Abstract
For honey bee and other social insect colonies the ‘queen substance’ regulates colony reproduction rendering workers functionally sterile. The evolution of worker reproductive altruism is explained by inclusive fitness theory, but little is known of the genes involved or how they regulate the phenotypic expression of altruism. We previously showed that application of honeybee queen pheromone to virgin fruit flies suppresses fecundity. Here we exploit this finding to identify genes associated with the perception of an ovary-inhibiting social pheromone. Mutational and RNAi approaches in Drosophila reveal that the olfactory co-factor Orco together with receptors Or49b, Or56a and Or98a are potentially involved in the perception of queen pheromone and the suppression of fecundity. One of these, Or98a, is known to mediate female fly mating behaviour, and its predicted ligand is structurally similar to a methyl component of the queen pheromone. Our novel approach to finding genes associated with pheromone-induced sterility implies conserved reproductive regulation between social and pre-social orders, and further helps to identify candidate orthologues from the pheromone-responsive pathway that may regulate honeybee worker sterility.
Similar content being viewed by others
Introduction
The evolution of altruism has long intrigued biologists interested in the origins of behavioural diversity. Reproductive altruism of the type typical of sterile worker and defensive castes of the social insects is obviously costly to the self-less individual, but nonetheless is predicted to evolve via indirect fitness effects1. Despite this central prediction from inclusive fitness theory we do not yet have a good understanding of which genes are under indirect selection or that are otherwise involved in mediating the expression of altruistic behaviour2,3,4. The European honey bee Apis mellifera is a post-genomic eusocial model5,6 that has been used to generate lists of genes implicated in worker sterility, mostly from microarray screens for genes responsive to ovary-inhibiting queen mandibular pheromone7,8,9. QMP is an honest signal of queen fecundity to which workers respond by de-activating their ovaries and otherwise adopting alloparental roles within their kin-based colonies10,11. The progress from array and other honeybee genomic studies12 has only begun to identify the most up-stream pheromone-responsive genetic elements through which workers regulate their ovaries in response to QMP. We have recently discovered however that application of QMP to virgin fruit flies suppresses fecundity in a manner comparable to its normal effect on honeybees. Treated flies tend to have smaller ovaries that contain fewer mature eggs13,14. This worker-like response from Drosophila melanogaster (Diptera) to a eusocial honeybee (Hymenoptera) pheromone implies conserved reproductive regulation between social and pre-social orders and introduces Drosophila as a proxy model for the action of sterility-inducing pheromones15.
In this study we use a Drosophila model to screen for loci involved in the olfactory response to queen mandibular pheromone. First, we use a bioassay to quantify the extent to which the fly’s response to QMP is strictly olfactory, as opposed to gustatory or tactile. Second, we screen the near-full complement of Drosophila melanogaster olfactory receptors (ORs) via RNAi-mediated knock-downs. Individual ORs that block the fly’s conspicuous worker-like response to QMP represent functional candidates for the olfactory perception of ovary-inhibiting queen pheromone. Finally, to the extent that receptors identified from the fly are homologous to those from the bee, we use our Drosophila model to identify, for the first time, candidates from the pheromone-responsive pathway that may regulate honeybee worker sterility.
Results and Discussion
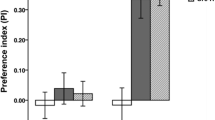
To determine whether the action of QMP in flies required direct physical contact, we set up two trails. Under “full access” trails we used custom-built chambers that permitted full physical contact with a pheromone-treated filter paper. Under “limited access” trails, by contrast, we fit a screen within chambers that prevented contact with the pheromone (Fig. 1A). Flies exposed to QMP consistently showed a worker-like response. In full access trials females are observed to regularly make contact with the pheromone-treated filter paper and produced smaller ovaries (F2,53 = 16.74, P < 0.001) that contained fewer eggs (F2,67 = 78.91, P < 0.001) than did untreated controls. Even with limited access (separated by ~4 cm) to QMP females yielded smaller ovaries (F2,52 = 17.19, P < 0.001) with fewer eggs (F2,67 = 37.95, P < 0.001) than did non-exposed controls (Fig. 1B). This latter response strongly implicates QMP as a near-distance olfactory cue for suppression of direct fitness.
Exposure and response to QMP.
(a) We used ‘full’ or ‘limited’ access chambers to expose groups (n = 5) of wild type flies to filter paper containing queen mandibular pheromone (QMP) or a no-QMP control. In full access chambers, flies could touch the filter paper; under limited access, they could not. (b) Response of wild type flies to QMP under full (gray bars) or limited (white bars) access. Both response variables (egg number, ovary area) decrease by 20 to 64% under QMP treatment, and this response holds in the limited access condition. Error bars indicated 95% confidence intervals.
To test whether olfaction is required for perception of QMP we compared the ovarian response of small groups (n = 5) of Ore-R females against two mutant genotypes deficient for the major olfactory co-factor Orco (formerly, Or83b). The mutant genotypes are: w1118; Orco1 and w1118; Orco2. They are each homozygous for loss-of-function alleles16 characterized by a coding region deletion, and both are effective at blocking a wide range of olfactory stimuli17. Orco females have inherently smaller ovaries18 and did not respond to QMP, unlike the background (w1118) or wild types (Ore-R) that did contain smaller ovaries (F3,145 = 7.26, P < 0.001) with fewer eggs (F3,144 = 8.90, P < 0.001; Fig. 2). This lack-of-response suggests that Orco – a 1-to-1 orthologue with the bee’s AmOr219, which itself is up-regulated in sterile workers7 – is essential for the perception of QMP and its downstream effect on ovaries.
Measuring Orco mutant response to QMP.
(a) Confocal images of Drosophila ovaries stained with DAPI. Wild type (Ore-R) and background control (W1118) genotypes have relatively large, well-developed ovaries under a zero ‘queen-equivalent’ dose of QMP that regress upon exposure to a [20] qe dose. The ovaries of Orco mutants (Orco1, Orco2), by contrast, are not affected by QMP. (b) Bar graphs summarize how control lines (Ore-R, black; w1118 solid grey) respond to QMP while mutant lines (Orco1 white; Orco2 striped) do not. This genotype × treatment effect is significant for both measures of fecundity (egg number, ovary area). Scale bar = 200 μm.
To identify olfactory receptors (ORs) that “tune” Orco to the specific perception of QMP we systematically knocked down individual ORs via Gal4-driven RNAi insertions20 available from Vienna Drosophila RNAi Center. We crossed UAS-RNAi males of either P-element RNAi (“GD Library”) or phiC31 (“KK Library”) genetic background with virgin w1118, elav-Gal4; UAS-dcr2 females to generate F1s that express OR-specific knockdowns (Fig. 3). We proceeded with lines only for which the background had a minimal effect on ovary phenotype. To select for this, we compared ovary scores for each RNAi line to background control F1s produced from crossing elav-Gal4; UAS-dcr2 females to GD or KK males (with no QMP). From each pairwise comparison (Supplementary Table 1), we considered only those knockdown lines for which the standardized difference in mean ovary scores was ‘small’ - i.e., Hedge’s g less than 0.5 (Supplementary Table 2). The majority of lines screened met this criterion (34 of 45 lines for egg number; 26 of 45 lines for ovary area) and were deemed suitable for testing the knockdown effect of specific ORs against QMP. We again measured this effect using Hedge’s g, except in this case by simply comparing QMP treated vs. untreated flies.
Exposure of RNAi flies to queen bee pheromone.
We crossed elav-GAL4; UAS-dcr2 females to males with specific UAS-OR-RNAi genotypes, corresponding to the n = 48 OR knockdowns available for Drosophila (from Vienna Drosophila RNAi Stock Center). We collected groups of n = 30 F1 larvae and reared them for ~10 days to maturity. We then exposed small groups (n = 5) of mature same-aged (within 1 hr) females to the QMP treatment. Flies received either no-QMP, or a low [13.3 queen-equivalents] or high [20 queen-equivalents] dose. Finally, after 48 hrs we dissected complete sets of ovaries, stained them with DAPI, and scored them against an established scale38 for assessing reproductive readiness, and did so via digitized confocal images.
For egg number, n = 23 lines had no appreciable knockdown effect on the perception of QMP – that is, females continued to show a worker-like response. The remaining lines did, however, show a strong knockdown effect as evidenced by lack-of-response to QMP (Fig. 4). For ovary area, n = 10 RNAi lines had no appreciable knockdown effect and thus continued to show the worker-like response. The remaining lines did, however, show a strong knockdown effect on the expected response to QMP. In total 16 unique ORs are identified from the two ovary-related assays, with some overlap. We noticed that three of the olfactory receptors – named Or49b, Or56a and Or98a – are retrieved from both assays independently, and have particularly strong knockdown effects as evidenced by lack-of-response to pheromone (g < 0.2; Fig. 4A,B). These three Drosophila receptors are therefore strongly and consistently associated with pheromone-induced ‘sterility’, and cluster on a genealogy21 with Hymenopteran OR gene subfamilies “B”, “C”, “D” and “E” (so named in ref. 21). Collectively, these gene subfamilies each potentially represent one OR gene copy in the common ancestor of Hymenoptera and contain a mere eight (of ~170)19 extent Apis mellifera olfactory receptor orthologues, which are: AmOr116, AmOr119 and AmOr68-AmOr73 (Fig. 5). Thus in addition to AmOr2 identified above, our screen implicates a clear set of n = 8 honeybee genes in the QMP-responsive pathway that may regulate Apis mellifera worker sterility. It is not yet feasible to knockdown individual olfactory receptor genes in the sensory tissues of the bee itself. This technology is being developed22,23 and we predict individual or collective knockdown of these eight bee receptors in a native context will inhibit the worker’s altruistic response to queen pheromone.
RNAi screen for olfactory receptors responsive to queen pheromone.
For each Drosophila olfactory receptor RNAi knockdown line (n = 48) we show the statistical effect size (Hedge’s g) of pheromone treatment (x-axis) and genetic background (y-axis) on egg number (a) and ovary area (b). RNAi lines that map to the lower left quadrant are hardly responsive to QMP (g < 0.5 on x) and have low genetic backgrounds (g < 0.5 on y). Three ORs in red have particularly strong knockdown effects (g < 0.2 on x) and are retrieved from both assays independently. We superimpose the Orco mutant effect for comparison. To validate the RNAi-implied function of the three highlighted receptors we used promoters for each of Or49b, Or56a and Or98a fused to GAL4 to drive expression of a tetanus toxin transgene (i.e., -TNTG) alongside an inactive toxin control (i.e., -TNTVIF). We find that this targeted disruption of neural activity effectively mimics the RNAi-knockdown effect for two of the three receptors tested: Or56a and Or98a are required to mediate the full pheromone response on egg number (c) and ovary area (d), while Or49b is seemingly required only for ovary area g < 0.5, (d) only. The wild type w1118-TNTG and inactive toxin lines (Or56a-TNTVIF, etc) show the typical very large effect of queen pheromone on fly egg number and ovary area.
Most highly related Apis olfactory receptors to the candidate Drosophila ORs identified in our screen.
Genealogical relationship between Hymenopteran olfactory receptor families (A-P, 9-exon, Orco) and the Drosophila olfactory receptors identified from the present screen (n = 4, incl. Orco). Red shows Drosophila genes (Or49b, Or56a, Or98a) embedded within the Hymenopteran genealogy, with dashes indicating 1-to-1 orthology between Orco and AmOr2. These relationships are re-drawn from Zhou et al21 (Supplementary Figure 3 therein) and are here used to identify the most-closely related Apis genes (n = 9, incl. AmOr2).
We independently tested the RNAi-derived results in the fly model using promoters for each of Or49b, Or56a and Or98a receptors fused to GAL4 to drive expression of the tetanus toxin, and scored the female’s reproductive response to pheromone. As predicted, the receptor-specific disruption of synaptic function dampens the response to queen pheromone, producing females with enlarged ovaries (F1 = 53.63, P < 0.001) that contain more eggs (F1 = 87.86, P < 0.001) than do non-toxic transgene controls (Fig. 4C,D). Why olfactory receptive flies respond to bee pheromone is an open question13,14, with one possibility being that social insect queen pheromones act on conserved regulatory pathways that were already present in solitary ancestors24. Regardless, our tetanus toxin-based validation suggests that the original RNAi screen did capture a functional subset of receptors with a capacity to perceive the ovary-inhibiting pheromone. Our receptor- and neuron-specific rescue is simply not expected under a non-specific pharmacological response to a pheromone of any type. Further testing of pheromone-treatment specificity is possible – for example, by testing flies for any gains-of-function predicted from honey bee biology upon exposure to QMP (e.g., behavioral attraction to pheromone). Or49b and Or98a are broadly tuned receptors that respond to a range of ecologically relevant odors25. Or56a, by contrast, is apparently very narrowly tuned but its function is nonetheless linked to ovary inhibition26, which is clearly relevant to ‘sterility’. An electrophysiology or activity assay27 will help clarify if fly olfactory neurons expressing these three receptors are actively and specifically detecting the bee pheromone.
Finally, we used the on-line Database of Odorant Receptors28 and the maximum common sub-structure method of Cao et al.29 to predict the affinity of predicted receptor ligands to any of the five components of QMP (Supplementary Table 3). Or98a, identified above, showed the single highest sub-structural similarity score to any component of QMP (to methyl p-hydroxybenzoate, HOB; Fig. 6). The conspicuous response from the fly to honeybee pheromone may, therefore, lie in conserved olfactory or other30 signaling mechanisms that remain linked to female fecundity and ovary de-activation, as predicted by socio-evolutionary hypotheses24,31,32,33. Or98a and HOB may indeed be linked to reproduction: the former mediates female fly mating behaviour34 and the latter molecule varies in its expression as a function of reproductive caste across Apis spp.35. Whether HOB or other single components of the queen pheromone blend can singlehandedly induce sterility in flies has not been widely tested36, but data emerging from honeybees does suggest that some single-components can suppress worker fertility24.
Structural similarity between QMP molecular components and predicted receptor ligands.
Queen mandibular pheromone consists of five organic components (9-ODA, cis/trans 9-HDA, HOB and HVA) that show sub-structural similarity to the principle ligands of Drosophila olfactory receptors, as measured here using Tanimoto’s coefficient (Supplementary Table 3). The ligand for Or98a (in red) has a strong affinity for the HOB component of queen pheromone.
Conclusions
Our findings advance insect sociobiology in two ways. First, we demonstrate how a novel Drosophila model can accelerate discovery of genes relevant to the evolution and expression of socially-mediated reproduction. Second, we implicate Orco and its Or49b, Or56a and Or98a receptors as functional orthologues in the pheromone-responsive pathways that may regulate honeybee worker sterility – a pathway of major significance to insect sociobiology.
Methods
Fly rearing
We reared all strains of Drosophila melanogaster under standard conditions (25 °C, 60% humidity and a 12 h:12 h light: dark cycle) in an insect growth chamber (Caron Inc., Marietta, OH) on a standard cornmeal diet, as described in ref. 14. We synchronized adult emergence by first housing (for 24 hrs) a small reproductive population (n = 30 males and n = 30 females) in collection cages (60 mm; Diamed, Mississauga, Canada) fitted with nutrient (grape juice and agar) plates. We then collected and transferred day-old larvae to fresh food vials (28.5 × 95 mm, VWR International, Radnar, PA) at a density of n = 30 larvae per vial. Finally, we collected same-age (within 1 h) adult virgin females approximately 10 days later, and dissected their ovaries after a further 48 hours.
Pheromone treatment
First, we diluted a 500 mg stock of synthetic QMP (Contech Ltd, Victoria, Canada) with 100% ethanol into two working concentrations; a relatively low dose of approximately 13 queen equivalent (qe) units37 and a higher dose of approximately 20 qe units. We use doses that are nominally high as prepared within the fly food (yeast and sugar) medium, but the amount actually received by flies coming in contact with the medium-soaked filter paper (in groups of five) is presumably much less. We reason that if female flies consume roughly 2 μl of food per day then the oral dose of QMP would be ~1.3 queen equivalents (13 QE in 20 μl @ 2 μl = 1.3 QE), or less via olfaction alone. Therefore, the effective dose is estimated to be within a biologically relevant range that has been shown to effectively suppress fly ovaries in a manner comparable to its normal effect on worker bees14. Moreover, the effect of pheromone treatment at these doses does appear to specifically affect fly ovaries, as opposed to other aspects of female reproduction (pupation) or survivorship14. Lack-of-response to a control pheromone (7-triclosene), and variable response among reproduction-related genotypes13, further suggests that this ovary-specific response is not simply a pharmacological side-effect to a pheromone of any type. Second, we warmed working aliquots to 50 °C in a water bath, and exposed flies to QMP in one of following two ways. Under “full access” we exposed flies to QMP within chambers that permitted full physical contact with the pheromone-treated filter paper. Under “limited access”, by contrast, we exposed flies to QMP within chambers fitted with a screen that prevented physical contact with the pheromone-treated filter paper.
For full access trials we placed n = 5 flies into a 50 ml Falcon tube modified to administer pheromone, as described in ref. 14. Briefly, we cut the bottom tip of the tube to insert a standard fly plug, then custom fit a piece of filter paper (grade 413: VWR International, Radnar, PA) saturated with a yeast and sugar solution (0.1 g yeast, 0.15 g sugar, in 5 ml of 5% ethanol) under the screw cap. To treat flies, we pipetted 20 μl of QMP-EtOH solution, or the equivalent volume of just-EtOH control, onto the paper, and incubated the whole chamber for a period of 48 hrs. For limited access trials, we performed a comparable procedure, except used a mesh barrier to prevent flies from touching the filter paper. Following treatment, we dissected the ovaries of individual flies, and scored the approximate level of activation in two ways; by counting the number of mature eggs38 per female (both ovaries) and by estimating the total ovary area, as inferred from on-screen measurements of digitized confocal microscope images.
Scoring the level of ovary activation
For all assayed flies, we exposed females within chambers for 48 hrs. We then CO2 anesthetized them and dissected complete pairs of ovaries from individual females using ultra-fine forceps under an Olympus S7 × 7 stereomicroscope (Olympus, Richmond Hill, Canada) that we fitted with a cold light source (KL300 LED; Leica Microsystems, Wetzlar, Germany). We dissected ovaries in 1X Dulbecco’s phosphate-buffered saline (1 × D-PBS; Invitrogen, Carlsbad, CA), then fixed stained tissue in a 4%-formaldehyde in D-PBS solution for a period of 20 min. We then washed [1 × D-PBS and 0.5% PBT (0.1% Triton X 100 in 1 × D-PBS)] and DAPI-stained (1:2000) ovaries prior to mounting (7% glycerol in D-PBS) and visualized them using a Zeiss LSM 5 Duo Vario confocal microscope (Zeiss, Oberkochen, Germany). Finally, we scored ovaries against two biological criteria that capture QMPs effect on reproductive readiness. First, we counted the number of mature (stage 14)38 eggs within each ovary. We then estimated ovary area (μm2) from confocal images using the ‘thresholding’ function of Image-Pro Premier 9.1 software (Version 9.1, Media Cybernetics, Bethesda, MD). For this part of the analysis, we excluded any ovaries that were inadvertently damaged during dissection or that otherwise had weak imaging (a minority, ~3–5%).
Electrochemical disruption of synapse via GAL4-driven tetanus toxin
To validate the RNAi knockdown effects on individual olfactory receptors, we drove a tetanus toxin transgene using promoters specific to receptors Or49b, Or56a and Or98a. These Or49b-GAL4, Or56a-GAL4, Or98a-GAL4 driver lines were obtained from Bloomington Stock Center. The active form of the tetanus toxin transgene (TNTG) inhibits neural function by blocking synaptic transmission39. The tissue-specific delivery of this effect into our top three ORs of interest (See Fig. 4A,B) provides an independent test of their physiological capacity to mediate the ovary-inhibiting effect of queen pheromone. Specifically, we crossed w; Or-GAL4 males with w; UAS-TNTG females to generate F1s that express the OR-specific neural disruption (e.g., w; Or98a-GAL4/UAS-TNTG). As a background control, we likewise crossed w; Or-GAL4 males with w; UAS-TNTVIF females carrying the inactive form of the toxin, and proceeded with only those lines for which the background had a very minimal effect on ovary phenotype (Hedges g less than 0.2). We predict that the synaptic disruption of specific olfactory receptors via tetanus toxin will mimic the original RNAi knockdown effect to render female flies significantly less responsive to ovary-inhibiting pheromone. The rearing, treatment and scoring of fly populations was identical to that for the RNAi experiment (above).
Calculating statistical effect size
Effect size comparisons were computed using

an unbiased version of Cohen’s d index of effect size for sample sizes smaller than n = 20 (Hedges, L. V. 1981. Distribution theory for Glass’s estimator of effect size and related estimators. Journal of Educational Statistics, 6, 107–128), whereby M is the population mean of each response variable, the pooled standard deviation SD of both samples is

Stoichiometric analysis of olfactory ligands to pheromone components
Following our screen, we compared the structural similarity of candidate olfactory receptor ligands to the five components40 of QMP: 9-ODA (E)-9-oxodec-2-enoic acid), HOB (methyl p-hydroxybenzoate), HVA (4-hydroxy-3-methoxyphenylethanol) and cis and trans 9-HDA (9-hydroxydec-2-enoic acid). First, we identified the dominant ligand for each candidate OR using the on-line Database of Odorant Receptors28. In four cases (corresponding to Or47a, Or43a, Or43b and Or22a) there was more than one probable ligand (Supplementary Table 3). We then used the maximum common sub-structure (MCS) method of Cao et al.29 to predict the affinity of ligands to individual components of QMP. For each test, we used the ChemMine application41 that generates a Tanimoto42 ‘similarity score’ for each pair of compounds (MSC Ts). In this context, a high score (maximum of ‘1’) implies a higher chemical identity between fly ligand and bee pheromone. In total we identified n = 21 ligands corresponding to the 16 unique receptors identified from our RNAi screen. The structural similarity scores between ligand and QMP component ranged from 0.20–0.83, suggesting that sub-structural analysis contains ample variation to predict biological affinity.
Additional Information
How to cite this article: Camiletti, A. L. et al. A novel screen for genes associated with pheromone-induced sterility. Sci. Rep. 6, 36041; doi: 10.1038/srep36041 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hamilton, W. D. The genetical evolution of social behaviour, I and II. J. Theoret. Biol. 7, 1–52 (1964).
Thompson, G. J., Hurd, P. L. & Crespi, B. J. Genes underlying altruism. Biol. Lett. 9, 20130395 (2013).
Keller, L. Adaptation and the genetics of social behaviour. Philos. Trans. R. Soc. B-Biol. Sci. 364, 3209–3216 (2009).
Linksvayer, T. A. The molecular and evolutionary genetic implications of being truly social for the social insects. Adv. Insect Physiol. 48, 271–292 (2015).
Denison, R. & Raymond-Delpech. Insights into the molecular basis of social behaviour from studies on the honeybee, Apis mellifera. Invert. Neurosci. 8, 1–9 (2008).
Weinstock, G. M. et al. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949 (2006).
Cardoen, D. et al. Genome-wide analysis of alternative reproductive phenotypes in honeybee workers. Mol. Ecol. 20, 4070–4084 (2011).
Grozinger, C. M., Fan, Y. L., Hoover, S. E. R. & Winston, M. L. Genome-wide analysis reveals differences in brain gene expression patterns associated with caste and reproductive status in honey bees (Apis mellifera). Mol. Ecol. 16, 4837–4848 (2007).
Thompson, G. J., Kucharski, R., Maleszka, R. & Oldroyd, B. P. Genome-wide analysis of genes related to ovary activation in worker honey bees. Ins. Mol. Biol. 17, 657–665 (2008).
Ratnieks, F. L. W. & Helantera, H. The evolution of extreme altruism and inequality in insect societies. Philos. Trans. R. Soc. B-Biol. Sci. 364, 3169–3179 (2009).
Peso, M., Elgar, M. A. & Barron, A. B. Pheromonal control: reconciling physiological mechanism with signalling theory. Biol. Rev. 90, 542–559 (2015).
Smith, C. R., Toth, A. L., Suarez, A. V. & Robinson, G. E. Genetic and genomic analyses of the division of labour in insect societies. Nat. Rev. Genet. 9, 735–748 (2008).
Camiletti, A. L., Awde, D. N. & Thompson, G. J. How flies respond to honey bee pheromone: the role of the foraging gene on reproductive response to queen mandibular pheromone. Naturwissenschaften 101, 25–31 (2014).
Camiletti, A. L., Percival-Smith, A. & Thompson, G. J. Honey bee queen mandibular pheromone inhibits ovary development and fecundity in a fruit fly. Entomologia Experimentalis et Applicata 147, 262–268 (2013).
Camiletti, A. L. & Thompson, G. J. Drosophila as a genetically tractable model for social insect behaviour. Front. Ecol. Evol. 4, 40 (2016).
Larsson, M. C. et al. Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 43, 703–714 (2004).
Steck, K. et al. A high-throughput behavioral paradigm for Drosophila olfaction - The Flywalk. Scientific Reports 2, 361 (2012).
Libert, S. et al. Regulation of Drosophila life span by olfaction and food-derived odors. Science 315, 1133–1137 (2007).
Robertson, H. M. & Wanner, K. W. The chemoreceptor superfamily in the honey bee, Apis mellifera: Expansion of the odorant, but not gustatory, receptor family. Genome Res. 16, 1395–1403 (2006).
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Zhou, X. F. et al. Phylogenetic and transcriptomic analysis of chemosensory receptors in a pair of divergent ant species reveals sex-specific signatures of odor coding. PLoS Genet. 8, e1002930 (2012).
Schulte, C., Theilenberg, E., Muller-Borg, M., Gempe, T. & Beye, M. Highly efficient integration and expression of piggyBac-derived cassettes in the honeybee (Apis mellifera). Proc. Natl. Acad. Sci. USA 111, 9003–9008 (2014).
Ben-Shahar, Y. A piggyBac route to transgenic honeybees. Proc. Natl. Acad. Sci. USA 111, 8708–8709 (2014).
Oi, C. A. et al. The origin and evolution of social insect queen pheromones: Novel hypotheses and outstanding problems. Bioessays 37, 808–821 (2015).
Hallem, E. A. & Carlson, J. R. Coding of odors by a receptor repertoire. Cell 125, 143–160 (2006).
Stensmyr, M. C. et al. A conserved dedicated olfactory circuit for detecting harmful microbes in Drosophila. Cell 151, 1345–1357 (2012).
Benton, R. & Dahanukar, A. Electrophysiological recording from Drosophila olfactory sensilla. Cold Spring Harb. Protoc. 7, 824–838 (2011).
Galizia, C. G., Munch, D., Strauch, M., Nissler, A. & Ma, S. W. Integrating heterogeneous odor response data into a common response model: A DoOR to the complete olfactome. Chem. Senses 35, 551–563 (2010).
Cao, Y. Q., Jiang, T. & Girke, T. A maximum common substructure-based algorithm for searching and predicting drug-like compounds. Bioinformatics 24, I366–I374 (2008).
Beggs, K. T. et al. Queen pheromone modulates brain dopamine function in worker honey bees. Proc. Natl. Acad. Sci. USA 104, 2460–2464 (2007).
Rehan, S. M. & Toth, A. L. Climbing the social ladder: the molecular evolution of sociality. Trends Ecol. Evol. 30, 426–433 (2015).
Amdam, G. V., Csondes, A., Fondrk, M. K. & Page, R. E. Complex social behaviour derived from maternal reproductive traits. Nature 439, 76–78 (2006).
Van Oystaeyen, A. et al. Conserved class of queen pheromones stops social insect workers from reproducing. Science 343, 287–290 (2014).
Sakurai, A., Koganezawa, M., Yasunaga, K., Emoto, K. & Yamamoto, D. Select interneuron clusters determine female sexual receptivity in Drosophila. Nat. Commun. 4, 1825 (2013).
Plettner, E. et al. Species- and caste-determined mandibular gland signals in honeybees (Apis). J. Chem. Ecol. 23, 363–377 (1997).
Sannasi, A. Inhibition of ovary development of the fruit-fly, Drosophila melanogaster by synthetic ‘queen substance’. Life Sciences 8, 785–789 (1969).
Pankiw, T. et al. Mandibular gland components of European and Africanized honey bee queens (Apis mellifera L). J. Chem. Ecol. 22, 605–615 (1996).
King, R. C. Ovarian development in Drosophila melanogaster. Pp. 227 (Academic Press, 1970).
Sweeney, S. T., Broadie, K., Keane, J., Niemann, H. & Okane, C. J. Targeted expression of tetannus toxin light-chain in Drosophila specifically eliminates synaptic transmission and causes behavioral defects. Neuron 14, 341–351 (1995).
Slessor, K. N., Kaminski, L. A., King, G. G. S. & Winston, M. L. Semiochemicals of the honeybee queen mandibular glands. J. Chem. Ecol. 16, 851–860 (1990).
Backman, T. W. H., Cao, Y. Q. & Girke, T. ChemMine tools: an online service for analyzing and clustering small molecules. Nucleic Acids Res. 39, W486–W491 (2011).
Rogers, D. J. & Tanimoto, T. T. Computer program for classifying plants. Science 132, 1115–1118 (1960).
Acknowledgements
We thank Emma K Mullen, Onyka Gairey, Zachary Dloomy, Brandon Budhram, Christine Scharf, Anne F Simon, Jeremy N McNeil and all members of the Social Biology Group at Western University (Canada) for help and discussion. This work was funded by an Ontario Graduate Scholarship to A.L.C. and a Natural Sciences and Engineering Research Council of Canada Discovery Grant to G.J.T. In addition, we thank the Herbette Foundation (La Fondation Herbette) for facilitating G.J.T.’s stay at the University of Lausanne.
Author information
Authors and Affiliations
Contributions
A.L.C., A.P.-S. and G.J.T. conceived the study. A.L.C. performed the experiments, with assistance from J.R.C., A.L.C. and G.J.T. performed the analyses and wrote the paper. All authors edited and finalized the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Camiletti, A., Percival-Smith, A., Croft, J. et al. A novel screen for genes associated with pheromone-induced sterility. Sci Rep 6, 36041 (2016). https://doi.org/10.1038/srep36041
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36041
- Springer Nature Limited
This article is cited by
-
Sexual response of male Drosophila to honey bee queen mandibular pheromone: implications for genetic studies of social insects
Journal of Comparative Physiology A (2017)