Abstract
The morphological diversity of insects is important for their survival; in essence, it results from the differential expression of genes during development of the insect body. The silkworm apodal (ap) mutant has degraded thoracic legs making crawling and eating difficult and the female is sterile, which is an ideal subject for studying the molecular mechanisms of morphogenesis. Here, we confirmed that the infertility of ap female moths is a result of the degradation of the bursa copulatrix. Positional cloning of ap locus and expression analyses reveal that the Bombyx mori sister of odd and bowl (Bmsob) gene is a strong candidate for the ap mutant. The expression of Bmsob is down-regulated, while the corresponding Hox genes are up-regulated in the ap mutant compared to the wild type. Analyses with the dual luciferase assay present a declined activity of the Bmsob promoter in the ap mutant. Furthermore, we demonstrate that Bmsob can inhibit Hox gene expression directly and by suppressing the expression of other genes, including the BmDsp gene. The results of this study are an important contribution to our understanding of the diversification of insect body plan.
Similar content being viewed by others
Introduction
The morphological diversity of insects makes them able to adapt rapidly to environmental changes and provides good models for studying body plan diversity1. The identification of key genes and the elucidation of pathways relating to the morphogenesis and pattern formation are of great interest. Insects share some common features; for example, the trunk is composed of segments that can be grouped into a head, a thorax and an abdomen. By contrast, insect appendages have morphological and functional differences despite the same basic types of appendage along the body axis2,3. Based on studies of Drosophila, genes involved in the formation of body segments can be divided into four conceptual categories; maternal effect, gap, pair-rule and segment polarity4,5,6. Segmental identity was determined along the anterior-posterior body axis by the homeotic (Hox) genes, whose expression patterns largely determine the morphology and development of tissues and organs7,8,9. Changes in Hox genes can lead to changes in insect body plan and morphology1,10. Despite our increased knowledge and understanding of insect morphogenesis, details of genes responsible for segmentation and their regulatory networks in non-Drosophilia insects remain obscure.
In Drosophila, the odd-skipped gene family consists of odd-skipped (odd), brother of odd with entrails limited (bowl), sister of odd and bowl (sob) and drumstick (drm), which have key roles during embryogenesis11,12,13. The odd gene, which was identified as a member of the pair-rule genes, encodes a C2H2 zinc finger transcription factor regulating the development of multiple tissues14,15,16,17,18. The bowl and drm genes are required for leg, notum and hindgut pattern formation17,19,20. The identification and analysis of related developmental mutants has contributed to our understanding of the odd-skipped family. Although recent studies suggested the sob gene might be involved in the development of antennae, legs and wings21,22,23,24, the particular role of this gene in insect morphogenesis is not clear.
Wings, ventral limbs and the reproductive system have pivotal roles in insects; hence, the identification of the corresponding developmental genes is of particular importance. In the silkworm Bombyx mori, the apodal (ap) mutant exhibits marked multiple morphological variations, including degraded thoracic legs and wings as well as female sterility, which is an ideal subject for the study of the molecular mechanisms for morphogenesis. The spontaneous ap mutant was identified by the presence of degraded thoracic legs throughout development and the ap adults exhibited smaller or deformed wings seriously affecting their ability to mate (Fig. 1). For all of these reasons, the ap mutant could not mate naturally and required artificial mating to produce normal offspring. Genetic analysis has revealed that it is located at 22.3 centimorgans (cM) on the third linkage in the silkworm genetic linkage map and controlled by a single recessive gene25. Moreover, the silkworm E complex mutants, located on chromosome 6 corresponding to the Hox genes, display ectopic legs, abnormal wings and genital system25,26, implying a relationship between the ap mutant and Hox genes.
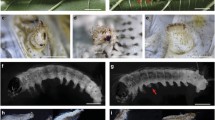
Phenotypes of wild type and apodal (ap) mutant.
(a) Wild-type (ap/+ap) embryo. The arrow indicates the normal thoracic legs. Bar = 0.5 mm. (b) The ap mutant (ap/ap) embryo. The arrow indicates degraded thoracic legs. Bar = 0.5 mm. (c) Wild-type (ap/+ap) larva. The arrow indicates the normal thoracic legs. Bar = 1 cm. (d) The ap mutant (ap/ap) larva. The arrow indicates degraded thoracic legs. Bar = 1 cm. (c′) Enlargement of the thoracic leg in (c). Bar = 1 mm. (d′) Enlargement of the degraded thoracic leg in (d). Bar = 1 mm. (e) Dorsal side of the wild-type (ap/+ap) adult. Bar = 1 cm. (f) Lateral view of the wild-type (ap/+ap) adult. The arrow indicates the normal thoracic legs. Bar = 1 cm. (g) Dorasl side of the ap mutant (ap/ap) adult. Bar = 1 cm. The wings of the ap mutant adult were smaller compared to the wild type (e). (h) Ventral side of the ap mutant (ap/ap) adult. The arrow indicates degraded thoracic legs. Bar = 1 cm.
In this study, we investigated the physiological cause of ap female infertility and then identified and characterized the strong candidate gene responsible for the ap mutant. We confirmed that the infertility of ap female moths was due to the degradation of the bursa copulatrix and demonstrated that the abnormal expression of the Bmsob gene encoding C2H2 zinc finger-containing transcriptional factor should be probably responsible for ap mutant. Finally, we found that the Hox genes in the ap mutant are abnormally expressed and verified the Hox genes can be regulated by the Bmsob gene.
Results
The female ap mutant exhibited degenerated bursa copulatrix
In addition to the degraded thoracic legs and abnormal wings of the ap mutant (Fig. 1), another striking feature of the ap mutant was the female sterility, which had a significant effect on fertility. To explore the underlying cause of female sterility, further, we examined the reproductive system of ap mutant female moths from several aspects. We verified that the ap female moth could make eggs like the wild type, although the number of eggs varied more between ap individuals (Supplementary Fig. S1 and Table S1). We investigated the micropyle to make sure sperm can enter the egg smoothly and found that the micropyles of ap eggs were normal (Supplementary Fig. S2). We used artificial parthenogenesis on wild type and ap eggs and found about 50% of both can develop normally (Supplementary Fig. S3 and Table S2), suggesting that the contents of ap eggs are normal. According to these comparative experiments, our results indicated that ap female sterility was not caused by malformation of eggs. Next, we searched for evidence of structural variation in ap female internal genitalia. We found that the bursa copulatrix, which can store seminal fluid temporarily during the mating process, was degenerate in the ap mutant female moths (Fig. 2a,b). The normal bursa copulatrix of wild type B. mori is a sac-like structure and becomes club-shaped after mating (Fig. 2c,e), whereas the ap bursa copulatrix had only vestiges (Fig. 2d–f). Collectively, these studies revealed that ap female sterility was due to the degenerate structure of the bursa copulatrix.
Genital glands of the wild type and ap mutant female adult.
(a) The genital gland of the ap/+ap female adult. The female moths were mated for 6 h before dissection. The arrow indicates the normal bursa copulatrix with seminal fluid. (b) The genital gland of the ap/ap female adult. The female moths were mated for 6 h before dissection. The arrow indicates the degraded bursa copulatrix. (c) Enlargement of the bursa copulatrix in (a). (d) Enlargement of the bursa copulatrix in (b). (e) Bursa copulatrix of the ap/+ap female moth without mating. (f) Bursa copulatrix of the ap/ap female moth without mating. The green dots indicated the outline of the bursa copulatrix and the blue dots indicated the sperm canal. Bar = 1 mm.
Positional cloning of the ap locus
We used a map-based cloning approach to investigate the B. mori genome sequence in an attempt to identify the gene responsible for the ap locus. In the B. mori linkage map, the ap locus was approximately 22.3 cM from the distal lem locus, which was reported to result from the mutation of BmSpr, a gene located on Scaffold 293125,27 (Fig. 3a,b). In addition, the Ze locus was reported to be linked tightly with the FL0827 marker, which was located on Scaffold 293028 (Fig. 3a,b). Hence, we used the BmSpr gene and FL0827 as anchor markers of the genome sequence to find new markers.
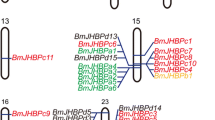
Mapping of the ap locus on the B. mori linkage group 3.
(a) The 3rd linkage group of B. mori. The mutation symbols are shown above the map and the corresponding locus in centiMorgans (cM) are shown below the map. (b) Genomic scaffolds on chromosome 6. Orange boxes represent the assembled scaffolds and the names of the scaffolds are shown below. (c) Scaffold map and gene model of the nscaf2930 and nscaf2924 scaffolds on chromosome 3. The markers are identified above the map and the numerals below the map indicate the number of recombinants identified in 384 F2 progeny. Blue arrows represent the predicted genes in SilkDB and red arrows represent the validated genes.
To map the ap locus more precisely, we generated an F2 population of 384 individuals with the ap mutant phenotype produced by selfing F1 progeny of a cross between a Dazao female and an ap male. We mapped the ap locus to a region between two markers, A60 and A73 and the markers A54 and A85 were linked tightly with the ap locus (Fig. 3c). Within the region between the markers A60 and A73, only two genes BGIBMGA008843 and BGIBMGA008844 were predicted to be present, while there is a gap region that has not been sequenced on the scaffolds (Fig. 3c). Both of the predicted genes had high levels of homology to sister of odd and bowl (sob) of Drosophila melanogaster, which is one member of the odd-skipped family that involved in Drosophila leg development17. Comparison and assembly of sequences of these two predicted genes and expressed sequence tags (ESTs) resulted in the dependable conclusion that they were actually one gene, designated as Bombyx mori sister of odd and bowl (Bmsob). Therefore, we next focused on the Bmsob gene that is the unique gene having sequence information within the mapped region.
Subsequently, according to the two predicted genes and ESTs, we obtained the full-length cDNA of Bmsob gene in the wild type and the ap mutant using rapid amplification of cDNA ends (RACE) technology, 1923 bp and 1924 bp, respectively (Supplementary Fig. S4). The length of the coding sequence region was 1311 bp, encoding 436 amino acids with five C2H2-type zinc finger structures. The 5′-untranslated region (5′-UTR) of both was 85 bp long and the 3′-UTR was 527 bp and 528 bp long, respectively and both had two exons. There were 15 single base mutations in Bmsob open reading frames (ORFs) between Dazao and the ap mutant, of which 13 were synonymous mutations and two were missense mutations (P → A, S → P). In addition, there were eight single base mutations and one base insertion in the Bmsob UTR in the ap mutant. In order to study the relationship between the candidate gene and the ap mutant further, we undertook a more detailed analysis of the Bmsob gene.
Expression profiles of the Bmsob gene
Multiple defects associated with the ap mutant suggest its candidate gene is expressed in particular spatial temporal patterns. Therefore, we undertook a detailed analysis of the temporal expression pattern of the Bmsob gene. Given that the embryonic period is crucial for the development of limbs, we examined the temporal expression pattern of Bmsob mRNA at nine different developmental stages of the embryo and newly hatched larvae. The semi-quantitative RT-PCR analysis showed that expression of the Bmsob gene was detected first after oviposition and then increased gradually; it reached a peak level at 4 days after egg laying and decreased gradually thereafter (Fig. 4a). Remarkably, the high-level expression of Bmsob gene in stages 2, 3 and 4 days was consistent with silkworm limb development, suggesting that the Bmsob gene is involved in the early limb development.
Temporal expression profiles of the Bmsob gene.
(a) Expression profile of the Bmsob gene during silkworm embryonic development. NHL, newly hatched larvae. (b) Temporal expression of Bmsob in the silkworm from the 4th instar larvae to the pupa stage. Total RNA samples were isolated from the whole bodies. (c) Temporal expression profile of the Bmsob gene in the development of the wing. Total RNA samples were isolated from the wing discs of larvae and the wings of pupae and moths. The Bombyx mori Actin 3 gene was used as internal control. d, day; 4th, fourth instar; 5th, fifth instar; W, wandering stage; P, pupa stage; P-M, the day before the moth; M, moth.
As the silkworm undergoes complete metamorphosis and the late larval stage to pupal stage is especially important for metamorphosis, we examined the temporal expression pattern of Bmsob mRNA from the 4th instar day 4 larva to the day 5 pupae. Our results showed that high-level expression of Bmsob gene appeared at the third day of the wandering stage and the first day of the pupal stage (Fig. 4b). This period is the key stage in the metamorphosis from larva to pupa and the important stage of genital gland development, implying that the Bmsob gene has an important role in the development of the silkworm genital gland. In addition, analysis of Bmsob gene expression pattern during wing development showed that the expression level of Bmsob increased gradually with development of the wing primordium (Fig. 4c). The level of Bmsob expression was highest in the final stage of the 5th instar, implying that the Bmsob gene is involved in the development and formation of silkworm wings. Together, complex patterns of expression suggested extensive roles for the Bmsob gene during development of the silkworm.
Expression level of Bmsob gene in the ap mutant compared to wild-type strain Dazao
To assess the relationship between the ap mutant and the Bmsob mutated state, we compared the expression of Bmsob between ap and Dazao at two stages of development, day 4 of the embryo and the first day of pupation, when Bmsob normally showed its highest level of activity. The results of the quantitative RT-PCR (qRT-PCR) showed that the expression level of Bmsob was decreased markedly in the ap mutant compared to the wild type at both stages (Fig. 5a,b), while there was no significant difference in the expressions of Bmdrm and Bmbowl genes between the wild type and ap mutant (Supplementary Fig. S5). Because there was no significant difference in the coding region of the Bmsob gene in the ap mutant compared to the wild type, we examined the promoter region to explore the reason for the reduction of Bmsob expression in the ap mutant. We found sequence polymorphisms in the Bmsob promoter region in the ap mutant and in the Dazao stain (Supplementary Fig. S6). Then we detected the promoter activity in the wild type and the ap mutant using the dual luciferase report system and found that the level of activity of the Bmsob promoter in the ap mutant was decreased significantly (Fig. 5c,d). Accordingly, we speculated that the sequence variation of the Bmsob promoter region might be a significant reason for the decreased expression level of the Bmsob gene.
Analysis of the Bmsob gene and its promoter region in the wild type (Dazao) and the ap mutant.
(a) Relative quantitative analysis of Bmsob in Dazao and ap mutant embryos. (b) Relative quantitative analysis of Bmsob in Dazao and ap mutant pupae (1 d). (c) A diagram of the construction of the luciferase reporter gene vectors. (d) The relative activity of the Bmsob promoter in Dazao and the ap mutant. DZ, Dazao; **P < 0.01, Student’s t-test, n = 3.
Bmsob is involved in maintenance of expression of the Hox genes
The published results of studies on the functions of Hox genes led us to speculate that the appearance of abnormal thoracic legs, wings and bursa copulatrix in ap mutant might be related to the B. mori Hox genes. We used qRT-PCR to examine the expression levels of Hox genes BmAntp, BmUbx and BmAbd-B; the results showed that the expression of these three genes was up-regulated in the ap mutant compared to the wild type (Fig. 6a). To confirm that the Bmsob gene was responsible for the abnormal expression levels of these three Hox genes, we used electrophoretic mobility shift assay (EMSA) experiments to validate interactions between the Bmsob protein and the BmAntp, BmUbx and BmAbd-B genes. The results showed that the Bmsob protein could bind directly to the BmUbx gene but not to the BmAntp or BmAbd-B genes (Fig. 6b–d).
Expression profiles of Hox genes in Dazao and the ap mutant and the interaction between the Bmsob protein and Hox genes.
(a) Expression profiles of Hox genes in Dazao and the ap mutant. **P < 0.01, Student’s t-test, n = 3. Detection by EMSA of (b) the interaction between the Bmsob protein and the BmAntp gene; (c) the interaction between the Bmsob protein and the BmUbx gene; (d) the interaction between the Bmsob protein and the BmAbd-B gene. The amount of GST protein used in the reactions was 1 μg and the amount of Bmsob protein was 0.8 μg. The amount of biotin labeling probe, in turn, was 1, 2 and 6 pmol. The amount of unlabeled probe, in turn, was 2, 20 and 100 pmol. The arrow indicates the protein-DNA complex.
Further, in order to verify whether Hox genes can be regulated indirectly by the Bmsob gene, we screened the differentially expressed genes by microarray comparison of the wild type and the ap mutant (see Supplementary data for a complete list). Notably, the Dsp gene, which encodes a high-mobility group-like protein, appeared on our microarray list and exhibited a marked increase of Dsp expression in the ap mutant. The qRT-PCR result was consistent with the microarray data (Fig. 7a). In particular, its Drosophila homologous gene dsp1 participated in the regulation of Hox genes29. The EMSA results confirmed that the Bmsob protein can bind to the BmDsp gene (Fig. 7b) and further results showed that the BmDsp protein can bind to the BmAntp, BmUbx and BmAbd-B genes (Fig. 7c–e). These findings led to the hypothesis that the Bmsob gene can inhibit Hox gene expression directly and indirectly through suppressing the expression of other genes, such as the BmDsp gene, which can promote expression of the Hox gene (Fig. 7f).
Expression profiles of the BmDsp gene in Dazao and the ap mutant, the interaction between the Bmsob protein and the BmDsp gene and the interaction between the BmDsp protein and the Hox genes.
(a) Expression profiles of BmDsp genes in Dazao and the ap mutant. **P < 0.01, Student’s t-test, n = 3. Detection by EMSA of (b) the interaction between the Bmsob protein and the BmDsp gene; (c) the interaction between the BmDsp protein and the BmAntp gene; (d) the interaction between the BmDsp protein and the BmUbx gene; (e) the interaction between the BmDsp protein and the BmAbd-B gene. The amount of protein used in each reaction was: GST 1 μg; Bmsob 0.8 μg; BmDsp 0.8 μg. The amount of biotin labeling probe, in turn, was 1, 2 and 6 pmol. The amount of unlabeled probe, in turn, was 2, 20 and 100 pmol. The arrow indicates the protein-DNA complex. (f) Predicted regulation relationship between the Bmsob, BmDsp and Hox genes.
Discussion
In this study, we demonstrated that the Bmsob gene is a strong candidate for the B. mori ap mutant which generates multiple defects by fine mapping and gene expression analysis. We confirmed that the infertility of ap female moths was due to the degradation of the bursa copulatrix. Moreover, we examined the relationship between the candidate gene Bmsob and the Hox genes, whose expression are essential for the proper organization of the animal body plan during development, indicating that Bmsob gene can inhibit Hox gene expression directly and indirectly through suppressing other genes, such as BmDsp. The results suggest that this regulatory network has an important role in the precise expression pattern of the Hox genes during development of the silkworm body plan.
The odd-skipped family genes encode zinc-finger transcription factors and are conserved in insects and vertebrates. Phylogenetic analysis showed that the silkworm-conserved odd gene family contained four genes belonging to four subfamilies, all of which had a single copy (Supplementary Fig. S7). Earlier studies showed that odd gene has widespread roles in embryonic development16,17,18, but few reports on the sob gene are available. Recent studies of T. castaneum by RNA interference (RNAi) screening showed that the sob gene might be involved in the development of legs and wings22,24. Similarly, microarray comparison analysis indicated that the Drosophila sob gene participates in the wing development23. Our results indicated that Bmsob as the only predicted gene on the scaffolds within our mapping region should be probably responsible for the multiple mutation phenotypes of the ap mutant, suggesting that Bmsob has important roles in the development of legs, wings and the reproductive system in the silkworm. Furthermore, the expression patterns of the Bmsob gene are consistent with the development of the embryo and wings (Fig. 4a,c). Especially, the massively high expression levels of the Bmsob gene occurred at the W-3d and P-1d stages, when the reproductive system was undergoing a violent metamorphosis (Fig. 4b). These data further implied an important role of the Bmsob gene during the development of these tissues and, according to its specific expression patterns in different tissues, we suggest that there are some gene-specific cis-regulatory elements for the Bmsob gene.
Although we have not identified large changes within the Bmsob gene in the ap mutant that predominantly affects its structure, we identified 15 single-base mutations in the Bmsob gene ORF between the wild type and the ap mutant, two of which cause missense mutations (P → A, S → P) and eight single base mutations and one base insertion into the UTR (Supplementary Fig. S4). Considering the decreased expression of the Bmsob gene in the ap mutant, we detected the promoter activity of Bmsob and the results showed that the activity of the Bmsob promoter in the ap mutant was decreased significantly (Fig. 5d). These data suggest that the ap mutant was not caused by large changes in the Bmsob coding region, such as deletions or insertions, but rather by changes with a few single nucleotide and/or in the Bmsob gene upstream regulatory sequences. Based on our mapping data, future studies will be aimed at identifying novel regulatory elements of the Bmsob gene in the responsible region. In addition, we found that the three genes, drm, sob and odd, were clustered in D. melanogaster, Tribolium castaneum and Heliconius melpomene30,31,32. Therefore, the Bmodd gene probably locates in the gap region. Considering of the fact that we did not obtain any significant phenotypes by Bmsob gene RNAi, we can’t rule out the possibility that the Bmodd gene may also contribute to ap mutant. On the basis of these results, our future research will pay some attention to test the possibility.
Numerous studies have demonstrated that the function of a Hox gene is related closely to its expression patterns33. In Drosophila, the Antp, Ubx and Abd-B genes were involved in development of the thoracic leg, wing and gonad respectively, which corresponds to their functional domain34,35,36. Similarly, our previous study suggested that the Antp gene is required for the organization of thoracic legs in B. mori37. These results suggested that multiple mutation phenotypes of the ap mutant might be related to the expression of the corresponding Hox gene and provide clues for studying the mechanism underlying Hox gene expression. In the present study, our data showed that the expression of the Antp, Ubx and Abd-B genes was abnormal in the ap mutants, suggesting that there might be a regulatory relationship between them and the Bmsob gene. Subsequently, EMSA analysis revealed that the Bmsob gene could bind to the predicted transcription factor binding sites upstream of the Ubx gene (Fig. 6c), indicating that it might be regulated directly by the Bmsob gene. By contrast, our results showed that Bmsob might regulate the expression of Hox genes indirectly through other genes, such as BmDsp. Taken together, our results indicate that the product of the Bmsob gene would act as a repressor of target gene expression and the Hox genes, especially Antp, Ubx and Abd-B could be its important target genes in B. mori.
Uncovering the molecular genetic basis of mutations provides particularly significant clues for understanding the underlying mechanisms of animal development. Conspicuously, the recessive ap mutations have multiple developmental defects that are related to their survival and reproductive capacity, such as degraded thoracic legs, wings and bursa copulatrix, providing good targets for the study of pest control. In particular, studying female sterility can be used to make improvements to a highly effective area-wide method of pest control, known as the sterile insect technique38,39. Based on these results, our future work will concentrate on the inducement of pest sterility by genetic manipulation.
Methods
Silkworm strains and cell lines
The wild-type strain Dazao and the ap mutant strain (ap/ap) were obtained from the Silkworm Gene Bank at Southwest University, Chongqing, China. Silkworms were reared with fresh mulberry leaves at 25 °C under a photoperiod of 12h light/12h dark. The B. mori ovary cell line BmN-SWU140 was cultured in TC-100 (US Biological) medium containing 10% (v/v) fetal bovine serum (Gibco), penicillin G (200 U/ml) and streptomycin sulfate (200 U/ml) at 27 °C.
Scanning electron microscopy and artificial parthenogenesis
Eggs from non-mated females were dissected and rinsed with double-distilled water. For scanning electron microscopic observation, the eggs were processed as previously described37 and were coated with gold before observation under a scanning electron microscope (S-3000N, Hitachi, Tokyo, Japan). For artificial parthenogenesis, the newly dissected eggs were treated according to the following steps: 1) incubated at 25 °C for 12 h; 2) placed in the hot water at 46 °C for 18 min; 3) taken out of the water and incubated at 25 °C for 10 min; 4) transferred to 16 °C for 3 days; 5) treated with hydrochloric acid (sp. gr. 1.075) at 46 °C for 5 min; 6) incubated at 25 °C with adequate humidity.
Positional cloning of the ap locus
Owing to the female sterility of the ap mutant, we used a single-pair cross between a Dazao female and an ap male to produce F1 offspring. Because there is no recombination in female silkworms, 22 progeny from a single-pair backcross between an F1 female and an ap male were used for the linkage analysis and 384 F2 progeny with the ap phenotype from selfing F1 (Dazao × ap) × (Dazao × ap) were used for the recombination analysis. Genomic DNA was extracted from parent moths, F1 moths and each F2 individual of third instar larvae as previously described37. Primer sets were designed from the B. mori genome sequence41 and the markers showing polymorphisms between Dazao and ap were used for linkage analysis (Supplementary Table S3) with MAPMAKER/EXP 3.042 using the Kosambi function43. The bioinformatics analysis of candidate genes was done as previously described37.
Rapid amplification of cDNA ends (RACE)
Total RNA of 7 days old Dazao embryos was isolated as previously described37. The 5′ and 3′ RACE was performed using Gene Racer Kit (Invitrogen, Carlsbad, CA, USA), according to the manufacturers’ protocol. PCR products were cloned into the pMD19-T vector (TaKaRa, Dalian, China) and sequenced. The primers used for PCR are listed in Supplementary Table S3.
RT-PCR and quantitative RT-PCR (qRT-PCR)
To analyze gene expression patterns, the semi-quantitative RT-PCR experiments were performed. Total RNAs were isolated with TRIzol® reagent (Invitrogen) from Dazao wild-type embryos of different developmental stages (from newly laid eggs to newly hatched larvae), the whole body of several developmental stages (from day 4 of the 4th instar larvae to day 5 of the pupae) and the wing discs of larvae and the wings of pupae and moths (from day 1 of the 5th instar larvae to the adult moth). The cDNA was synthesized using the PrimeScript reagent Kit with gDNA Eraser (TaKaRa) according to the manufacturer’s protocol. The gene for Actin3 of B. mori was used as an internal control. For qRT-PCR, total RNA was isolated from 4 days old Dazao and ap embryos and reverse transcription was carried out as described above. The ap/ap embryos were from ap/+ females × ap/ap males and we distinguished ap/ap embryos from ap/+ by the apodal phenotype. The eukaryotic translation initiation factor 4A (silkworm microarray probe ID: sw22934) was used as an internal control. The qRT-PCR experiments were performed using the CFX96 Real-Time System (Bio-Rad, Hercules, CA, USA) with an iTaq Universal SYBR Green Supermix (Bio-Rad), according to the manufacturer’s recommended procedure. The specific primers for RT-PCR and qRT-PCR are listed in Supplementary Table S3.
Phylogenetic analysis
Protein sequences were downloaded from the following websites: National Center for Biotechnology Information (NCBI) Protein Database (http://www.ncbi.nlm.nih.gov/protein), SilkDB41 (http://silkworm.genomics.org.cn/), Flybase30 (http://flybase.org/), BeetleBase31 (http://beetlebase.org/), VectorBase44 (https://www.vectorbase.org/) and Heliconius Genome32 (http://www.butterflygenome.org/). Accession numbers are listed in Supplementary Table S4. A multiple sequence alignment of the complete protein sequences was performed using MUSCEL program with default parameters. The phylogenetic tree was constructed using the Bayesian approach. Bayesian inferences were performed using MrBayes v3.1.245 with the VT model estimated by protest-3.2 software46. We performed 2,500,000 generations and other parameters were set as default. The phylogenetic tree was visualized with MEGA 647.
Dual luciferase reporter assays
We constructed recombinant plasmids from a pfsLSV40 vector (a gift from Dr. Wang, Southwest University) that contained the luciferase gene driven by a Bmactin4 promoter. The promoter region of the sob gene amplified from the Dazao and ap strains was cloned into pfsLSV40 to replace the Bmactin4 promoter, respectively. Primers are listed in Supplementary Table S3. The resulting pfsLSV40-Bmsob-Dazao and pfsLSV40-Bmsob-ap plasmids were used for analyzing the promoter activity. BmN-SWU1 cells (1 × 106) were seeded in 24-well culture plates (Corning Glass Works, Corning, NY, USA). After 24 h, cells were co-transfected with 400 ng pfsLSV40-Bmsob-Dazao plasmid or pfsLSV40-Bmsob-ap plasmid and 100 ng pIZ-Rluc control plasmid. At 48 h after transfection, a GloMax-Multi Detection System and a Dual-Glo Luciferase Assay Kit (Promega, Madison, WI, USA) were used to quantify luciferase activity. All assays were performed in triplicate.
Microarray analysis
Total RNA was extracted from the wild type and ap mutant embryos as described above. For the hybridization experiment, we mixed equal amounts of total RNA from at least seven biological replicates to create one sample. Microarray hybridization and raw data normalization were carried out by Capital Bio Corp. (Beijing, China) as described48. For the microarray data analysis, a gene was considered as up- or downregulated in the ap mutant compared to the wild type if it displayed at least a twofold change.
Recombinant expression and protein purification
The ORF of Bmsob and BmDsp were cloned into the pGEX-4T-1 vector with a glutathione S-transferase (GST) tag and pET-28a (+) vector with a His tag, respectively. Primers are listed in Supplementary Table S3. The proven clones were transformed into Escherichia coli strain BL21 to express GST-tagged Bmsob (GST-sob) and His-tagged BmDsp (His-Dsp) proteins. The GST-sob protein and His-Dsp protein were purified using a GSTrap FF column and a HisTrap HP column (GE Healthcare, Freiburg, Germany), respectively, according to the manufacturers’ protocol. Purified proteins were verified through western blotting as described previously49 with anti-GST tag and anti-His tag antibodies (Beyotime, Jiangsu, China), respectively.
Electrophoretic mobility shift assay (EMSA)
For the prediction of Bmsob and BmDsp binding sites, we used two different transcription factor binding site prediction programs, Tfsitescan (http://www.ifti.org/cgi-bin/ifti/Tfsitescan.pl) and PROMO (http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3). The features related to the gene binding sites are G+C-rich and A+T-rich sequences for Bmsob and BmDsp, respectively, in accordance with their structural domains50,51. For EMSA, the probes were 5′-labeled by using biotin (Invitrogen) and then the labeled oligonucleotides were annealed to produce a double-stranded probe. The probes are listed in the Supplementary Table S3. The EMSA assay was performed with a LightShift Chemiluminescent EMSA Kit (Pierce, Rockford, IL, USA) according to the manufacturers’ protocol. After incubation, reaction products were loaded on to 5% (w/v) polyacrylamide non-denaturing gel (acrylamide:bis-acrylamide 19/1, w/w) and electrophoresed in TBE buffer (89 mM Tris, 89 mM boric acid, 2 mM EDTA, pH 8.3) for approximately 1.5 h at 100V. Then, the protein-DNA complexes were transferred electrophoretically from the gel onto nylon membranes (Invitrogen) for approximately 45 min at 380 mA and visualized with streptavidin-horseradish peroxidase conjugate (Pierce) according to the manufacturer’s protocol.
Additional Information
How to cite this article: Chen, P. et al. Molecular mapping and characterization of the silkworm apodal mutant. Sci. Rep. 6, 18956; doi: 10.1038/srep18956 (2016).
References
Heffer, A. & Pick, L. Conservation and variation in Hox genes: how insect models pioneered the evo-devo field. Annu Rev Entomol 58, 161–179 (2013).
Williams, J. A. & Carroll, S. B. The origin, patterning and evolution of insect appendages. Bioessays 15, 567–577 (1993).
Angelini, D. R. & Kaufman, T. C. Insect appendages and comparative ontogenetics. Dev Biol 286, 57–77 (2005).
Erwin, D. H. & Davidson, E. H. The evolution of hierarchical gene regulatory networks. Nat Rev Genet 10, 141–148 (2009).
Gehring, W. J. The animal body plan, the prototypic body segment and eye evolution. Evol Dev 14, 34–46 (2012).
Green, J. & Akam, M. Evolution of the pair rule gene network: Insights from a centipede. Dev Biol 382, 235–245 (2013).
Gellon, G. & McGinnis, W. Shaping animal body plans in development and evolution by modulation of Hox expression patterns. Bioessays 20, 116–125 (1998).
Ronshaugen, M., McGinnis, N. & McGinnis, W. Hox protein mutation and macroevolution of the insect body plan. Nature 415, 914–917 (2002).
Pearson, J. C., Lemons, D. & McGinnis, W. Modulating Hox gene functions during animal body patterning. Nat Rev Genet 6, 893–904 (2005).
Hughes, C. L. & Kaufman, T. C. Hox genes and the evolution of the arthropod body plan. Evol Dev 4, 459–499 (2002).
Hart, M. C., Wang, L. & Coulter, D. E. Comparison of the structure and expression of odd-skipped and two related genes that encode a new family of zinc finger proteins in Drosophila. Genetics 144, 171–182 (1996).
Wang, L. & Coulter, D. E. bowel, an odd-skipped homolog, functions in the terminal pathway during Drosophila embryogenesis. EMBO J 15, 3182–3196 (1996).
Green, R. B., Hatini, V., Johansen, K. A., Liu, X. J. & Lengyel, J. A. Drumstick is a zinc finger protein that antagonizes Lines to control patterning and morphogenesis of the Drosophila hindgut. Development 129, 3645–3656 (2002).
Nüsslein-Volhard, C. & Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801 (1980).
Coulter, D. E. et al. Molecular analysis of odd-skipped, a zinc finger encoding segmentation gene with a novel pair-rule expression pattern. EMBO J 9, 3795–3804 (1990).
Ward, E. J. & Coulter, D. E. odd-skipped is expressed in multiple tissues during Drosophila embryogenesis. Mech Dev 96, 233–236 (2000).
Hao, I., Green, R. B., Dunaevsky, O., Lengyel, J. A. & Rauskolb, C. The odd-skipped family of zinc finger genes promotes Drosophila leg segmentation. Dev Biol 263, 282–295 (2003).
Gao, H., Wu, X. & Fossett, N. Odd-skipped maintains prohemocyte potency and blocks blood cell development in Drosophila. Genesis 49, 105–116 (2011).
Iwaki, D. D., Johansen, K. A., Singer, J. B. & Lengyel, J. A. drumstick, bowl and lines are required for patterning and cell rearrangement in the Drosophila embryonic hindgut. Dev Biol 240, 611–626 (2001).
Del Signore, S. J., Hayashi, T. & Hatini, V. odd-skipped genes and lines organize the notum anterior-posterior axis using autonomous and non-autonomous mechanisms. Mech Dev 129, 147–161 (2012).
Angelini, D. R., Kikuchi, M. & Jockusch, E. L. Genetic patterning in the adult capitate antenna of the beetle Tribolium castaneum. Dev Biol 327, 240–251 (2009).
Angelini, D. R., Smith, F. W. & Jockusch, E. L. Extent with modification: leg patterning in the beetle Tribolium castaneum and the evolution of serial homologs. G3-Genes Genom Genet 2, 235–248 (2012).
Ibrahim, D. M., Biehs, B., Kornberg, T. B. & Klebes, A. Microarray comparison of anterior and posterior Drosophila wing imaginal disc cells identifies novel wing genes. G3-Genes Genom Genet 3, 1353–1362 (2013).
Linz, D. M. & Tomoyasu, Y. RNAi screening of developmental toolkit genes: a search for novel wing genes in the red flour beetle, Tribolium castaneum. Dev Genes Evol 225, 11–22 (2015).
Banno, Y., Fujii, H., Kawaguchi, Y., Yamamoto, K. & Nishikawa, K. A Guide to the Silkworm Mutants: 2005. Gene Name and Gene Symbol. Kyusyu University, Fukuoka (2005).
Ueno, K., Hui, C. C., Fukuta, M. & Suzuki, Y. Molecular analysis of the deletion mutants in the E homeotic complex of the silkworm Bombyx mori. Development 114, 555–563 (1992).
Meng, Y. et al. The silkworm mutant lemon (lemon lethal) is a potential insect model for human sepiapterin reductase deficiency. J Biol Chem 284, 11698–11705 (2009).
Zhan, S. et al. An integrated genetic linkage map for silkworms with three parental combinations and its application to the mapping of single genes and QTL. BMC Genomics 10, 389 (2009).
Decoville, M., Giacomello, E., Leng, M. & Locker, D. DSP1, an HMG-like protein, is involved in the regulation of homeotic genes. Genetics 157, 237–244 (2001).
dos Santos, G. et al. FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res 43, D690–D697 (2015).
Kim, H. S. et al. BeetleBase in 2010: revisions to provide comprehensive genomic information for Tribolium castaneum. Nucleic Acids Res 38, D437–D442 (2010).
Heliconius Genome Consortium. Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487, 94–98 (2012).
Choo, S. W. & Russell, S. Genomic approaches to understanding Hox gene function. Adv Genet 76, 55–91 (2011).
Gibson, G. & Gehring, W. J. Head and thoracic transformations caused by ectopic expression of Antennapedia during Drosophila development. Development 102, 657–675 (1988).
DeFalco, T., Le Bras, S. & Van Doren, M. Abdominal-B is essential for proper sexually dimorphic development of the Drosophila gonad. Mech Dev 121, 1323–1333 (2004).
Hersh, B. M. et al. The UBX-regulated network in the haltere imaginal disc of D. melanogaster. Dev Biol 302, 717–727 (2007).
Chen, P. et al. Antennapedia is involved in the development of thoracic legs and segmentation in the silkworm, Bombyx mori. Heredity 111, 182–188 (2013).
Alphey, L. Re-engineering the sterile insect technique. Insect Biochem Mol Biol 32, 1243–1247 (2002).
Robinson, A. S. Mutations and their use in insect control. Mutat Res 511, 113–132 (2002).
Pan, M. H. et al. Establishment and characterization of an ovarian cell line of the silkworm, Bombyx mori. Tissue Cell 42, 42–46 (2010).
Duan, J. et al. SilkDB v2.0: a platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res 38, D453–D456 (2010).
Lander, E. S. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 (1987).
Kosambi, D. D. The estimation of map distances from recombination values. Ann Eugen 12, 172–175 (1944).
Giraldo-Calderón, G. I. et al. VectorBase: an updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res 43, D707–713 (2015).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30, 2725–2729 (2013).
Liang, J., Zhang, L., Xiang, Z. & He, N. Expression profile of cuticular genes of silkworm, Bombyx mori. BMC Genomics 11, 173 (2010).
Chen, P. et al. Fine mapping of a supernumerary proleg mutant (ECs-l) and comparative expression analysis of the abdominal-A gene in silkworm, Bombyx mori. Insect Mol Biol 22, 497–504 (2013).
McClure, R. F., Heppelmann, C. J. & Paya, C. V. Constitutive Fas ligand gene transcription in Sertoli cells is regulated by Sp1. J Biol Chem 274, 7756–7762 (1999).
Solomon, M. J., Strauss, F. & Varshavsky, A. A mammalian high mobility group protein recognizes any stretch of six AT base pairs in duplex DNA. Proc Natl Acad Sci USA 83, 1276–1280 (1986).
Acknowledgements
This work was funded by Hi-Tech Research and Development 863 Program of China Grant (No. 2013AA102507), National Basic Research 973 Program of China Grant (No. 2012CB114600), National Natural Science Foundation of China (No. 31472153, No. 31372379), Fundamental Research Funds for the Central Universities in China (No. XDJK2015C115, No. XDJK2013A001) and Chongqing Youth Science and Technology Talent Training Project (cstc2014kjrc-qnrc80001).
Author information
Authors and Affiliations
Contributions
P.C., X.L.T., F.Y.D. and C.L. conceived and designed the experiments, P.C. and M.Y.F. performed the experiments, P.C., X.L.T., M.Y.F. and S.Z.H. analyzed the data, H.H., J.B.S. and T.T.G. contributed materials and analysis tools, P.C. and X.L.T. wrote the manuscript, F.Y.D. and C.L. edited and proofread the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Chen, P., Tong, XL., Fu, MY. et al. Molecular mapping and characterization of the silkworm apodal mutant. Sci Rep 6, 18956 (2016). https://doi.org/10.1038/srep18956
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18956
- Springer Nature Limited