low counts, typically 1–100 cells per mL of blood23. A number of microfluidic techniques are capable of antibody-based capture and release of CTCs24,25,26. In addition to these immunocapture methods, several label-free methods based on size and deformability also exist to separate CTCs25,27,28,29. More recently, clusters of CTCs have also been isolated using microfluidic approaches30.
The advent of numerous technologies for efficiently isolating CTCs opens unique opportunities for using CTCs for drug discovery and probing drug resistance. However, technical hurdles still exist for conducting drug investigations using CTCs. First, even though microfluidic technologies are available for efficiently isolating and collecting CTCs, conducting drug assays downstream can be challenging due to potential loss of the rare cells while handling them using pipettes and multiwell plates. Second, although ex vivo culture methods are beginning to emerge to culture CTCs for drug assays31,32,33, the molecular heterogeneity of individual CTCs and clusters is often lost during the bulk expansion process making it difficult to identify drug resistant cells.
In this study, we present a pipette-based microfluidic cell isolation (MCI) technology that is capable of conducting single cell resolution drug assays with a small number of tumor cells or their clusters present in small sample volumes (e.g. 10–100 cells in 10 μL). The method is based on digitizing the sample volume containing tumor cells into an array of nanoliter-scale droplets by simply using a pipette and a microfluidic device. The sample digitization occurs in the device in such a way that an array of static droplets is created in which tumor cells and their clusters are isolated. This approach also allows automated imaging of tumor cells stored in the droplets.
To establish proof-of-principle of our pipette-based MCI method for CTC research, we use breast cancer cells (MCF-7) and a chemotherapy drug, doxorubicin. Doxorubicin is an FDA approved cytotoxic drug used widely in cancer chemotherapy34,35 and this was chosen in this study as it is the most active single agent available for the treatment of breast cancer36. Using this system, we demonstrate that (i) individual MCF-7 cells can be isolated without any loss during the pipetting and digitization steps (ii) our method can isolate clustered tumor cells containing 2–22 cells per cluster (iii) the uptake profile of doxorubicin is heterogeneous in individual tumor cells and that a critical amount of intracellular uptake of doxorubicin determines the onset of apoptosis (iv) clustered tumor cells show a higher viability than their single-cell counterparts when exposed to doxorubicin suggesting that being in a cluster can alter a cancer cell’s response to chemotherapy drugs. The overall viability of clusters might be improved due to the coordinated resistivity provided by the group of cells. Overall, our MCI technology is a promising tool for chemotherapy drug testing on CTCs and their clusters.
Results and Discussion
Basic description of the MCI technology
In recent years, drop-based microfluidics has emerged as a powerful technology to compartmentalize cells in volumes down to picoliters37,38,39. The standard practice has been to encapsulate cells within trains of individual droplets produced in T-junction or flow-focusing devices, using syringe pumps. Subsequently, these droplets are stored in a variety of ways including on-chip incubation chambers/traps40,41 and off-chip incubation39. Although such methods have been shown to efficiently isolate individual cells for single-cell assays39, these techniques are incompatible with the needs of rare cell handling and manipulation, where only 10–100 cells are present in about 10–100 μL of sample volume. Specific reasons for the incompatibility include (i) establishing a stable flow and consistent drop production by syringe pumps requires stabilization time, which necessitates the use of more sample volume, typically >0.1 mL (ii) the inevitable presence of dead volumes resulting in not all of the sample being digitized into droplets and (iii) finally, there is a chance of losing cells in the syringe and tubing due to sedimentation and non-specific adhesion. Thus, the standard practice of using syringe pumps for droplet generation is ill suited for compartmentalizing rare cell samples.
We describe a simple method that resolves the above issues making it particularly suited for assaying rare cells. As shown in Fig. 1, our MCI technology involves using a pipette as a fluid-handling tool42 in conjunction with a microfluidic device that is capable of storing or parking arrays of nanoliter droplets. This approach is much more simplified than techniques presented in our previous work, where we used syringe pumps to create droplet arrays43,44. The current technique involves a highly robust method of creating an off-chip oil-sample cartridge in a pipette tip by sequentially aspirating oil and sample, followed by a single step dispensing into the microfluidic device that has been prefilled with oil.
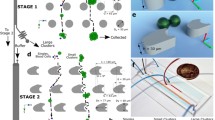
(a) Schematic showing preparation of cartridge by sequential aspiration of oil and cell-laden sample, followed by a single step dispensing into the microfluidic device that has been prefilled with oil. The microfluidic device contains 60 parking loops. Each loop has a bypass channel and a lower branch containing a trap that can park a droplet of volume 30 nL. (b) Time-stamped images of plug motion through the device. The sample plug (blue dye solution) fills the channels and traps and is sequentially digitized by the oil plug producing an array of static droplets of uniform size.
The mechanism of sample digitization is dictated by the geometry of the microfluidic parking network43,44,45. As shown in Fig. 1b, the device contains several interconnected parking loops, with each loop containing a trap chamber and a bypass channel. As the plug travels through the microfluidic device, the long aqueous plug breaks at the junctions of the parking loop, leaving remnants of the plug parked in the trap chambers. The manual pipetting does not introduce large error in droplet volumes since the trapped volume is preset by the size of the trap. This process creates an array of sixty drops of uniform volumes (30 nL in Fig. 1b, with a polydispersity of <5%) in less than 30 sec, using only 2 μL sample. Thus, our MCI approach enables the production of static droplet arrays (SDAs) with 90% of the sample being digitized and stored on the device, by simply using a pipette and avoiding the complications of syringe pumps.
Isolation of single tumor cells (Sl-TC)
Given that we can efficiently digitize the sample volume, we tested our approach as to whether (i) cells can be stored in the device without loss during sample manipulation, and (ii) single cells can be efficiently isolated in individual drops. This testing is important to ensure that rare CTCs can be isolated individually without cell loss. As a proof-of-principle, we used the human breast cancer cell line MCF-7. Experimental protocols used to test the efficacy of isolating MCF-7 cells are schematically illustrated in Fig. 2a. Four serially diluted solutions (500, 250, 100, and 50 cells per 10 μl) of cell samples were prepared from a stock solution of 3.5 × 106 cells/mL. These diluted samples were analyzed for cell counts both off-chip and in the device. For off-chip analysis, 2 μL of cell samples were pipetted on a glass slide and the number of cells were counted. For on-chip analysis, cell cartridges were prepared by aspiration of 5 μL of the biocompatible oil (FC-40) followed by aspiration of 2 μL of cell samples. This cell cartridge was then hand-dispensed into the microfluidic device that was preloaded with the biocompatible oil. The number of cells in each 30 nL drop was counted to determine the total number of cells in the microfluidic device. The results from the bulk sample analysis (n = 5) are compared with the counts obtained from microfluidic cell separation (n = 3) (Fig. 2b).
(a) Experimental protocol used to test the loss-less separation of MCF-7 cells. Cell samples were drawn from four serially diluted solutions and their concentration was measured by pipetting 2 μL on a glass slide and counting the number of cells. The same cell samples were loaded into the microfluidic device. The number of cells in each 30 nL drop was counted to determine the total number of cells in the microfluidic device. (b) The results from the bulk off-chip sample analysis (n = 5) are compared with the counts obtained from microfluidic cell separation (n = 3). The numbers in brackets denote the actual cell counts obtained from both the methods. The error bars represent standard deviation resulting from the trials conducted. (c) Efficacy of isolating single cells using microfluidics. For three different cell samples, the number of cells were counted in each 30 nL drop, and a probability distribution was generated from a total of 60 drops. The error bars represent standard deviation from n = 3 trials. The lines are predicted curves from Poisson distribution.
As shown in Fig. 2b, the data obtained from the two methods are in very good agreement. For example, the MCI method has 0% loss for the sample with 250 cells/10 μl, 5% loss for the sample with 100 cells/10 μl and maximum 15% loss for the sample with 500 cells/10 μl. Thus, the MCI technology showed 85–100% effectiveness for loss-less separation of cells from small sample volumes. Our results suggest that there is little or no adhesion of cells to pipette tips. However, a recent study suggests that nonspecific cell adhesion can occur due to the hydrophobic nature of most commercially available pipette tips46. We believe that non-specific adhesion of cells to pipette tips is mitigated in the MCI approach due to the presence of a thin biocompatible oil film in the pipette tip, which prevents cell loss during pipette-based fluid handling.
We also tested whether our method for single cell isolation produces Poisson statistics. Previously, single particle encapsulation in microfluidic droplets using T-junction and flow-focusing devices40 have shown that the encapsulation process follows a Poison distribution:

where x is the number of cells present in the droplet and λ is the expected average number of cells per droplet, which is adjusted by controlling the stock cell density.
In microfluidic droplet generation methods such as T and cross-junction, cells are encapsulated at sub-second time scales, however, in our case the timescale for encapsulating cells in droplets is tens of seconds (see Fig. 1b) causing potential cell sedimentation issues44, which might alter the cell encapsulation statistics from a Poisson distribution. To test the applicability of Poisson statistics for the MCI method, we used three different cell samples (500, 250, 100 cells/10 μl). For these cell concentrations, the expected average numbers of cells per droplet (λ) are 1.5, 0.75, and 0.3. The probability distribution P(x) was calculated by counting the number of cells per droplet (x) from a total of 60 drops for each cell sample. Using Eqn. (1), we also estimated the theoretical Poisson distribution considering the number of cells/per trap (x = 0, 1, 2, 3, 4…..) for each expected average number of cells per droplet (λ). Figure 2c shows the statistics of the isolation process from the experiment (symbols) and it’s comparison to theoretical prediction (dashed lines). We find that the experimental cell distribution closely follows Poisson statistics and about 25–35% drops in the microfluidic device contain single cells.
Overall, the MCI technology is capable of efficient isolation of individual cells from minute samples that contain a small number of cells, without much cell loss. The MCI approach addresses the current issues associated with handling rare cells using pump-based droplet microfluidics. Likewise, it outperforms traditional pipette and multiwell plate fluid handing method that have difficulty in handling nanoliter volumes and isolating single cells. Thus, the MCI technology has significant potential for handling small number of cells in minute samples that is typical for assays with isolated CTCs or primary human cells.
Influence of microfluidic drop environment on tumor cell viability
A potential concern of the MCI technology for drug testing is that the drop environment might adversely affect the viability of tumor cell suspension. For example, MCF-7 cells that are cultured in the laboratory in the adherent state, may undergo anoikis in the microfluidic drop environment, and therefore potentially introduce bias in our drug testing data. To address this concern, we conducted microfluidic experiments with CCRF-CEM leukemia cells whose natural as well as laboratory culture environment is liquid media. This allowed us to compare the level of anoikis of these CEM cells with that of MCF-7 cells incubated in microfluidic droplets. As shown in Fig. 3a, both the MCF-7 and CEM cells showed greater than 90% viability in droplets across 24 hours. Statistical analysis reveals that the percentage viability between the two cells lines is indistinguishable, suggesting that the level of anoikis is not very different between MCF-7 and CEM.
(a) Comparison of cell viability of MCF-7 breast tumor cells and CCRF-CEM leukemia cells in the liquid drop environment, measured over 24 hrs. For all time points, the difference in viability between the two cell lines is not statistically significant (p > 0.05: ns (not significant)) suggesting that the level of anoikis is not very different between the two cell lines. The error bars represent standard deviation from n = 12 trials. (b) Cell viability of MCF-7 cells in drops accessed over 7 days without any media change. Up to 3 days, more than 90% of cells survived. From day 4, the viability of cells decreased. Cell viability analysis was based on 1287 cells, with each drop containing ~10 cells. (c) Changes in drop volume stored in microfluidic device over 7 days. The drop volume (V) compared to the initial drop volume (V0) is less than 11% over 7 days. The average volume ratios (V/V0) and standard deviations were calculated from 3 replicates.
We also conducted long-term viability studies for up to 7 days, with MCF-7 cells in the microfluidic drops (see Fig. 3b). We observed that more than 90% cells are viable up to 72 hrs. On days 4, 5, 6 and 7, we found that cell viability is 75, 62, 43 and 21% respectively. We also recorded changes in drop volume due to evaporation, and find that the change in volume in 72 hrs is less than 10%, after which the drop volume remains almost unchanged (Fig. 3c). Thus, significant reduction in cell viability after day 3 is potentially due to lack of nutrients and anoikis. These results suggest that cells can be cultured successfully for drug testing up to 72 hrs, using the MCI technology.
Comparison of performance of MCI technology against multiwell plates
To assess the performance of our MCI technology vis-à-vis conventional method, we conducted a dose response assay using multiwell plates and the MCI technique, and compared the two methods. Muliwell plates are the most commonly used approach for conducting cell-based drug assays, Fig. 4 shows the comparison between the two approaches. We find that the dose-response curve obtained from the MCI technique is statistically indistinguishable from that of the 96-well plate data. This result suggests that the performance of our approach is comparable to that of multiwell plates, but with marked advantages of (i) isolating single cells without much cell loss and (ii) conducting dose-response assays with ~O(102) cells, rather than the O(104) cells that are typically used in multi-well plates.
Dose response curves with doxorubicin and MCF-7 cells were generated using both approaches, after 24 hrs of incubation. Cell viability analysis at each drug concentration was based on ~600 cells, in both the approaches. Each data point represents the mean response and the error bar corresponds to standard deviation from 3 replicates. The solid line is the average dose response curve, which was generated by fitting the standard Hill equation. The half maximal inhibitory concentrations (IC50) for 96-wellplate and microfluidic device are not statistically significant (p > 0.05).
Kinetics of Dox uptake and its relationship to tumor cell apoptosis
Given that the MCI technology has the capacity to isolate single cells and the microfluidic drop environment does not adversely affect cell viability, we studied the kinetics of uptake of doxorubicin by individual breast cancer cells. Doxorubicin can induce cancer cell death through different mechanisms: (i) inhibition of topoisomerase II47; (ii) intercalation to base pairs of the DNA double helix, blocking the synthesis of DNA and RNA, and DNA strand scission48; (iii) binding to cellular membranes to alter fluidity and ion transport; and (iv) generation of semiquinone free radicals and oxygen free radicals through an enzyme-mediated reductive process49.
Despite the ability of Dox to induce cell death through several mechanisms, breast cancer cells (MCF-7) can still show drug resistance to Dox after first chemotherapy50,51. The uptake rate of a drug by cancer cells is usually controlled by the resistivity of the cell population5,52. As a result, there is a need to identify which cells in a population are resistant to Dox uptake and how the uptake kinetics correlates with cellular apoptosis. Such investigations are difficult to pursue with bulk drug assays. Microfluidic approaches are suitable for single cell drug assays53,54, however, a survey of literature shows that Dox uptake kinetics has not been correlated to apoptotic susceptibility in MCF-7 cells. Given that we have demonstrated effective isolation of single cells using MCI technology, here we study single-cell kinetics of Dox uptake, identify Dox sensitive and insensitive cells and establish a correlation between accumulation of Dox and apoptotic susceptibility.
To quantify Dox uptake, the amount of auto fluorescence due to presence of Dox in the nucleus of individual cells was measured from droplets that contained only single cells (n = 118) over a period of 9 hrs. Similar to previous studies53,55, we observed significant heterogeneity among MCF-7 cells in terms of Dox uptake and retention. Figure 5a shows representative images of Dox accumulation in the nuclei of individual tumor cells (in red) co-stained with Annexin V (in green) for apoptosis. In general, we observe that cells that uptake high amounts of Dox undergo apoptosis. Importantly, we found eight different Dox uptake and retention profiles depending on how the Dox concentration, [C]Dox, changes between different arbitrary time points of 1, 3, 6 and 9 hrs. As shown in Fig. 5b, we observed cells in which [C]Dox continually decreases (d-d-d) or increases (i-i-i) or other combinations (e.g. d-d-i; d-i-i; i-d-d; i-i-d; i-d-i and, d-i-d). Among these most of the cells showed i-d-d (27%), d-d-i (15%) and, i-i-d (14%) uptake profiles and very few cells showed i-i-i (2%) and d-i-i (6%) profiles. Collectively, these results indicate significant heterogeneity in the uptake kinetics of Dox in MCF-7 cells.
(a) The top row images show representative images of Dox fluorescence in the nucleus (red) and the bottom row shows the corresponding co-stained images for apoptosis (green) for the same cell inside a droplet. The red box highlights the timepoint at which apoptosis was noted. Based on the Dox concentration after 1 hour of incubation, all cells were categorized in three groups defined as: high ([C]dox > 900 au), medium ([C]dox = 301–900 au) and, low ([C]dox < 300 au). (b) Dox uptake patterns of 118 single cells over 9 hrs. Total 8 different Dox uptake patterns were observed. Heterogeneity of individual MCF-7 cells was observed in terms of intracellular Dox uptake (represented by Dox concentration in arbitrary unit, au). The legends of subplots indicate the type of Dox uptake pattern; where‘d’ and ‘i’ represent decrease and increase in Dox concentration respectively. (c) Distribution of intercellular Dox concentration at four different time points. The red dashed line represents the critical Dox concentration needed for cellular apoptosis delineating the surviving and non-surviving cells.
Given the heterogeneous uptake profiles, we sought to investigate the correlation between Dox uptake and apoptotic susceptibility. We used two-color fluorescence imaging to establish this relationship and representative images are shown in Fig. 5a. We evaluated if there was a critical Dox uptake concentration at which tumor cells undergo apoptosis, irrespective of whether they uptake the drug slowly or rapidly. To address this, in Fig. 5c we plot the [C]Dox distribution at different time points and identified the [C*]Dox ≈ 825–900 at which apoptosis is initiated (vertical lines in Fig. 5c). Interestingly, we find that the critical intracellular Dox concentration needed to trigger apoptosis decreases only slightly with increase in incubation time. However, the percentage of apoptotic cells increases significantly from 35% to 58% at time points of 1 hr and 9 hr respectively. Taken together, these results suggest that a critical intracellular concentration of the chemotherapy drug Doxorubicin is needed to induce cell death in MCF-7 breast tumor cells and cells that do not uptake the critical Dox concentration survive.
The surviving and non-surviving cells show interesting uptake patterns. The non-surviving cells undergoing apoptosis uptake a Dox concentration higher than [C*]Dox at initial time points, but the Dox concentration decreases towards the later time points. The surviving cells mostly follow the d-i-i, d-d-i, i-i-i, and i-d-i patterns, indicating that towards later time points the Dox concentration increases, but still remaining mostly below [C*]Dox.
Isolation and drug response of clustered tumor cells (Cl-TC)
Most of the CTCs in patients are found as single cells. However, a very small fraction (2–5%) of CTCs are found as clusters with the number of cells within a cluster ranging from 2–5019,30. These clusters show 50 times more metastatic potential than single CTCs19,56,57,58. A major issue with handling CTC clusters is their sensitivity to rupture under shear. For example, the CTC-cluster chip uses an estimated shear rate of ≈700 s−1 to isolate tumor cell clusters from cancer patient blood30. The shear rate during the pipetting and loading/trapping process in the MCI technology is about 10–20 s−1, making it a promising candidate for handling clusters of tumor cells. We therefore generated clusters of MCF-7 tumor cells30 and tested the capability of the MCI method to efficiently encapsulate these clusters in the droplet array and conduct drug assays. We focused on addressing two questions: (i) What is the probability of generating clusters containing a given number of cells (ii) Do single tumor cells show different drug response than cells present in clusters?
Size distribution of tumor cell clusters
To generate clusters, we chose a stock cell density such that the average number of cells per droplet, λ = 10. The detailed protocol for preparing clusters is discussed in the Experimental section. Using this protocol, we trapped both clustered tumor cells (Cl-TC) and single tumor cells (Sl-TC) in drops (Fig. 6a, top). As shown in the images of Fig. 6a, we obtain clusters with Nc = 2–22 cells, where Nc is the number of tumor cells in a cluster. To characterize the probability distribution of obtaining a cluster with a given Nc, we interrogated 360 drops (from 6 microfluidic devices) and the data is shown in Fig. 6b. We find that clusters with Nc = 2 and 3 are the most predominant with probabilities of 30% and 26% respectively. Tumor cell clusters with Nc ≥ 4 have a lower probability of occurrence, about 1–12%. In addition, we note that the cluster size probability distribution depends on how cells detach during trypsinization and the extent of pipette-induced mixing, since the cluster distribution obtained from each of the 6 devices were not the same (data not shown).
(a) Isolation of tumor cell clusters along with single tumor cells in trapped drops of volume 30 nl (top). Physical morphology of different sizes of cell clusters (bottom). The number of cells in the cluster varies from 2–22. Each image is composite image of bright field (grey) and fluorescence (DAPI-stained blue nuclei). Scale bars represent 100 μm. (b) Probability distribution of clusters of different sizes. Data was taken from a total of 360 drops.
Drug response of single tumor cell (Sl-TC) vs. clustered tumor cell (Cl-TC)
Figure 7a shows representative images of clusters of different sizes that have been exposed to Dox. In general, we find that a significant number of clusters contain live cells even after 20 hrs of drug exposure. To investigate whether isolated cells show a different drug response than cells present in clusters, we tested the response of Dox on two different batches as shown in Fig. 7b. Batch [A] contained only single tumor cells (Sl-TCA) and Batch [B] had a mixture of clustered tumor cells (Cl-TCB) and single tumor cells (Sl-TCB).
(a) Images showing the viability of tumor cells in clusters of different sizes when exposed to 10 μM dox. Each image is a composite rendition showing live (green) and dead (red) cells. (b) The viability of two batches of MCF-7 cells were measured over time when exposed to 10 μM doxorubicin. Batch [A] contained only single tumor cells (Sl-TCA, n = 2275 and Batch [B] contained a mixed population of single tumor cells (Sl-TCB, n = 1054) and clustered tumor cells (Cl-TCB, n = 1621 cells out of 620 clusters). To test whether the clustered cells showed different response to drug compared to single cells, we performed standard unpaired t- test with 95% confidence interval at 10, 20 and 24 hrs. Statistical significance (p value) is represented with asterisks (*) with the following convention - *P < 0.05, **P < 0.01, ***P < 0.001. For all time points, cells in clusters (Cl-TCB) showed statistically higher viability than the viability of single individual cells [(Sl-TCA) and (Sl-TCB].
We assessed cell viability at 0, 10, 20 and 24 hrs in both batches. To test whether the clustered tumor cells show different drug response than single cells, we performed standard unpaired t- test for time points 10, 20 and 24 hrs. At these three time points, clustered tumor cells showed statistically higher viability than that of the independently generated single cell population as well as the co-existing single cell population. We checked for anoikis with clusters, and we did find it to contribute significantly to cell death (see Fig. S1). Thus, even though the single and the clustered cells were from the same population, the number of cells surviving drug exposure depended on whether they were present as individuals or clusters.
The Dox response of tumor cell clusters is different from that of single cells probably because the uptake kinetics of the drug could be different for a group of cells than individuals, especially when they are confined in microfluidic droplets. It is also possible that the overall drug resistivity of clustered tumor cells might be different compared to the resistivity of individual cells. From a cancer metastasis perspective, our result suggests that a CTC cluster has a better chance of surviving chemotherapy drug treatment than a single CTC.
Conclusions
Patient-derived cells from human tissues and biofluids are increasingly being used to predict patient outcomes as they are a superior model of the in vivo situation10,13,16. For example, recent advances to separate CTCs from blood have opened up opportunities for downstream assays of rare CTCs. However, to harness the full benefits of these downstream assays of rare CTCs requires manipulating a small number of cells without their loss during handling. Even loss of one diseased CTC from a pool of cells, might incorrectly predict the patient outcome. In this context, our simple MCI method offers benefits of i) reproducible sample discretization, ii) control over distribution of minute amount of cells/clusters without much loss of cells, iii) assessing heterogeneity at single cell level, iv) easy identification of drug response and, v) long time observation of intracellular events while preserving identity of each cell.
To demonstrate the power of the method, we have conducted proof-of-principle drug studies and provided useful insights into the response of single isolated tumor cells and clusters of tumor cells. Importantly, we have shown that individual cancer cells show a rich variety of drug uptake patterns and that clusters are more resistant to drug treatment. These results obtained from in vitro tumor cell lines need to be validated with actual CTCs from patient samples. The capabilities of the MCI technology can be further expanded to replace media from the droplets44 for long term drug studies and also characterize the influence of drugs on co-culture of cells (e.g. different proportions of tumor and stromal fibroblast cells59.
Methods
Fabrication of microfluidic devices
The design of the devices used in this study is identical to that used in our previous works43,44. Briefly, the device contains a microfluidic network of 60 loops (Fig. 1). Each loop has a bypass channel and a lower branch containing a trap that can park a droplet of volume 30 nL (Fig. 1a). In this microfluidic device44, all the channel widths and diameters of the trap chambers are 200 μm and 450 μm respectively. The height of all the features in the device is 170 μm. Microfluidic devices were fabricated using soft lithography60. Polydimethyl siloxane (PDMS) prepolymer and curing agent were mixed in a 10:1 ratio, degassed and poured on the mold and cured for a minimum of two hours at 65 °C. Subsequently the PDMS replica was cut with a scalpel and peeled. Inlet and outlet reservoirs were defined by punching holes. The replica was placed on a PDMS coated glass slide that was partially cured in the oven at 65 °C for 10 minutes. The entire assembly was further cured at 65 °C for two hours to create an irreversible seal.
Cell culture
The human breast cancer cell line (MCF-7) obtained from American Tissue and Culture Collection (ATCC, Manassas, VA, USA) was maintained and grown in DMEM medium containing 10% fetal bovine serum, 1% Sodium Pyruvate, and 1% Penicillin/Streptomycin at 37 °C in 5% CO2. Cell culture chemicals were purchased from Fisher Scientific. Different cell densities of MCF-7 cells were used as samples in the cell assay experiments. For all experiments, the initial cell concentration was 1 × 106 cells/ml and desired final concentrations were achieved by diluting with DMEM media.
Preparation of cell samples
To produce the cell samples for single tumor cell assays, we used a cell concentration of 0.35 × 106 cells/ml. The dilution and staining process were done by pipette mixing multiple times (at least 10 times) followed by vortex mixing to ensure that no clustered cells are disrupted.
To produce clustered tumor cell clusters (Cl-TC), we used the same concentration as the single cell assays, but we modified the mixing process to preserve cell clusters/groups. The total dilution and staining process were done by only three times mixing with pipette. This ensured mixing without significantly disrupting cell clusters. However, in this process each of the traps of the microfluidic device contained a mixture of clustered tumor cells and single tumor cells.
Drug assays
We conducted drug assays on the tumor cells in our microfluidic device, using the anti-cancer drug doxorubicin hydrochloride (Dox) (Sigma Aldrich). For all assays, Dox was used at a concentration of 10 μM. Initially, Dox was mixed with distilled water to make a 1 mM stock solution of drug and stored at 4 °C. For all experiments, stock solution of Dox was freshly diluted with DMEM media to a concentration of 10 μM. For drug assays, Dox was mixed with the cell samples and immediately loaded into the microfluidic device and imaged. The drug mixing and loading process were done within 2 minutes. In between different imaging time points the microfluidic devices were incubated at 37 °C in 5% CO2.
Pipetting procedure for loading cell samples
All the samples including cell suspensions and drug treated cells were handled in the microfluidic device using a multi-channel VIAFLO Electronic Pipette (0.5–12.5 μL, INTEGRA Biosciences Corp., NH, USA) and Grip-tips pipette tips (12.5 μL, INTEGRA Biosciences Corp., NH, USA). We tested several pipette tips (10 μL, 12.5 μL, 100 μL, 200 μL) and dispensed volumes (5 μL–10 μL) on a manual pipettor. We found that 12.5 μL, pipette tips with dispensed volumes of 7 μL worked the best in terms of uniform trapped volumes. The microfluidic devices were primed with a carrier fluid (FC-40, Sigma Aldrich) containing 0.001% w/w biocompatible fluorosurfactant (KrytoxFSH-PEG600-KrytoxFSH, RAN Biotechnologies, Inc., MA). Then, an off-chip oil-sample cartridge was prepared in the pipette tip using the multichannel automated pipette. The cartridge contains 5 μL of oil and 2 μL of sample. This oil-sample cartridge was then injected, using the electronic pipette, into the device that was prefilled with carrier fluid.
Viability and apoptosis detection
LIVE/DEAD® Cell Imaging Kit (Life technologies) was used to quantify cell viability. The ‘Live’ and ‘Dead’ parts of the stain were mixed following the manufacturer protocol. The mixed solution was diluted 3 times with cell media. This 3x-diluted solution was mixed with cell samples at 50/50 volume ratio to stain the cells. NucBlue® Live ReadyProbes® Reagent (Life technologies) were used to identify cell nuclei to accurately quantify cell counts.
Apoptosis was detected using Alexa Fuor® 488 Annexin V (Thermo Fisher Scientific). The dilution of cell samples and Dox were done with 1X Annexin- binding buffer (Thermo Fisher Scientific). Dox was mixed with the cell samples as described for other drug assays at a concentration of 10 μM. 25 μl of Alexa Fuor® 488 Annexin V (Thermo Fisher Scientific) was added to 100 μl cell-drug samples as an early-stage apoptosis marker. The cell sample with the Alexa Fuor® 488 Annexin V was incubated at room temperature for 15 min, loaded to the microfluidic device and then incubated at 37 °C in 5% CO2 for 45 mins. Fluorescence images (FITC and TRITC) were taken at time points of 1, 3, 6 and 9 hrs. In between imaging, the devices were incubated at 37 °C in 5% CO2.
Image acquisition and processing
Brightfield and fluorescence images of cells in droplets were captured using Olympus IX 81 microscope (Massachusetts, USA) equipped with a Thorlab automated stage (New Jersey, USA) controlled by SlideBook 6.1 (3i Intelligent Imaging Innovations, Inc., Denver, USA). Images were acquired with a Hamamatsu digital camera (ImagEM X2 EM-CCD, New Jersey, USA), 10–60x objectives and standard FITC/DAPI/TRITC filters. The exposure times used in the experiments ranged from 20–430 ms.
The uptake of doxorubicin i. e. concentration of Dox, [C]Dox, was quantified from the time-lapse images of each cell. Image processing was done with ImageJ (http://rsb.info.nih.gov/ij/). The [C]dox in arbitrary units (a.u.) was calculated after background correction by the following equation, [C]dox = Mean fluorescence of cell − Mean fluorescence of background. A composite image from FITC (live) and TRITC (dead) filters was used to assess live/dead cells in ImageJ.
Additional Information
How to cite this article: Bithi, S. S. and Vanapalli, S. A. Microfluidic cell isolation technology for drug testing of single tumor cells and their clusters. Sci. Rep. 7, 41707; doi: 10.1038/srep41707 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Collins, I. & Workman, P. New approaches to molecular cancer therapeutics. Nat Chem Biol 2, 689–700 (2006).
Kamb, A., Wee, S. & Lengauer, C. Why is cancer drug discovery so difficult? Nat Rev Drug Discov 6, 115–120 (2007).
Luo, J., Solimini, N. L. & Elledge, S. J. Principles of Cancer Therapy: Oncogene and Non-oncogene Addiction. Cell 136, 823–837 (2009).
Hoelder, S., Clarke, P. A. & Workman, P. Discovery of small molecule cancer drugs: Successes, challenges and opportunities. Molecular Oncology 6, 155–176 (2012).
Zahreddine, H. & Borden, K. L. B. Mechanisms and insights into drug resistance in cancer. Frontiers in Pharmacology 4, 28 (2013).
Raguz, S. & Yague, E. Resistance to chemotherapy: new treatments and novel insights into an old problem. Br J Cancer 99, 387–391 (2008).
Thiery, J. P. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2, 442–454 (2002).
Craene, B. D. & Berx, G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer 13, 97–110 (2013).
Longley, D. B. & Johnston, P. G. Molecular mechanisms of drug resistance. The Journal of Pathology 205, 275–292 (2005).
Burdall, S. E., Hanby, A. M., Lansdown, M. R. J. & Speirs, V. Breast cancer cell lines: friend or foe? Breast Cancer Research 5, 89–95 (2003).
Holliday, D. L. & Speirs, V. Choosing the right cell line for breast cancer research. Breast Cancer Research: BCR 13, 215–215 (2011).
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Keller, P. J. et al. Mapping the cellular and molecular heterogeneity of normal and malignant breast tissues and cultured cell lines. Breast Cancer Res 12, R87 (2010).
Stingl, J. & Caldas, C. Molecular heterogeneity of breast carcinomas and the cancer stem cell hypothesis. Nat Rev Cancer 7, 791–799 (2007).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100, 3983–3988 (2003).
Speirs, V. et al. Short-term primary culture of epithelial cells derived from human breast tumours. Br J Cancer 78, 1421–1429 (1998).
Dairkee, S. H. et al. A molecular ‘signature’ of primary breast cancer cultures; patterns resembling tumor tissue. BMC Genomics 5, 47 (2004).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Aceto, N. et al. Circulating Tumor Cell Clusters Are Oligoclonal Precursors of Breast Cancer Metastasis. Cell 158, 1110–1122 (2014).
Cho, E. H. et al. Characterization of circulating tumor cell aggregates identified in patients with epithelial tumors. Phys Biol 9, 016001 (2012).
Yu, M. et al. Circulating Breast Tumor Cells Exhibit Dynamic Changes in Epithelial and Mesenchymal Composition. Science 339, 580–584 (2013).
Polzer, B. et al. Molecular profiling of single circulating tumor cells with diagnostic intention. EMBO Molecular Medicine 6, 1371–1386 (2014).
Li, P. et al. Acoustic separation of circulating tumor cells. Proceedings of the National Academy of Sciences 112, 4970–4975 (2015).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Chaudhuri, P. K., Ebrahimi Warkiani, M., Jing, T., Kenry & Lim, C. T. Microfluidics for research and applications in oncology. Analyst 141, 504–524 (2016).
Joosse, S. A., Gorges, T. M. & Pantel, K. Biology, detection, and clinical implications of circulating tumor cells. EMBO Molecular Medicine 7, 1–11 (2015).
Huang, S. B. et al. High-purity and label-free isolation of circulating tumor cells (CTCs) in a microfluidic platform by using optically-induced-dielectrophoretic (ODEP) force. Lab Chip 13, 1371–1383 (2013).
Sollier, E. et al. Size-selective collection of circulating tumor cells using Vortex technology. Lab Chip 14, 63–77 (2014).
Hou, H. W. et al. Isolation and retrieval of circulating tumor cells using centrifugal forces. Scientific reports 3 (2013).
Sarioglu, A. F. et al. A microfluidic device for label-free, physical capture of circulating tumor cell clusters. Nature methods 12, 685–691 (2015).
Yu, M. et al. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science 345, 216–220 (2014).
Maheswaran, S. & Haber, D. A. Ex vivo culture of CTCs: an emerging resource to guide cancer therapy. Cancer research 75, 2411–2415 (2015).
Khoo, B. L. et al. Liquid biopsy and therapeutic response: Circulating tumor cell cultures for evaluation of anticancer treatment. Science Advances 2 (2016).
Minotti, G., Menna, P., Salvatorelli, E., Cairo, G. & Gianni, L. Anthracyclines: molecular advances and pharmacologic developments in antitumor activity and cardiotoxicity. Pharmacological reviews 56, 185–229 (2004).
Cristina, C. et al. Doxorubicin: The Good, the Bad and the Ugly Effect. Current Medicinal Chemistry 16, 3267–3285 (2009).
Tassone, P. et al. BRCA1 expression modulates chemosensitivity of BRCA1-defective HCC1937 human breast cancer cells. Br J Cancer 88, 1285–1291 (2003).
Teh, S. Y., Lin, R., Hung, L. H. & Lee, A. P. Droplet microfluidics. Lab Chip 8, 198–220 (2008).
Guo, M. T., Rotem, A., Heyman, J. A. & Weitz, D. A. Droplet microfluidics for high-throughput biological assays. Lab Chip 12, 2146–2155 (2012).
Brouzes, E. et al. Droplet microfluidic technology for single-cell high-throughput screening. Proceedings of the National Academy of Sciences 106, 14195–14200 (2009).
Koster, S. et al. Drop-based microfluidic devices for encapsulation of single cells. Lab Chip 8, 1110–1115 (2008).
Song, H., Tice, J. D. & Ismagilov, R. F. A microfluidic system for controlling reaction networks in time. Angew Chem Int Ed Engl 42, 768–772 (2003).
Derzsi, L., Kaminski, T. S. & Garstecki, P. Antibiograms in five pipetting steps: precise dilution assays in sub-microliter volumes with a conventional pipette. Lab Chip 16, 893–901 (2016).
Sun, M., Bithi, S. S. & Vanapalli, S. A. Microfluidic static droplet arrays with tuneable gradients in material composition. Lab Chip 11, 3949–3952 (2011).
Bithi, S. S., Wang, W. S., Sun, M., Blawzdziewicz, J. & Vanapalli, S. A. Coalescing drops in microfluidic parking networks: A multifunctional platform for drop-based microfluidics. Biomicrofluidics 8, 034118 (2014).
Bithi, S. S. & Vanapalli, S. A. Collective dynamics of non-coalescing and coalescing droplets in microfluidic parking networks. Soft Matter 11, 5122–5132 (2015).
Goebel-Stengel, M., Stengel, A., Taché, Y. & Reeve, J. R. The importance of using the optimal plastic and glassware in studies involving peptides. Analytical Biochemistry 414, 38–46 (2011).
Villman, K. et al. TOP2A and HER2 gene amplification as predictors of response to anthracycline treatment in breast cancer. Acta Oncol 45, 590–596 (2006).
Hilmer, S. N., Cogger, V. C., Muller, M. & Le Couteur, D. G. The hepatic pharmacokinetics of doxorubicin and liposomal doxorubicin. Drug metabolism and disposition 32, 794–799 (2004).
Gewirtz, D. A. A critical evaluation of the mechanisms of action proposed for the antitumor effects of the anthracycline antibiotics adriamycin and daunorubicin. Biochem Pharmacol 57, 727–741 (1999).
Aas, T. et al. Specific P53 mutations are associated with de novo resistance to doxorubicin in breast cancer patients. Nat Med 2, 811–814 (1996).
Taylor, C. W. et al. Different mechanisms of decreased drug accumulation in doxorubicin and mitoxantrone resistant variants of the MCF7 human breast cancer cell line. Br J Cancer 63, 923–929 (1991).
Housman, G. et al. Drug Resistance in Cancer: An Overview. Cancers 6, 1769–1792 (2014).
Sarkar, S., Cohen, N., Sabhachandani, P. & Konry, T. Phenotypic drug profiling in droplet microfluidics for better targeting of drug-resistant tumors. Lab Chip 15, 4441–4450 (2015).
Wood, D. K., Weingeist, D. M., Bhatia, S. N. & Engelward, B. P. Single cell trapping and DNA damage analysis using microwell arrays. Proc Natl Acad Sci USA 107, 10008–10013 (2010).
AbuHammad, S. & Zihlif, M. Gene expression alterations in doxorubicin resistant MCF7 breast cancer cell line. Genomics 101, 213–220 (2013).
Au, S. H. et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proceedings of the National Academy of Sciences 113, 4947–4952 (2016).
Cheung, K. J. et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc Natl Acad Sci USA 113, E854–863 (2016).
Aceto, N., Toner, M., Maheswaran, S. & Haber, D. A. En Route to Metastasis: Circulating Tumor Cell Clusters and Epithelial-to-Mesenchymal Transition. Trends in Cancer 1, 44–52 (2015).
Jaganathan, H. et al. Three-Dimensional In Vitro Co-Culture Model of Breast Tumor using Magnetic Levitation. Scientific Reports 4, 6468 (2014).
Duffy, D. C., McDonald, J. C., Schueller, O. J. & Whitesides, G. M. Rapid Prototyping of Microfluidic Systems in Poly(dimethylsiloxane). Anal Chem 70, 4974–4984 (1998).
Acknowledgements
We are thankful to the National Science Foundation (Grant Nos: 1150836 and 1445070) and the Cancer Prevention and Research Institute of Texas (Grant Nos: 140298 and 140840) for partially supporting this work. We are also thankful to Brandon Palomo and Jacqueline Berry for assistance with experiments.
Author information
Authors and Affiliations
Contributions
S.S.B. and S.A.V. designed the study, analyzed the data and wrote the manuscript. S.S.B. conducted all the experiments, and S.A.V. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Bithi, S., Vanapalli, S. Microfluidic cell isolation technology for drug testing of single tumor cells and their clusters. Sci Rep 7, 41707 (2017). https://doi.org/10.1038/srep41707
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41707
- Springer Nature Limited
This article is cited by
-
In vitro evidence for the potential of EGFR inhibitors to decrease the TGF-β1-induced dispersal of circulating tumour cell clusters mediated by EGFR overexpression
Scientific Reports (2024)
-
Intensity Histogram-Based Reliable Image Analysis Method for Bead-Based Fluorescence Immunoassay
BioChip Journal (2024)
-
A model for the dissemination of circulating tumour cell clusters involving platelet recruitment and a plastic switch between cooperative and individual behaviours
BMC Ecology and Evolution (2023)
-
Is cancer an intelligent species?
Cancer and Metastasis Reviews (2023)
-
A microfluidic device enabling drug resistance analysis of leukemia cells via coupled dielectrophoretic detection and impedimetric counting
Scientific Reports (2021)