Abstract
Human rhinovirus species C (HRV-C) was recently discovered and this virus has been associated with various acute respiratory illnesses (ARI). However, the molecular evolution of the major antigens of this virus, including VP1, VP2 and VP3, is unknown. Thus, we performed complete VP1, VP2, and VP3 gene analyses of 139 clinical HRV-C strains using RT-PCR with newly designed primer sets and next-generation sequencing. We assessed the time-scale evolution and evolutionary rate of these genes using the Bayesian Markov chain Monte Carlo method. In addition, we calculated the pairwise distance and confirmed the positive/negative selection sites in these genes. The phylogenetic trees showed that the HRV-C strains analyzed using these genes could be dated back approximately 400 to 900 years and these strains exhibited high evolutionary rates (1.35 to 3.74 × 10−3 substitutions/site/year). Many genotypes (>40) were confirmed in the phylogenetic trees. Furthermore, no positively selected site was found in the VP1, VP2 and VP3 protein. Molecular modeling analysis combined with variation analysis suggested that the exterior surfaces of the VP1, VP2 and VP3 proteins are rich in loops and are highly variable. These results suggested that HRV-C may have an old history and unique antigenicity as an agent of various ARI.
Similar content being viewed by others
Introduction
Human rhinovirus (HRV) belongs to the genus Enterovirus and the family Picornaviridae. HRV is the common causative agent of acute respiratory illness (ARI), such as the common cold, bronchiolitis and pneumonia1. HRV is classified into three species: HRV-A, -B and -C1. HRV-C was recently discovered and this virus is not easily cultured by conventional methods2. Additionally, molecular epidemiological studies have suggested that HRV-C may have wide genetic diversity3. However, their virology is not exactly understood.
In general, enteroviruses, including HRV, possess the structural viral proteins (VP) 1 to 4. These are role-sharing proteins and of these proteins, VP1, VP2 and VP3 are both major structural proteins and essential antigens1, whereas VP4 is a scaffold protein of the viral capsid4. Additionally, the divergence of the VP1, VP2 and VP3 proteins is linked to the antigenicity of the various enteroviruses, resulting in different serotypes5. However, the essential knowledge of antigenicity regarding HRV-C is poorly understood.
Viral molecular evolution is different for each virus and it may be responsible for their genome size and genome polarity6. The positive pressure of the host may lead to changes in the selection site, leading to changes in antigenicity faster than if spontaneous mutations had occurred7. Therefore, such factors should be considered when we refer to the evolution of viral genes7.
Recent phylogenetic technologies, including the Bayesian Markov chain Monte Carlo (MCMC) method, enable us to perform cluster analysis and time scale evolution of the various viral genes8. For example, Tsukagoshi et al. showed that the evolution rate of the C-terminal 3rd hypervariable region in the glycoprotein gene, which is a major antigen of respiratory syncytial virus (RSV), was rapid, as was the hemagglutinin (H) gene in the influenza virus subtype AH19. Furthermore, Mizuta et al. showed that the evolutionary rate of the hemagglutinin-neuraminidase (HN) gene in human parainfluenza virus type 1 was nearly equal to the H gene of the measles virus10,11. However, the molecular evolution of the major structural proteins, such as the VP1, VP2 and VP3 proteins, in HRV-C is still unknown due to the wide genetic divergence of the VP protein coding regions in HRV-C; this divergence makes it hard to analyze their genetic properties. We were able to overcome that difficulty by analyzing the major antigen coding regions of HRV-C using next-generation sequencing (NGS) with thoroughly prepared primers. Additionally, we calculated the structure of the VP1, VP2 and VP3 proteins in silico. Using these advanced methods, we studied the molecular evolution of the VP1, VP2, and VP3 genes in HRV-C (139 strains) detected in various ARI patients.
Results
HRV-C specific RT-PCR and amplicon sequencing using NGS
To amplify the highly variable HRV-C genomic RNA by RT-PCR, suitable degenerate primers were designed using the following procedures. Publicly available HRV-C complete genome sequences (23 strains) were aligned by MUSCLE in the MEGA 5.0 software and a potential primer site was determined for a possible common reverse primer to lower variation. The availability of the degenerate primer HRV-C_6410R was evaluated with 8 randomly selected clinical specimens; a 5.8-kb RT-PCR product was observed for all 8 tested samples, suggesting that the primer could efficiently amplify the long range of the HRV-C genome (Suppl. Fig. S1). The amplicons theoretically contain an incomplete VP4 coding region, complete VP2 to 3C regions and an incomplete 3D region. To investigate the molecular evolution of the major antigens, we focused on the analyses of the VP1, VP2 and VP3 genes.
Using the above-described a primer set for PCR amplification, 187 amplicons were obtained and the amplicons were de novo sequenced using NGS with multiple indexing (~100,000 reads for each sample ID); these sequences were then assembled into the respective 5.8 kb HRV-C contig sequences. Homology searches of these contigs suggested that 139 of the 187 amplicons (74.3%) derived from clinical specimens were able to analyze as HRV-C strains. To characterize the genetic evolution and immunogenicity of HRV-C, the potential coding regions of the VP1, VP2 and VP3 genes in the obtained contigs were aligned with the reference HRV-C sequences.
Phylogenetic analysis of the VP1 gene in HRV-C
We analyzed the full sequence of the VP1 coding region (804–846 nt) in 139 strains of HRV-C detected in ARI patients. Using the Bayesian MCMC method, we constructed a phylogenetic tree with time scale evolution (Fig. 1). First, the present strains comprised three major lineages. When we genotyped the samples using 13% nucleotide divergence3,12, 44 genotypes were confirmed. Lineages 1, 2 and 3 contained 16, 16 and 12 genotypes, respectively. The most recent common ancestor (tMRCA) of all of the strains was found in 1652 (95% highest posterior density (HPD) 1522 to 1769). The lineages 1, 2 and 3 were dated back to 1744 (95% HPD 1640 to 1832), 1745 (95% HPD 1642 to 1832) and 1710 (95% HPD 1594 to 1812), respectively. The evolution rate was 3.48 × 10−3 substitutions/site/year (95% HPD 2.36 × 10−3 to 4.60 × 10−3). The results suggested that the VP1 gene in HRV-C showed a wide range of genetic divergence with a high rate of evolution.
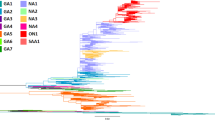
Phylogenetic tree of the VP1 gene in HRV-C constructed using the Bayesian MCMC method.
A phylogenetic tree based on VP1 gene sequences. In the tree, the triangle size expresses the number of strains, while only one representative strain is listed for each genotype. The strains are presented as follows: virus species/GenBank accession no./year. The GenBank accession numbers of the reference strains are indicated in bold letters. The scale bar represents the unit of time (year).
Phylogenetic analysis of the VP2 gene in HRV-C
As well as VP1 gene, we analyzed the full sequence of the VP2 coding region (783–816 nt) in 139 strains and a phylogenetic tree was constructed with time scale evolution (Fig. 1B). First, the present strains comprised four major lineages, similar to the phylogenetic tree based on the full VP1 coding region (Fig. 2). When we genotyped the strains using 13% nucleotide divergence3,12, 43 genotypes were confirmed. Lineages 1, 2, 3 and 4 contained 16, 16, 6 and 5 genotypes, respectively. tMRCA of all of the strains was found in 1125 (95% HPD 570 to 1548). Lineages 1, 2, 3 and 4 were dated back to 1294 (95% HPD 853 to 1643), 1361 (95% HPD 955 to 1681), 1545 (95% HPD 1244 to 1775) and 1586 (95% HPD 1312 to 1803), respectively. The evolution rate was 1.35 × 10−3 substitutions/site/year (95% HPD 2.12 × 10−3 to 6.46 × 10−4). The results suggested that HRV-C VP2 gene of the present strains had a wide range of genetic divergence with a high rate of evolution as well as their VP1 gene.
Phylogenetic tree of the VP2 gene in HRV-C constructed using the Bayesian MCMC method.
A phylogenetic tree based on VP2 gene sequences. In the tree, the triangle size expresses the number of strains, while only one representative strain is listed for each genotype. The strains are presented as follows: virus species/GenBank accession no./year. The GenBank accession numbers of the reference strains are indicated in bold letters. The scale bar represents the unit of time (year).
Phylogenetic analysis of the VP3 gene in HRV-C
We analyzed the full sequence of the VP3 coding region (699–717 nt) in 139 strains of HRV-C. We constructed a phylogenetic tree with time scale evolution as well as VP1 and VP2 genes (Fig. 3). First, the present strains comprised three major lineages, similar to the phylogenetic tree based on the full VP1 coding region (Fig. 3). When we genotyped the samples using 13% nucleotide divergence3,12, 42 genotypes were confirmed. Lineages 1, 2 and 3 contained 16, 15 and 11 genotypes, respectively. tMRCA of all of the strains was found in 1628 (95% HPD 1494 to 1746). Lineages 1, 2 and 3 were dated back to 1698 (95% HPD 1579 to 1797), 1749 (95% HPD 1654 to 1829) and 1695 (95% HPD 1581 to 1796), respectively. The evolution rate was 3.74 × 10−3 substitutions/site/year (95% HPD 2.63 × 10−3 to 4.91 × 10−3). The results suggested that the VP3 gene in HRV-C showed a wide range of genetic divergence with a high rate of evolution as well as the VP1 and VP2 genes.
Phylogenetic tree of the VP3 gene in HRV-C constructed using the Bayesian MCMC method.
A phylogenetic tree based on VP3 gene sequences. In the tree, the triangle size expresses the number of strains, while only one representative strain is listed for each genotype. The strains are presented as follows: virus species/GenBank accession no./year. The GenBank accession numbers of the reference strains are indicated in bold letters. The scale bar represents the unit of time (year).
Genetic divergence of the VP1 gene in HRV-C
We calculated the nucleotide identity of the VP1 gene in 139 strains of HRV-C. Among the present strains, the nucleotide identity was 60.2 to 100% and the deduced amino acid identity was 58.5 to 100%. Among the present strains in lineages 1, 2 and 3, the nucleotide identity was 65.4–100%, 63.0–100% and 61.2–100%, respectively, while among the present strains, the deduced amino acid identity was 68.5–100%, 67.0–100% and 59.0–100%, respectively. Furthermore, we calculated the p-distance of the present strains (Fig. 4 (a)). As a result, the p-distance of all strains was 0.34 ± 0.07 (mean ± standard deviation, SD) at the nucleotide level and 0.32 ± 0.08 at the deduced amino acid level. Among the present strains in lineages 1, 2 and 3, the values were 0.29 ± 0.07, 0.28 ± 0.10 and 0.29 ± 0.11 at the nucleotide level, respectively (Fig. 4 (b–d)) and the deduced amino acid p-distance values were 0.25 ± 0.08, 0.23 ± 0.09 and 0.27 ± 0.12, respectively.
Genetic divergence of the VP2 gene in HRV-C
We calculated the nucleotide identity of the VP2 gene in 139 strains of HRV-C. The nucleotide identity was 66.2 to 100% and the deduced amino acid identity was 70.7 to 100%. Lineages 1, 2, 3 and 4, the nucleotide identity was 67.0–100%, 69.3–100%, 72.2–100% and 73.8–100%, respectively, while the deduced amino acid identity was 71.3–100%, 75.7–100%, 82.3–100% and 82.6–100%, respectively. We also calculated the p-distance of the present strains (Fig. 5 (a)). The p-distance of all strains was 0.29 ± 0.06 (mean ± standard deviation, SD) at the nucleotide level and 0.23 ± 0.06 at the deduced amino acid level. Lineages 1, 2, 3 and 4, the values were 0.25 ± 0.09, 0.25 ± 0.07, 0.20 ± 0.09 and 0.17 ± 0.09 at the nucleotide level, respectively (Fig. 5 (b–e)) and the deduced amino acid p-distance values were 0.20 ± 0.08, 0.18 ± 0.06, 0.10 ± 0.06 and 0.09 ± 0.07, respectively.
Genetic divergence of the VP3 gene in HRV-C
We also calculated the nucleotide identity of the VP3 gene in 139 strains of HRV-C. Among the present strains, the nucleotide identity was 59.4 to 100% and the deduced amino acid identity was 58.3 to 100%. Lineages 1, 2 and 3, the nucleotide identity was 62.2–100%, 66.3–100% and 61.9–100%, respectively, while the deduced amino acid identity was 65.4–100%, 70.7–100% and 63.2–100%, respectively. Next, we calculated the p-distance of the present strains (Fig. 6 (a)). The p-distance of all strains was 0.33 ± 0.07 (mean ± SD) at the nucleotide level and 0.30 ± 0.08 at the deduced amino acid level. Among the present strains in lineages 1, 2 and 3, the values were 0.28 ± 0.10, 0.27 ± 0.07 and 0.28 ± 0.10 at the nucleotide level, respectively (Fig. 6 (b–d)) and 0.23 ± 0.10, 0.21 ± 0.08 and 0.24 ± 0.11, respectively, at the deduced amino acid level.
Simplot data and recombination analysis of the VP1, VP2 and VP3 genes in the present strains
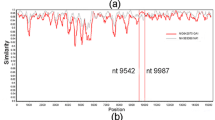
We calculated the similarity of the nucleotide sequences of the VP1, VP2 and VP3 genes in the present strains and in a prototype strain (HRV-QPM strain). As shown in Fig. 7 (a), an overall high similarity of the VP1 coding region was found when compared to the VP3 coding region. The minimum similarities of the VP2, VP3 and VP1 genes were approximately 70, 68 and 72%, respectively. Additionally, the similarities of the 5′-terminal VP3 coding region and the 3′-terminal VP1 coding region were low (approximately 70%), whereas the similarities of the 3′-terminal VP3 coding region and the 5′-terminal VP1 coding region were high (approximately 80%). Additionally, we calculated the similarity of the deduced amino acid sequences of the VP1 and VP3 genes in the present strains and the prototype strain (HRV-QPM strain) (Fig. 7 (b)). The minimum similarities of the VP2, VP3, and VP1 proteins were approximately 86, 83 and 87%, respectively. The Recombination Detection Program (RDP) found no evidence of a recombination event. The results suggested that high genetic divergence was found in the VP3 coding region compared to the VP2 and VP1 coding region.
Characterization of the VP1, VP2 and VP3 genes in HRV-C.
(a) Nucleotide similarity to HRV-QPM (prototype strain) was determined using SimPlot analysis. (b) Amino acid similarity to HRV-QPM (prototype strain) was determined using SimPlot analysis. The plots indicate the percentage of similarity to a 50% consensus sequence from each species' polyprotein compared to HRV-QPM.
Association between positive selection sites and the possible structures of the VP1, VP2 and VP3 proteins
Using SLAC, FEL and IFEL methods, we estimated the positive selection sites in the VP1, VP2 and VP3 proteins in HRV-C. No positively selected sites were detected in any position by any method, while many sites under negative selection (>100) were found. Next, we constructed a molecular model of the complex containing the HRV-C VP1, VP2 and VP3 proteins. The model showed that these proteins are rich in loop structures that are primarily positioned on the exterior surface of the capsid complex (Figs. 8 (a) and 8 (b)). Our Shannon entropy data show that these exterior loops of VP1, VP2 and VP3 are highly variable compared to the interior of the capsid within the HRV-C population analyzed in this study (Figs. 8 (a) and 8 (b)).
Shannon entropy scores expressed on the structural models of HRV-C VP1, VP2 and VP3 proteins.
The HRV-C VP1, VP2 and VP3 models were constructed by homology modeling as described in Materials and Methods and were superimposed on the VP1, VP2 and VP3 in those of the HRV2 capsid (PDB code: 3VDD). The entropy scores are expressed on the HRV-C VP1, VP2 and VP3 models. (a) Top view. (b) Side view.
Discussion
We studied the molecular evolution of the full length VP1, VP2 and VP3 genes in HRV-C detected from ARI. Analysis of the full sequences of the major 3 viral protein genes in the current HRV-C strains was performed using NGS with an improved RT-PCR method. Time-scaled phylogenetic analysis with evolution rate for the genes was analyzed using the Bayesian MCMC method. Additionally, we constructed the VP1, VP2 and VP3 proteins using an in silico method. First, the phylogenetic trees based on the VP1, VP2 and VP3 genes showed that the current HRV-C strains were classified into 3 or 4 major lineages and these lineages were subdivided into many genotypes (>40). The most recent common ancestor (tMRCA) of all the strains based on the VP1, VP2 and VP3 genes was found in the years 1652, 1125 and 1628, respectively. The evolution rates of both genes were fast. Similarity plot data showed high genetic divergence of the 5′-terminal VP3 coding region. Moreover, no positively selected site was found in the VP1, VP2 and VP3 proteins. Additionally, no recombination of the VP1, VP2 and VP3 genes was found in the studied strains. The exterior surfaces of the VP1, VP2 and VP3 proteins are rich in loops and are highly variable within the HRV-C population. The results suggested that the VP1, VP2 and VP3 genes, which encode major structural proteins of HRV-C, uniquely and rapidly evolved without positive selections.
Comprehensive molecular evolutionary and/or molecular epidemiologic studies of HRV-ABCs have been reported13,14. However, almost all studies were partially analyzed with regard to the VP4/VP2 coding region of HRV14,15. The genes coding the VP proteins have many hypervariable regions; thus, it is difficult to design common primers for the amplification of the VP1, VP2 and VP3 genes12,16. In addition, it may not be possible to isolate HRV-C using conventional methods at this time2. Thus, the antigenicity of HRV-C is still unknown. In the present study, we used an improved RT-PCR method with new primer sets and NGS and we detected and analyzed the full length VP1, VP2 and VP3 genes with a high probability (>70%). To the best of our knowledge, this may be the first large study of the complete VP1, VP2 and VP3 genes using many clinical strains.
Although HRV-C was recently discovered, it is thought that it may have a long history as a species. Indeed, Kiyota et al. showed that Japanese HRV-C strains could be dated back to the 1870s, according to the analysis of the VP4/VP2 coding region14. The present strains dated back approximately 400 to 900 years (Fig. 1,2,3). Previous reports have suggested that HRV-C and HRV-A are frequently detected in patients with various ARIs17. Additionally, with regards to the analysis of the VP4/VP2 coding region, both viruses exhibited large genetic divergence with many genotypes15. For example, Arakawa et al. showed that the VP4/VP2 coding region in HRV-ABCs had over 0.3 divergence17. In the present study, greater than 0.3 divergence was found in the VP1, VP2 and VP3 genes. High genetic divergence in the 5′-terminal VP3 coding region and the 3′-VP1 coding region was found. The results from the partial analysis were compatible with our findings12,18. These results suggested that HRV-C might have a long history dating back at least 100 years; however, further studies are needed to confirm this.
The VP proteins of picornaviruses, which include HRV, play roles in their biology7. VP1, VP2 and VP3 protein are located at the surface of the viral capsid and are exposed to immune pressure, whereas VP4 is located inside the capsid19. For example, the VP1, VP2 and VP3 protein of many types of enteroviruses, such as EV71, is essential for the virus's ability to infect the host cells and acts as a protective protein in the viral shell20. Additionally, these proteins are recognized as major antigens in the host20. Indeed, the VP1 protein of many EVs, including HRV-A, is a major antigen1. However, the VP1, VP2 and VP3 proteins are major antigens for some types of HEVs5. It has been suggested that positive pressure in the host is associated with positive selection sites in major antigens7. Positive selection shows a survival advantage under the selective constraints that confront the viral population7. In the studied HRV-C strains, positive selection site was not found in the VP1, VP2 and VP3 proteins. Thus, HRV-C may be hardly affected under positive selection in our immune system.
Next, many negative selection sites (>100) were found in VP proteins in the present HRV-C strains. In general, negative selection plays an important role in maintaining the long-term stability of biological structures by removing deleterious mutations7. In the present study, many negative selection sites (>200) were found in both genes. Kiyota et al. showed that over 100 sites were found in the VP4/VP2 coding region in HRV-C14. Thus, the negative selection sites in the VP1, VP2 and VP3 proteins may play the same roles as those in the VP4/VP2 coding regions21,22.
Our in silico structural analysis disclosed that the exterior surfaces of the VP1, VP2 and VP3 proteins are rich in loops, highly variable within the HRV-C population. It is conceivable that the exterior loops contain neutralization epitopes of HRV-C. In contrast, the interior regions of the VP1, VP2 and VP3 proteins were less diverse, suggesting the presence of functional and/or structural constraints on the diversity of this region. Some sites within these regions may be important for interactions with the infection receptor or the formation of a functional capsid complex structure. However, further studies may be needed to determine whether the VP1 protein is the major antigen in the infective HRV-C strains.
In conclusion, HRV-C was detected in various ARI patients and the virus exhibited large genetic divergence with a uniquely rapid evolution. Additionally, these viruses have been agents of ARI for a lot longer than previously thought.
Methods
Samples and patients
Nasopharyngeal swabs were collected from 2,922 patients with ARI between November 2007 and March 2013. ARI patients were diagnosed mainly with upper respiratory infection (URI) or lower respiratory infection (LRI; bronchitis, bronchiolitis and pneumonia). The samples were obtained by the local health authorities of the Fukui prefecture, Kumamoto prefecture, Tochigi prefecture and Yokohama Medical Center for the surveillance of viral diseases in Japan. Informed consent was obtained from the patients or their guardian for the donation of the samples.
RNA extraction, RT-PCR and de novo sequencing by NGS
Viral RNA was extracted from 140 μL of supernatant using the QIAamp Viral RNA Mini Kit without carrier RNA (Qiagen, Valencia, CA). RT-PCR was performed using the PrimeScript® II High Fidelity One Step RT-PCR Kit (TaKaRa Bio, Otsu, Japan) and the primer pair HRV-C_546F: 5′- CTACTTTGGGTGTCCGTGTT -3′ and HRV-C_6410R: 5′- CCRTCATARTTDGTRTARTCAAA -3′. The PCR reactions are described in Suppl. Fig. S1. The NGS DNA library was prepared using a Nextera XT DNA sample prep kit (Illumina, San Diego, CA) with 96 indexing, followed by 200-mer paired-end de novo sequencing with MiSeq (Illumina). The obtained sequencing reads were assembled using the A5 assembler with the default parameters23.
Phylogenetic analysis and estimation of the evolutionary rate using the Bayesian Markov chain Monte Carlo method
We aligned the nucleotide sequences of the VP1, VP2 and VP3 genes (positions; 2302-3126; 825 bp for HRV-QPM strain, positions 814-1602; 789 bp for HRV-QPM strain, positions 1603-2301; 699 bp for HRV-QPM strain) using CLUSTAL W [http://www.ddbj.nig.ac.jp/index-j.html]. To estimate the evolutionary rate and the time-scaled phylogeny, we used the Bayesian MCMC method in the BEAST package version 1.8.024. The dataset was analyzed with a strict clock using the general time reversible with gamma-distributed rates across sites (GTR + I−) substitution model25,26 selected by the Kakusan4 program version 4.0 [http://www.fifthdimension.jp/products/kakusan/]27. The MCMC chain was run for 50 million steps to achieve convergence, with sampling every 1000 steps. Convergence was assessed by the effective sample size (ESS) after a 10% burn-in using the Tracer program version 1.6 [http://tree.bio.ed.ac.uk/software/tracer]. Only parameters with an ESS above 200 were accepted. Uncertainty in the estimates was indicated by the 95% highest posterior density (HPD) intervals. The maximum clade credibility tree was obtained using the Tree Annotator program version 1.8.0 and the first 10% of the trees were removed as burn-in. The phylogenetic tree was viewed in the FigTree program version 1.5 [http://tree.bio.ed.ac.uk/software/figtree/].
Recombination analyses
Similarity plots showing the relationships between the aligned nucleotide sequences were generated using SimPlot, version 3.1 [http://sray.med.som.jhmi.edu/RaySoft/]28. The level of nucleotide similarity in each sequence, with a window size of 200 nt and a step size of 20 nt, was calculated using the Kimura 2-parameter method and similarity plot analyses based on the deduced amino acid sequences of the VP2, VP3 and VP1 proteins were performed with a window size of 100 aa. In this analysis, 1 aa was calculated using the Kimura 2-parameter method. The sequences were applied to RDP 3 [http://darwin.uvigo.es/rdp/rdp.html] to predict the recombination events using RDP, GENECONV, BootScan, Maxchi, Chimaera, SiSscan and 3Seq29,30.
Selective pressure analysis and the calculation of pairwise distances
To obtain estimates of the positively and negatively selected sites among the present strains during each season, we calculated the synonymous (dS) and nonsynonymous (dN) rates at every codon in the alignment using Datamonkey [http://www.datamonkey.org/]31. We used the following three different methods: single likelihood ancestor counting (SLAC), fixed effects likelihood (FEL) and internal fixed effects likelihood (IFEL). The SLAC and FEL methods were used to detect sites under selection at the external branches of the phylogenetic tree, while the IFEL method investigated sites along the internal branches. The SLAC method is best for large alignments but appears to underrate the substitution rate. Positively (dN > dS) and negatively (dN < dS) selected sites were determined by a p-value of <0.05 (SLAC, FEL, IFEL). Additionally, to assess the frequency distribution, we calculated the p-distance for the present strains, as previously described10.
Molecular modeling of the HRV-C VP1, VP2 and VP3 proteins and analysis of amino acid diversity
Three-dimensional (3-D) models of the HRV-C VP1, VP2 and VP3 complex were constructed by homology modeling using ‘MOE-Align’ and ‘MOE-Homology’ in the Molecular Operating Environment (MOE) (Chemical Computing Group Inc., Quebec, Canada) as described for norovirus capsid protein modeling32,33. The X-ray crystal structures of HRV2 VP1 (PDB code: 3VDD), rhinovirus 14 capsid (PDB code:1R08) and human coxsackievirus VP3 (PDB code: 4GB3)34 were used as the modeling templates for HRV-C VP1, VP2 and VP3 proteins, respectively, because these templates exhibited high scores with low E-values. The HRV-C VP1, VP2 and VP3 models were superimposed on VP1, VP2 and VP3 in the complex containing VP1, VP2, VP3 and VP4 of HRV2 strain (PDB code: 3VDD). Amino acid diversity at individual sites in the HRV-C sequences obtained in this study was analyzed with Shannon entropy scores as previously described32.
Ethical approval
The study was approved by the National Institute of Infectious Disease Ethics Committee (No. 495) and the study was conducted in compliance with the principles of the Declaration of Helsinki.
Nucleotide sequence accession numbers
The sequences generated in this study have been assigned the GenBank accession numbers LC004772 to LC004910.
References
Pallansch, M. A. & Roos, R. Enteroviruses: Polioviruses, Coxsackieviruses, Echoviruses and New Enteroviruses. Knipe, D. M., Howley, P. M., Griffin, D. E., Martin, M. A., Lamb, R. A., Roizman, B., Straus, S. E. (ed.), Fields Virology 839–893 Lippincott Willams & Wilkins, 2007.
Lee, W. M. et al. A diverse group of previously unrecognized human rhinoviruses are common causes of respiratory illnesses in infants. PLoS One 2, e966 (2007).
Simmonds, P. et al. Proposals for the classification of human rhinovirus species C into genotypically assigned types. J Gen Virol. 91, 2409–2419 (2010).
Lewis, J. K., Bothner, B., Smith, T. J. & Siuzdak, G. Antiviral agent blocks breathing of the common cold virus. Proc Natl Acad Sci U S A. 95, 6774–6778 (1998).
Caro, V., Guillot, S., Delpeyroux, F. & Crainic, R. Molecular strategy for 'serotyping' of human enteroviruses. J Gen Virol. 82, 79–91 (2001).
Sanjuán, R., Nebot, M. R., Chirico, N., Mansky, L. M. & Belshaw, R. Viral mutation rates. J Virol. 84, 9733–9748 (2010).
Domingo, E. Virus evolution. Knipe, D. M., Howley, P. M., Griffin, D. E., Martin, M. A., Lamb, R. A., Roizman, B., Straus, S. E. (ed.), Fields Virology. 389–422 Lippincott Willams & Wilkins, 2007.
Thorne, J. L., Kishino, H. & Painter, I. S. Estimating the rate of evolution of the rate of molecular evolution. Mol. Biol. Evol. 15, 1647–1657 (1998).
Kishibuchi, I. et al. Molecular evolution of attachment glycoprotein (G) gene in human respiratory syncytial virus detected in Japan 2008–2011. Infect Genet Evol. 18, 168–173 (2013).
Mizuta, K. et al. Detailed genetic analysis of hemagglutinin-neuraminidase glycoprotein gene in human parainfluenza virus type 1 isolates from patients with acute respiratory infection between 2002 and 2009 in Yamagata prefecture, Japan. Virol J. 8, 533 (2011).
Saitoh, M. et al. Molecular evolution of hemagglutinin (H) gene in measles virus genotypes D3, D5, D9 and H1. PLoS One 7, e50660 (2012).
McIntyre, C. L., McWilliam Leitch, E. C., Savolainen-Kopra, C., Hovi, T. & Simmonds, P. Analysis of genetic diversity and sites of recombination in human rhinovirus species C. J Virol. 84, 10297–10310 (2010).
McIntyre, C. L., Savolainen-Kopra, C., Hovi, T. & Simmonds, P. Recombination in the evolution of human rhinovirus genomes. Arch Virol. 158, 1497–1515 (2013).
Kiyota, N. et al. Genetic analysis of the VP4/VP2 coding region in human rhinovirus species C in patients with acute respiratory infection in Japan. J Med Microbiol. 62, 610–617 (2013).
Franco, D. et al. High genetic diversity and predominance of Rhinovirus A and C from Panamanian hospitalized children under five years with respiratory infections. Virol J. 9, 257 (2012).
Tapparel, C. et al. Rhinovirus genome variation during chronic upper and lower respiratory tract infections. PLoS One 6, e21163 (2011).
Arakawa, M. et al. Molecular epidemiological study of human rhinovirus species A, B and C from patients with acute respiratory illnesses in Japan. J Med Microbiol. 61, 410–419 (2012).
Linsuwanon, P. et al. Complete coding sequence characterization and comparative analysis of the putative novel human rhinovirus (HRV) species C and B. Virol J. 8, 5 (2011).
Racaniello, V. R. Picornaviridae: The viruses and their replication. Knipe, D. M., Howley, P. M., Griffin, D. E.,Martin,M. A., Lamb, R. A., Roizman, B., Straus, S. E. (ed.), Fields Virology 795–838 Lippincott Willams & Wilkins, 2007.
Zhang, D., Lu, J. & Lu, J. Enterovirus 71 vaccine: close but still far. Int J Infect Dis. 14, e739–e743 (2010).
Donker, N. C. & Kirkwood, C. D. Selection and evolutionary analysis in the nonstructural protein NSP2 of rotavirus A. Infect Genet Evol. 12, 1355–1361 (2012).
Yoshida, A. et al. Molecular epidemiology of the attachment glycoprotein (G) gene in respiratory syncytial virus in children with acute respiratory infection in Japan in 2009/2010. J Med Microbiol. 61, 820–829 (2012).
Tritt, A., Eisen, J. A., Facciotti, M. T. & Darling, A. E. An integrated pipeline for de novo assembly of microbial genomes. PLoS One 7, e42304 (2012).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 7, 214 (2007).
Drummond, A. J., Ho, S. Y., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006).
Hasegawa, M., Kishino, H. & Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol. 22, 160–174 (1985).
Tanabe, A. S. Kakusan4 and Aminosan: two programs for comparing nonpartitioned, proportional and separate models for combined molecular phylogenetic analyses of multilocus sequence data. Mol Ecol Resour. 11, 914–921 (2011).
Zhang, Y. et al. Complete genome analysis of the C4 subgenotype strains of enterovirus 71: predominant recombination C4 viruses persistently circulating in China for 14 years. PLoS One 8, e56341 (2013).
McVean, G. A. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584 (2004).
Martin, D. P. et al. RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics 26, 2462–2463 (2010).
Pond, S. L. & Frost, S. D. Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics. 21, 2531–2533 (2005).
Motomura, K. et al. Identification of monomorphic and divergent haplotypes in the 2006–2007 norovirus GII/4 epidemic population by genomewide tracing of evolutionary history. J Virol. 82,11247–11262 (2008).
Motomura, K. et al. Divergent evolution of norovirus GII/4 by genome recombination from May 2006 to February 2009 in Japan. J Virol. 84, 8085–8097 (2010).
Yoder, J. D., Cifuente, J. O., Pan, J., Bergelson, J. M. & Hafenstein, S. The crystal structure of a coxsackievirus B3-RD variant and a refined 9-angstrom cryo-electron microscopy reconstruction of the virus complexed with decay-accelerating factor (DAF) provide a new footprint of DAF on the virus surface. J Virol. 86, 12571–12581 (2012).
Acknowledgements
This work was partly supported by a Grant-in-Aid from the Japan Society for the Promotion of Science and for Research on Emerging and Re-emerging Infectious Diseases from the Ministry of Health, Labour and Welfare, Japan (H25-Shinko-Ippan-015 and H22-Shinko-Ippan-011). We would like to thank Ms. Izumi Kushibuchi and Dr. Keiji Funatogawa (Tochigi Prefectural Institute of Public Health) and Ms. Eiko Hirano (Fukui Prefectural Institute of Public Health and Environmental Science) for helpful discussion.
Author information
Authors and Affiliations
Contributions
M.K., A.R., M.T., K.O., H.I. and H.K. designed research; S.N., T.S., H.T., M.Y. and M.N. performed research; H.S., N.K., K.S. and M.O. contributed analytic tools, S.N., T.S., H.T., H.S., N.K., K.K., K.S. and T.K. analyzed data; M.K., N.S., S.H. and H.K. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kuroda, M., Niwa, S., Sekizuka, T. et al. Molecular evolution of the VP1, VP2 and VP3 genes in human rhinovirus species C. Sci Rep 5, 8185 (2015). https://doi.org/10.1038/srep08185
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08185
- Springer Nature Limited
This article is cited by
-
Targeting intercellular adhesion molecule-1 (ICAM-1) to reduce rhinovirus-induced acute exacerbations in chronic respiratory diseases
Inflammopharmacology (2022)
-
Genome sequencing and phylogenetic reconstruction reveal a potential fourth rhinovirus species and its worldwide distribution
Archives of Virology (2021)