Abstract
The DRE/DREF transcriptional regulatory system has been demonstrated to activate a wide variety of genes with various functions. In Drosophila, the Hippo pathway is known to suppress cell proliferation by inducing apoptosis and cell cycle arrest through inactivation of Yorkie, a transcription co-activator. In the present study, we found that half dose reduction of the hippo (hpo) gene induces ectopic DNA synthesis in eye discs that is suppressed by overexpression of DREF. Half reduction of the hpo gene dose reduced apoptosis in DREF-overexpressing flies. Consistent with these observations, overexpression of DREF increased the levels of hpo and phosphorylated Yorkie in eye discs. Interestingly, the diap1-lacZ reporter was seen to be significantly decreased by overexpression of DREF. Luciferase reporter assays in cultured S2 cells revealed that one of two DREs identified in the hpo gene promoter region was responsible for promoter activity in S2 cells. Furthermore, endogenous hpo mRNA was reduced in DREF knockdown S2 cells and chromatin immnunoprecipitation assays with anti-DREF antibodies proved that DREF binds specifically to the hpo gene promoter region containing DREs in vivo. Together, these results indicate that the DRE/DREF pathway is required for transcriptional activation of the hpo gene to positively control Hippo pathways.
Similar content being viewed by others
Introduction
The BED finger-type transcription factor DREF (DNA replication-related element-binding factor)1 binds to the DRE sequence (5′-TATCGATA), a highly conserved sequence in the core promoters of many Drosophila genes2. Target genes of DREF are involved in a variety of diverse processes3, such as DNA replication1,4,5,6, cell cycle regulation7,8, apoptosis9, protein synthesis10 and degradation11 and maintenance of chromatin structure12,13.
Analyses of the cellular function of DREF suggest that it is a multifunctional protein. As a transcription factor DREF has been shown to be important for Drosophila development. It is important for bristle development through its regulation of endoreplication in shaft cells14. It co-operates with Drosophila myeloid leukemia factor15,16 and the chromatin regulator XNP/dATRX17 in thorax development to regulate the JNK pathway. It also functions downstream of the Target-of-Rapamycin (TOR) pathway18 to modulate cell and organ growth in Drosophila. Genome-wide ChIP-sequence analyses also suggest that as well as functioning as a transcription factor DREF, in collaboration with insulator proteins may contribute to maintaining chromosome organization during the cell cycle, marking a subset of genomic sites for the assembly of pre-replication complexes and gene bookmarking during the M/G1 transition19. Furthermore, a DREF/TRF2 complex20 appears to be important for maintenance of telomere length in Drosophila21.
We have reported that the DRE/DREF pathway is required for transcriptional regulation of the warts (wts) gene, an essential component of the Hippo pathway22. The Hippo pathway itself was firstly identified in Drosophila as a tumor-suppressive signal cascade which plays a crucial role in controlling organ size23,24,25,26,27,28.This pathway is defined by a kinase cascade whereby the Ste-20-like kinase Hippo (Hpo), facilitated by the WW-domain-containing adaptor protein Salvador (Sav), phosphorylates Wts25,27,28,29. Activated Wts then phosphorylates and inactivates the transcriptional co-activator Yorkie (Yki)30,31,32, leading to transcriptional down-regulation of target genes such as the cell-cycle regulator cyclin E, the cell death inhibitor diap1, the Hippo pathway regulator expanded (ex) and the microRNA bantam33,34,35,36,37,38,39,40,41,42,43. Inactivation of the Hpo, Sav, or Wts tumor suppressors, or overexpression of Yki, results in massive tissue over-growth, characterised by excessive cell proliferation and diminished apoptosis. A recent study showed that the Hippo pathway also regulates normal proliferation of intestinal stem cells in Drosophila midgut, playing an essential role in maintaining homeostasis and regeneration in response to tissue damage44. Although, a variety of factors have been identified which interact with the Hippo pathway, indicating wide-ranging functions, the mechanisms of, transcriptional regulation of the genes encoding these factors is largely unknown and poorly studied.
By extensive genetic screening with DREF overexpressing flies, we previously identified several genes related to the Hippo pathway13. However, although some preliminary indications were obtained regarding the wts gene22 the biological significance of these interactions and the involvement of DREF in regulation of the Hippo pathway are poorly understood. In the present study, we established that overexpression of DREF induced increased hpo signals in eye imaginal discs coupled with apoptosis. We also observed significant increase of phospho-Yki in DREF-overexpressing flies and a significant reduction of the diap1-lacZ reporter in DREF-overexpressing eye discs. In S2 cells knockdown of DREF reduced the level of endogenous hpo mRNA. These results suggest that DREF positively regulates the Hippo pathway to restrict apoptosis. Consistent with this, at least one of the two DREs identified in the hpo gene promoter region was found to be responsible for promoter activity determined by a luciferase transient expression assay in Drosophila S2 cells. In addition, chromatin immunoprecipitation assays with anti-DREF antibodies revealed that DREF binds specifically to the hpo gene promoter region containing DREs in vivo. These results indicate that the DRE/DREF pathway is required for transcriptional activation of the hpo gene to positively control Hippo pathways.
Results
Half dose reduction of the hpo gene induces ectopic DNA synthesis in eye discs and this can be suppressed by overexpression of DREF
A number of previous studies have implicated both Hippo and DREF in the control of cell proliferation and apoptosis and both factors have been suggested to affect the balance of cell proliferation and apoptosis via p53. In addition we have recently used the fact that overexpression of DREF in eye discs induces a severe rough eye phenotype in adults without impairing viability or fertility45, to identify proteins that interact with DREF genetically, by screening for mutations that modify the rough eye phenotype. This analysis revealed several modifier genes related to Hippo pathway, such as ex and fat13. We have also reported that DRE/DREF pathway may be required for transcriptional regulation of the wts gene22, an essential component of the Hippo pathway.
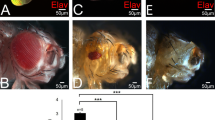
To further investigate the relationship between DREF and the Hippo pathway, we visualised DNA synthesis in the posterior region of the eye imaginal disc using an EdU incorporation assay and determined how this was affected by altered expression of DREF and Hippo. In eye discs of wild type flies, DNA synthesis was detected in the region anterior to the morphogenetic furrow (MF), where cells undergo non synchronous proliferation and also in the synchronized S phase zone behind the MF. In the eye discs of flies heterozygous for hpo mutation, ectopic DNA synthesis was detected in the region posterior to MF (Fig. 1E, G). Slight induction of DNA synthesis in the posterior region of the eye disc was observed in DREF-overexpressing flies, as reported previously45 (Fig. 1D, G). However, a similar extent of ectopic DNA synthesis was observed with GFP-overexpressing flies (Fig. 1C, G), suggesting that overexpression of DREF only marginally induced ectopic DNA synthesis. Overexpression of DREF and GFP was confirmed by immunostaining with specific antibodies (Fig. 1A, B). Immunostaining of eye discs with anti-DREF antibody further confirmed ubiquitous expression of endogenous DREF together with overexpression and knockdown of DREF by GMR-GAL4 driver (Supplementary Fig. S1 online). Interestingly, extensive induction of DNA synthesis in flies heterozygous for the hpo mutation was extensively suppressed by overexpression of DREF (Fig. 1F, G).
Half reduction of the hpo gene dose enhances cell proliferation which can be suppressed by over-expression of DREF.
(A) Eye imaginal discs of flies overexpressing UAS-GFP are stained with anti-GFP antibody (Red) and DAPI (Blue). (B) Eye discs expressing UAS-DREF are stained with anti-DREF antibody (Red) and DAPI (Blue). (For cell proliferation, eye imaginal discs were labeled with EdU (Red). (C) GMR-GAL4/+; UAS-GFP/+; +. (D) GMR-GAL4/+; UAS-DREF/+; +. (E) w*; +; hpoKC202/CyO. (F) GMR-GAL4/+; UAS-DREF/+; hpoKC202/+. (G) Quantification of the number of ectopic EdU cells in the posterior region of the eye discs. ***p<0.001, ****p<0.0001. White arrowheads indicate morphogenetic furrow (MF). The white border line indicates the posterior region (a) indicates anterior, (p) indicates posterior. Scale bars are 40 μm. The flies were reared at 28°C.
Half reduction of the hpo gene dose reduces the cell death program in DREF-overexpressing flies
We also examined the effects of alterations in these proteins on apoptosis using the Cell Event Caspase-3/7 Green Detection Reagent assay which uses a fluorogenic substrate to detect activated Caspases 3 and 7. This cell-permeant reagent consists of a four amino acid peptide (DEVD) conjugated to a nucleic acid binding dye. It is intrinsically non-fluorescent, because the DEVD peptide inhibits the ability of the dye to bind to DNA, however activation of caspase-3 or caspase-7 in apoptotic cells causes cleavage of the DEVD peptide, enabling the dye to bind to DNA and produce a bright, fluorogenic response. In eye imaginal discs of flies expressing either UAS-GFP (control) (Fig. 2A) or heterozygous for the hpo mutation (Fig. 2C), no apoptotic cells were detected. However, eye imaginal discs of DREF over-expression flies showed extensive cell death signals in the region posterior to the morphogenetic furrow (Fig. 2B) as compared to the control (Fig. 2A) and the hpo heterozygous mutant (Fig. 2C). Furthermore, these signals were dramatically reduced when heterozygous hpo mutation was combined with overexpression of DREF in the posterior region (Fig. 2D, E). These observations are consistent with other studies reporting that hpo mutation in Drosophila imaginal discs is associated with severe tumour-like phenotypes28 and resistance to apoptotic signals25,26,27,29,46,47,48.
Half dose reduction of the hpo gene reduces the cell death program in DREF over-expression flies.
Cell Event Caspase-3/7 Green Detection Reagent assays were performed in eye imaginal discs (Green). (A) GMR-GAL4/+; UAS-GFP/+; +. (B) GMR-GAL4/+; UAS-DREF/+; +. (C) w*; hpoKC202/CyO; +. (D) GMR-GAL4/+; UAS-DREF/+; hpoKC202/+. (E) Quantification of the number of apoptotic cells in the posterior region of the eye discs. ***p<0.001, ****p<0.0001. White arrowheads indicate the morphogenetic furrow (MF). The white border line indicates the posterior region. (a) indicates anterior and (p) indicates posterior. Scale bars are 40 μm. The flies were reared at 28°C.
Overexpression of DREF increases the hpo level in eye imaginal discs
The observations described above might be explained by activation of the Hippo pathway by overexpression of DREF. We therefore performed immunostaining of eye imaginal discs with anti-hpo antibodies. In eye discs of DREF overexpressing flies, hpo signals increased extensively in the posterior region to MF (Fig. 3B) as compared to control GFP-overexpressing flies (Fig. 3A). The hpo heterozygous mutant showed no detectable anti-hpo signal (Fig. 3C). With the combination of over-expression of DREF and hpo mutation, the anti-hpo signal was almost comparable to that in the controls (Fig. 3D, E). These results indicate that overexpression of DREF indeed increased the hpo level in eye discs.
Over-expression of DREF significantly enhances Hpo signals in eye imaginal discs.
Eye imaginal discs were immunostained with anti-hpo antibodies (Red). (A) GMR-GAL4/+; UAS-GFP/+; +. (B) GMR-GAL4/+; UAS-DREF/+; +. (C) w*; +; hpoKC202/CyO. (D) GMR-GAL4/+; UAS-DREF/+; hpoKC202/+. (E) Quantification of the number of Hpo positive signals in the posterior region of the eye discs. ***p<0.001, ****p<0.0001. White arrowheads indicate the morphogenetic furrow (MF). The white border line indicates the posterior region (a) indicates anterior, (p) indicates posterior. Scale bars are 40 μm. The flies were reared at 28°C.
To further confirm the effects of over-expression of DREF on the Hippo signal in eye discs, we performed flip-out experiments with DREF knockdown flies. The knockdown of DREF was confirmed by immunostaining of eye discs with the anti-DREF antibody in the flip-out experiments (Fig. 4A-C, 4A’-C’). The flip-out clones expressing GAL4 alone were used as a control (Fig. 4D–F). Immunostaining of eye imaginal discs of control flies over-expressing GFP with anti-hpo antibody revealed that Hippo is expressed as punctate pattern throughout eye discs (Fig. 5A, B). In the flip-out experiment, cells marked with GFP (Green) express DREF dsRNA (Fig. 5D, D’) and in these cells the hpo signal was significantly reduced (Fig. 5C, E). However, in the clone area without GFP, hpo signals (Red) were still detectable (Fig. 5C, E). These data further confirmed that DREF is required for expression of Hippo in the eye discs.
Confirmation of DREF knockdown in flip-out experiments.
(A) Eye imaginal discs of hs-flp; Act5C>FRT y FRT>GAL4, UAS-DREFIR flies are stained with anti-DREF antibody (Blue). (B) Cells expressing DREFdsRNA are marked with GFP (Green). (C) Merged image of anti-DREF and GFP signals in DREF knockdown eye discs. (A’), (B’) and (C’) are larger images of white box areas in (A), (B) and (C), respectively. In the control flies expressing GAL4 alone, the eye discs were stained with anti-DREF antibody (D) and anti-GFP antibody (E). (F) Merge image of anti-DREF and GFP signals in the control eye discs. The bars indicate 40 μm.
Expression of DREF dsRNA reduces hippo levels in eye discs.
(A) Eye imaginal discs of GMR>GFP flies are stained with anti-hpo antibody (Red). (B) Merge image of anti-hpo and GFP signals (Green) of the flies overexpressing GFP in the posterior region. (C) Eye discs of the flip-out experiment are stained with anti-hpo antibody (Red). (D) Cells expressing DREFdsRNA are marked with GFP (Green). (E) Merged image of anti-hpo and GFP signals in DREF knockdown eye discs. (C’), (D’) and (E’) are larger images of white box areas in (C), (D) and (E), respectively. Scale bars indicate 40 μm.
Overexpression of DREF increases the phospho-Yki level in eye imaginal discs
Yorkie (Yki) transcriptionally activates cycE and diap1, which are associated with cell proliferation and anti-apoptosis, respectively. Therefore, to investigate the effect of DREF downstream of the Hippo pathway, we performed immunostaining to determine how DREF affects the levels of anti-phospho-Yki, an inactive form of Yki, in eye discs. We found that overexpression of DREF increased phospho-Yki signals in the region posterior to the MF (Fig. 6B, E). The signals were reduced if DREF overexpression was combined with hpo mutation (Fig. 6D, E). In heterozygous hpo mutant flies, phospho-Yki signals were found to be lower than in control flies (Fig. 6C, E).
Over-expression of DREF increases the phospho-Yki level and reduces diap1-lacZ signals in eye imaginal discs.
Eye imaginal discs were immunostained with anti-phospho-Yki antibodies. (Red) (A) GMR-GAL4/+; UAS-GFP/+; +. (B) GMR-GAL4/+; UAS-DREF/+; +. (C) w*; +;hpoKC202/CyO. (D) GMR-GAL4/+; UAS-DREF/+; hpoKC202/+. (E) Quantification of the number of positive phospho-Yki signals in the posterior region of the eye discs. To examine expression of the diap1-lacZ reporter, eye discs were stained with anti-β-galactosidase antibody (Red) and DAPI (Blue). (F) GMR-GAL4/+; +; diap1-lacZ/+; (G) GMR-GAL4/+; UAS-DREF/+; diap1-lacZ/+. (H) Quantification of the number of β-galactosidase positive signals in the region posterior to the morphogenetic furrow. *p<0.05, ***p<0.001, ****p<0.0001. White arrowheads indicate the morphogenetic furrow (MF). The white border line indicates the posterior region. (a) indicates anterior and (p) indicates posterior. Scale bars are 40 μm. The flies were reared at 28°C.
To further investigate the effects of DREF on phospho-Yki level, the flip-out experiment with DREF knockdown flies was carried out. The phospho-Yki positive cells were detected mostly in the GFP-negative area where DREF was not knocked down (Fig. 7B-B’, E-E’, F-F’). In contrast, in the GFP-positive area, only a few phospho-Yki signals were detected (Fig. 7, white arrows). We found that these phospho-Yki-positive cells are also DREF-positive even in the GFP clone area (Fig. 7, white arrows), suggesting some variation in DREF-knockdown efficiency in individual cells within the same clone. It should be noted that these cells are still red even in merged image with GFP (Fig. 7E). The flip-out experiments were also carried out with the flies expressing GAL4 alone as a control to confirm these observations. The phospho-Yki signals were scattered throughout the whole eye discs including both GFP and non-GFP clones (Fig. 8B, C, D, F). Whereas, there was no change in the expression of DREF in the eye discs of these flies (Fig. 8A, D, E, F). In contrast to the data with the DREF-knockdown clone analysis, phospho-Yki signals become yellow in merged image with GFP in this control flip-out experiments (Fig. 8C). These data taken together suggest that DREF positively regulates the Hpo level and consequently increases the level of phospho-Yki in the eye disc.
Expression of DREF dsRNA reduces phospho-Yki levels in eye discs.
Eye imaginal discs of hs-flp; Act5C>FRT y FRT>GAL4, UAS-DREFIR flies are stained with anti-DREF antibody (Blue) (A) and with anti-phospho Yki antibody (Red) (C). (B) Merged image of DREF and phospho-Yki signals in DREF knockdown eye discs. (D) Cells expressing DREFdsRNA are marked with GFP (Green). (E) Merged image of phospho-Yki and GFP signals. (F) Merged image of DREF, phospho-Yki and GFP signals. (A’–F’) larger images of white box areas in (A–F). The white arrows indicate both DREF-positive and phospho Yki-positive cells in GFP clone area. The bars indicate 40 μm.
Distribution of endogenous phospho-Yki in the control flip-out experiments.
Eye imaginal discs of hs-flp; Act5C>FRT y FRT>GAL4, UAS-GFP flies are stained with anti-DREF antibody (Blue) (A) and with anti-phospho Yki antibody (Red) (B). (C) Merged image of phospho-Yki and GFP signals. (D) Merged image of DREF and phospho-Yki signals. (E) Merged image of DREF and GFP signals. (F) Merged image of DREF, phospho-Yki and GFP signals. The endogenous phospho-Yki signals scattered throughout the whole eye discs and there was no change in the level of DREF signals. The bars indicate 40 μm.
An increase in phospho-Yki should cause a decrease of the DIAP1 level. We therefore examined expression of DIAP1 by monitoring lacZ expression of the diap1-lacZ enhancer trap line, thj5c8 P[lacZ] that carries a P[lacZ] insertion in the 5′ untranslated region of diap1 (th)49. It is reported that the expression pattern of the lacZ mimics that of endogenous diap150. Expression of the diap1-lacZ reporter was significantly decreased in DREF-overexpressing flies (Fig. 6F, G, Supplementary Fig. S2 online). Quantification revealed that in eye discs overexpressing DREF, diap1-lacZ signals decreased by 73% of that of flies expressing GAL4 alone (Fig. 6H).
The 5′-flanking region of the hpo gene contains DRE and DRE-like sequences
The DRE sequence is an 8 bp sequence of which the central 6 bp (5′-ATCGAT) is essential for DREF binding and activation of promoters51 and previous studies have demonstrated that DRE sequences in the genomic region 5′ of the transcription initiation site can up-regulate transcription of many target genes.
To determine whether transcription of the hpo gene is directly regulated by DREF, we searched for DRE sequences within the 1.4 kb genome region 5′ of the transcription initiation site of the hpo gene as documented on the Drosophila genome database, Flybase. We found a perfect match to the DRE sequence, named DRE1 (5′-TATCGATA) located at nucleotide position from -136 to -129 with respect to the transcription initiation site and a DRE-like sequence with a 7 out of 8 bp match DRE2 (5′-TATCGATc) at -450 to -443 (Fig. 9A).
The hpo gene carries DRE and DRE-like sequences in the 5′-flanking region.
(A) Schematic of the 5′-flanking region of the hpo gene. The transcription initiation site is indicated by the arrow and designated as +1. The positions and nucleotide sequences of DRE1 and DRE2 are shown. Arrowheads indicate positions of the primers used for real-time PCR reactions. (B) Crosslinked chromatin of S2 cells was immunoprecipitated with either anti-DREF IgG or control rabbit IgG. Genomic regions containing hpoDRE1, hpoDRE2 and wtsDRE1, 2 were amplified by PCR and compared with amplicons from immunoprecipitates with control rabbit IgG.
DREF binds to the genomic region containing DRE and DRE-like sequences of the hpo gene in vivo
To test whether DREF binds to the DRE-containing 5′-flanking region of the hpo gene in S2 cells, chromatin immunoprecipitation assays with anti-DREF polyclonal antibodies were carried out (Fig. 9B). The wts locus, previously shown to bind DREF22, was used as a positive control and the upstream genomic devoid of DREs was used as a negative control. Amplification of the wts gene promoter region containing DRE in immunoprecipitates with anti-DREF IgG was 77-fold higher than that with control IgG (Fig. 9B). In contrast, no significant amplification of the upstream region was observed. Amplification of the hpo gene region containing DRE1 from immunoprecipitates with the anti-DREF IgG was about 9.9-fold higher than that with control IgG (Fig. 9B). We also could detect binding of DREF to the DRE2 region of the hpo promoter, but this binding was weaker than that for the DRE1 containing region. These results indicate that DREF mainly binds to the genomic region containing DRE1 and less effectively to DRE2 under cellular conditions.
DRE1 and DRE2 both influence hpo promoter activity
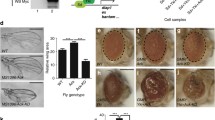
To investigate the role of the DREs in hpo promoter activity, we carried out transient luciferase expression assays in cultured Drosophila S2 cells. We constructed a hpo promoter: luciferase fusion plasmid and a set of derivatives carrying mutations in DRE1 and/or DRE2. These plasmids were transfected into S2 cells and luciferase activity was measured. A mutation in DRE1 (hpo-DRE1mut) and mutations in both DREs (hpo-DREallmut) reduced the hpo promoter activity by 85% and 70%, respectively (Fig. 10A). However, the mutation in DRE2 (hpo-DRE2mut) slightly increased the hpo promoter activity (Fig. 10A). These results indicate that DRE1 is mainly responsible for promoter activity and DRE2 may have some negative role in promoter activity.
DREF plays essential roles in hpo gene promoter activity in cultured S2 cells.
(A) Schematic features of the hpo promoter-luciferase fusion plasmid hpo-WT-luc and its base-substituted derivatives (hpo-DRE1mut-luc, hpo-DRE2mut-luc, hpo-DREallmut-luc). DRE is represented by an open box and mutated DRE by a closed box. Plasmids were transfected into S2 cells and luciferase activities measured at 48 hours thereafter. Luciferase activity was normalized to Renilla luciferease activity and expressed relative to that of hpo-WT-luc. Mean activities with standard deviations from three independent transfections are shown. (B) Effects of DREFdsRNA treatment on hpo gene promoter activity in S2 cells. Mean activities with standard deviations from three independent transfections are shown. (C) DREFdsRNA treatment reduces endogenous hpo mRNA levels in S2 cells. DREF mRNA, wts mRNA and hpo mRNA in DREF dsRNA-treated cells were measured by quantitative RT-PCR and compared with the values for non-dsRNA treated cells (Mock). mRNA for β-tubulin was used as a negative control and mRNA for wts as a positive control. * p<0.05.
DREF is required for hpo promoter activity in S2 cells
To further examine the requirement of DREF for hpo promoter activity, we carried out DREF RNA interference (RNAi) assays with cultured Drosophila S2 cells. We conducted transient luciferase expression assays with the wild-type hpo gene promoter-luciferase reporter gene, after treating S2 cells with DREFdsRNA, control YFPdsRNA or no dsRNA (Mock). Treatment of S2 cells with DREFdsRNA reduced hpo gene promoter activity by 50%, while control YFPdsRNA treatment exerted only a marginal effect (Fig. 10B). When DRE1 and 2 were mutated, no further decrease of hpo promoter activity was observed with knockdown of DREF (Fig. 10B). These results indicate that DREF is required for hpo promoter activity.
Knockdown of the DREF gene reduces endogenous hpo gene expression in cultured S2 cells
We also used the DREF knockdown S2 cells to examine whether DREF affects expression of the endogenous hpo gene. Total RNAs from dsRNA treated S2 cells were isolated at 5 days after addition of dsRNA for assessment by quantitative RT-PCR. The β-tubulin gene was used as a negative control and the wts gene as a positive control. The DREF mRNA level was reduced by 97% with DREFdsRNA treatment as compared with no dsRNA (mock) (Fig. 10C). The hpo mRNA level in DREFdsRNA treated cells was reduced to 58% relative to no dsRNA treated cells, while no significant change was observed with YFPdsRNA treated cells (Fig. 10C). Similar effects on hpo mRNA level were obtained for the positive control wts. These results indicate that DREF is truly required for hpo gene expression.
Discussion
In the present study, we demonstrated that the hpo gene is a major DREF-target, featuring one DRE and one DRE-like sequence in its 5′-flanking region. Luciferase transient expression assays showed hpo promoter activity to mainly depend on DRE1 and DREF levels. Consistent with this, chromatin immunoprecipitation assays showed DREF to bind to the genomic region of the hpo gene containing the DREs and the level of hpo mRNA was reduced in DREF knockdown S2 cells treated with DREFdsRNA. These data provide compelling evidence that DREF is a positive regulator of the hpo gene and its expression is directly activated by the DRE/DREF pathway. It has already been shown that DREF can simultaneously activate various apoptosis inducing genes for instance JNK pathway genes15 and p539. The increase of hpo level by overexpression of DREF resulted in increased levels of inactive phospho-Yki and a concomitant decrease of the DIAP1 level to further enhance apoptosis in eye discs (Fig. 11). In fact a half dose reduction of the hpo gene extensively decreased the apoptosis signals in eye discs of DREF-overexpressing flies.
A model of the role of DREF in regulation of Hippo, JNK and p53 pathways inducing apoptosis.
DREF binds to DRE in the hpo promoter region which activates hpo gene transcription. DREF upregulates both JNK and p53 pathways to induce apoptosis9,15. Activation of the Hippo pathway by DREF inactivates Yorkie to further enhance apoptosis.
Coordination of positive and negative regulation for cell proliferation is essential to achieve appropriate organ formation. During normal development and regeneration of organs after surgical excision, the organs reach their proper size by precisely controlling cell proliferation and its arrest. The Hippo pathway restricts cell proliferation via cell cycle arrest and inducing apoptosis. The major effect of DREF-overexpression in eye discs has been for a long time thought to be induction of ectopic DNA synthesis45. However, our analysis here, re-examining the influence of DREF-overexpression in eye discs, suggests that the major effect is the induction of apoptosis accompanied with induction of hpo expression and that the change in DNA synthesis appears to be only marginal. It should be noted that similarly strong induction of apoptosis was also observed when DREF was overexpressed in wing discs52. We therefore propose that the Hippo pathway is one of the major targets of the DRE/DREF regulatory system. In addition to Hippo pathway-related genes, DREF positively regulates the basket gene, a Drosophila JNK15. JNK exerts a protective function for the genome and promotes apoptosis just like p53, which is also a target of DRE/DREF9. In addition, the Hpo pathway and the JNK pathway cooperate in tissue growth and regeneration3,22. Thus DREF, in regulating both Hippo and JNK pathway-related genes appears to play a key role in coordination of these two important signal transduction systems (Fig. 11).
Methods
Fly stocks
Fly strains were maintained at 25°C on standard food. The UAS-DREF line was as described earlier as well as the transgenic fly line carrying GMR-GAL4 on the X chromosome13,51. The UAS-DREFIR fly line was obtained from the Vienna Drosophila Resource Center. The fly line hpoKC202, thj5c8P[lacZ], UAS-GFP, UAS-GFPIR and all other stocks used in this study were obtained from the Bloomington stock center (Indiana) and Drosophila Genetic Resource Center (Kyoto).
Flip-out experiments
RNAi clones in eye discs were generated with a flip-out system53. Female flies with hs-flp; Act5C>FRT y FRT>GAL4, UAS-GFP were crossed with UAS-DREFIR and clones were identified by the presence of green fluorescent protein (GFP) expressed under control of the Act5C promoter. In the control, the female flies with hs-flp; Act5C>FRT y FRT>GAL4, UAS-GFP were crossed with Canton S. Flip-out was induced by heat shock (60 min at 37°C) at 24-48 h after the eggs were laid.
Plasmid construction
To construct the plasmid p5′-hpowt-luc, PCR was performed using Drosophila genomic DNA as a template and the following primer combination:
hpo5′: 5′-ACTACGCGTGCATGTAGATTTCCCAAGCAC
hpo3′: 5′-TCACTCGAGGTGCCCTCGGCACTTTGCAA
PCR products were digested with Mlu1 and Xho1 and inserted between the f1 origin and luciferase (luc) coding region of the pGV-B plasmid (Toyo Ink). Base-substituted derivatives of p5′-hpowt-luc were constructed using a QuickChange Site-Directed Mutagenesis Kit (STRATAGENE) according to the manufacturer's protocol. The following oligonucleotide pairs carrying base substitutions in DRE were used as primers:
hpoDRE1mutF: 5′-GCCAAGAGAACTTCGCGGCAAATGGCTAATTCTGC
hpoDRE1mutR: 5′-GCAGAATTAGCCATTTGCCGCGAAGTTCTCTTGGC
hpoDRE2mutF: 5′-GATGAACAACAACTCGCGGCCCATAAAAGCAATTT
hpoDRE2mutR: 5′-AAATTGCTTTTATGGGCCGCGAGTTGTTGTTCATC
DNA transfection into S2 cells and luciferase assays
Approximately 2 × 105 cells were plated 24 h before plasmid transfection, then 500 ng of reporter plasmid and 1 ng of pAct5C-seapanzy as an internal control were co-transfected into cells using CellFectin II reagent (Invitrogen). After 48 h from transfection, S2 cells were harvested and luciferase activities were normalized to Renilla luciferase activity. Transfections were performed several times with at least two independent plasmid preparations.
For dsRNA interference (RNAi) experiments, 1 × 105 cells were plated in 24-well dishes in the presence of 3 μg/well of DREF-dsRNA, YFP-dsRNA and mock for control (dsRNA free incubation) in fetal bovine serum (FBS) free M3 medium for 1 h. After the incubation, three volumes of M3 containing 10% FBS were added to each well. Four days after RNAi treatment, cells were co-transfected with the luciferase reporter plasmid (500 ng) and pAct5C-seapanzy plasmid (1 ng) as an internal control, then harvested after 48 h for measurement of luciferase activities according to the standard instructions for the kit (Promega).
Quantitative RT-PCR
1 × 106 S2 cells were plated in 6-well dishes in 2 ml M3 medium containing 30 μg/well of double-strand RNAs for DREF (DREFdsRNA) or β-tubulin (β-tubulindsRNA) for 1 h. After incubation, 3 ml of M3 medium containing 10% FBS was added to each well. At 5 days after the RNAi treatment, total RNA was isolated from cells using Trizol Reagent (Invitrogen) and 1 μg aliquots were reverse transcribed with an oligo(dT) primer using a Takara high fidelity RNA PCR kit (Takara). Real-time PCR was performed with SYBR Green I kit (Takara) and the Applied Biosystems 7500 Real-time PCR system using 1 μl of reverse transcribed sample per reaction. β-tubulin mRNA was chosen as a negative control and Ribosomal protein L32 (RpL32/Rp49) as an endogenous reference gene. To carry out quantitative real-time PCR, the following PCR primers were chemically synthesized:
Hpo2nd Primer: 5′-CCTCTTCGGCAGCATCTC
Hpo3rd Primer: 5′-CCGAATCGGAGTTGATTACCATA
DREF-F: 5′-GGCAATCTCCGTTGAATGACG
DREF-R: 5′-TTCACCTCCGAGAAGCCCTT
β-tubulin-F: 5′-AGTTCACCGCTATGTTCA
β-tubulin-R: 5′-CGCAAAACATTGATCGAG
Wts-F: 5′-GCGAGAACAAAGAGGCGAAA
Wts-R: 5′-GGGAATTCTTCCACTCGCATTC
RP49-F: 5′-GCTTCTGGTTTCCGGCAAGCTTCAAG
RP49-R: 5′-GACCTCCAGCTCGCGCACGTTGTGCACCAGGAAC
Chromatin immunoprecipitation
We performed chromatin immunoprecipitation using a Chip assay kit using the protocol recommended by the manufacturer (Millipore). Approximately 1 × 107 S2 cells were fixed in 1% formaldehyde at 37°C for 10 min and then quenched in 125 mM glycine for 5 min at 25°C. Cells were washed twice in PBS containing protease inhibitors (1 mM PMSF, 1 μg/ml aprotinin and 1 μg/ml pepstain A), lysed in 200 μl of SDS lysis buffer, sonicated to break DNA into fragments of less than 1 kb and centrifuged at 15,300 × g for 10 min at 4°C. The sonicated cell supernatants were diluted 10-fold in Chip Dilution Buffer and pre-cleared with 80 μl salmon sperm DNA/protein A agarose-50% slurry for 30 min at 4°C. After brief centrifugation, each supernatant was incubated with 1 μg of normal rabbit IgG or rabbit anti-DREF IgG13 for 16 h at 4°C. Salmon sperm DNA/protein A agarose-50% slurry was added with incubation for 1 h at 4°C. After washing, immunoprecipitated DNA was eluted with elution buffer containing 1% SDS and 0.1 M NaHCO3. The protein-DNA crosslinks were reversed by heating at 65°C for 4 h. After deproteinization with proteinase K, DNA was recovered by phenol-chloroform extraction and ethanol precipitation. Quantitative real-time PCR was performed with the following PCR primers:
ChIPhpoDRE15′Primer: 5′-AGGTAAGCATGCGTGAGTTG
ChIPhpoDRE13′Primer: 5′-GAACACCGCAGAATGTAAACAAAG
ChIPhpoDRE25′Primer: 5′-ACAACGACAAATTGGCACAG
ChIPhpoDRE23′Primer: 5′-TGGATCGATTGTTGTTGTTCATC
ChIPhpoupstreamregion5′Primer: 5′-CACCACCCAAATGACATACG
ChIPhpoupstreamregion3′Primer: 5′-CGCTGCAATCGGAAAGTTA
ChIPwtsDRE1,2 5′Primer: 5′-CCGATAACATTTACTTGCTCTC
ChIPwtsDRE1,2 3′Primer: 5′-GACCGATATCGATAGACAGAG
Immunostaining
For immunostaining, third instar larval eye imaginal discs were dissected and fixed in 4% paraformaldehyde in PBS for 15 min at 25°C. After washing with PBS containing 0.3% Triton X-100 (PBST), samples were blocked with PBS containing 0.15% Triton X-100 and 10% normal goat serum for 30 min at 25°C and incubated with diluted primary antibodies in PBS or in PBS containing 0.15% Triton X-100 and 10% normal goat serum for 16 h at 4°C. The following antibodies were used as primary antibodies: mouse anti-DREF IgG (diluted at 1:200)54, rabbit anti-hpo IgG (1:200; a kind gift from Dr. Helen McNeil)42, rabbit anti-phosphoYki S168 IgG (1:200; a kind gift from Dr. Duojia Pan)32, mouse anti-GFP IgG (1:400; Nacalai tesque), mouse anti-β-galactosidase (1:400, DSHB). After extensive washing with PBST, samples were incubated with secondary antibodies labeled with Alexa 647, Alexa594 and Alexa488 (1:400; Invitrogen) for 3 h at 25°C. After further washing with PBST and PBS, samples were mounted in Vectashield Mounting Medium (Vector laboratories) and inspected with a confocal laser scanning microscope (Olympus FLUOVIEW FV10i).
5-ethynyl-2′-deoxyuridine (EdU) labeling
Detection of cells in S phase was performed using an EdU-labeling kit from Invitrogen (Click-iT EdU Alexa Fluor 594 Imaging Kit). Third instar larvae were dissected in PBS and the imaginal discs were suspended in Grace's insect medium in the presence of 10 μM EdU for 60 min at 25°C. The samples then were fixed with 3.7% formaldehyde in PBS for 15 min at 25°C. After fixing, samples were washed with 3% BSA in PBS and permeabilized in 0.5% Triton X-100 in PBS for 20 min at 25°C. Samples were washed with 3% BSA in PBS and incubated with Click-iT reaction cocktails for 30 min at 25°C according to the manufacturer's instructions. After further washing with 3% BSA in PBS and PBS, samples were stained with Hoechst 33342 (Invitrogen) for labeling DNA and finally samples were mounted and observed as described in immunostaining section.
Apoptosis assay
Detection of apoptotic cells was performed using a Cell Event Caspase-3/7 Green Detection Reagent (Molecular Probes, Invitrogen). Third instar larvae were dissected in PBS and their imaginal discs were incubated in Grace's insect medium containing 5 μM Cell Event Caspase3/7 Green Detection Reagent for 30 min at 37°C. Then, the eye discs were fixed with 3.7% formaldehyde for 15 min at 25°C. After fixing, samples were washed with PBS and permeabilized in 0.5% Triton X-100 in PBS for 20 min at 25°C. After several washings with PBS, samples were mounted and observed as described in the immunostaining section.
Quantification and statistics analysis
EdU signals, apoptosis signals and immunostaining signals in the region posterior to the MF were counted and measured from six independent eye imaginal discs using MetaMorph software (Molecular Devices). The experiments were repeated at least three times. Then, statistical analysis was conducted, as indicated in the figure legends, using GraphPad Prism 6. Every single set of data was assessed using Welch's t-test. Significance is described in the figure legends as *p<0.05, **p<0.01, ***p < 0.001; ****p < 0.0001.
References
Hirose, F., Yamaguchi, M., Handa, H., Inomata, Y. & Matsukage, A. Novel 8-base pair sequence (Drosophila DNA replication-related element) and specific binding factor involved in the expression of Drosophila genes for DNA polymerase alpha and proliferating cell nuclear antigen. J. Biol. Chem. 268, 2092–2099 (1993).
Ohler, U., Liao, G. C., Niemann, H. & Rubin, G. M. Computational analysis of core promoters in the Drosophila genome. Genome Biol. 3, RESEARCH0087 (2002).
Matsukage, A., Hirose, F., Yoo, M. A. & Yamaguchi, M. The DRE/DREF transcriptional regulatory system: a master key for cell proliferation. Biochim. Biophys. Acta. 1779, 81–89, 10.1016/j.bbagrm.2007.11.011 (2008).
Yamaguchi, M., Hayashi, Y. & Matsukage, A. Essential role of E2F recognition sites in regulation of the proliferating cell nuclear antigen gene promoter during Drosophila development. J. Biol. Chem. 270, 25159–25165 (1995).
Okudaira, K. et al. Transcriptional regulation of the Drosophilaorc2 gene by the DREF pathway. Biochim. Biophys. Acta. 1732, 23–30, 10.1016/j.bbaexp.2005.10.009 (2005).
Tsuchiya, A. et al. Transcriptional regulation of the Drosophilarfc1 gene by the DRE-DREF pathway. FEBS J. 274, 1818–1832, 10.1111/j.1742-4658.2007.05730.x (2007).
Ohno, K., Hirose, F., Sakaguchi, K., Nishida, Y. & Matsukage, A. Transcriptional regulation of the Drosophila CycA gene by the DNA replication-related element (DRE) and DRE binding factor (DREF). Nucleic Acids Res. 24, 3942–3946 (1996).
Sawado, T. et al. The DNA replication-related element (DRE)/DRE-binding factor system is a transcriptional regulator of the DrosophilaE2F gene. J. Biol. Chem. 273, 26042–26051 (1998).
Trong-Tue, N., Thao, D. T. P. & Yamaguchi, M. Role of DREF in transcriptional regulation of the Drosophilap53 gene. Oncogene 29, 2060–2069, 10.1038/Onc.2009.483 (2010).
Ida, H., Yoshida, H., Nakamura, K. & Yamaguchi, M. Identification of the DrosophilaeIF4A gene as a target of the DREF transcription factor. Exp. Cell Res. 313, 4208–4220, 10.1016/j.yexcr.2007.08.016 (2007).
Dang, T. P. T., Ida, H., Yoshida, H. & Yamaguchi, M. Identification of the Drosophila skpA gene as a novel target of the transcription factor DREF. Exp. Cell Res. 312, 3641–3650, 10.1016/J.Yexcr.2006.08.005 (2006).
Nakamura, K., Ida, H. & Yamaguchi, M. Transcriptional regulation of the Drosophila moira and osa genes by the DREF pathway. Nucleic Acids Res. 36, 3905–3915, 10.1093/Nar/Gkn291 (2008).
Ida, H. et al. Genetic screening for modifiers of the DREF pathway in Drosophila melanogaster: identification and characterization of HP6 as a novel target of DREF. Nucleic Acids Res. 37, 1423–1437, 10.1093/nar/gkn1068 (2009).
Kawamori, A. & Yamaguchi, M. DREF is critical for Drosophila bristle development by regulating endoreplication in shaft cells. Cell Struct. Funct. 36, 103–119 (2011).
Yoshioka, Y., Nguyen, T. T., Fujiwara, S., Matsuda, R., Valadez-Graham, V. et al. Drosophila DREF acting via the JNK pathway is required for thorax development. Genesis 50, 599–611, 10.1002/dvg.22017 (2012).
Yanai, H. et al. Drosophila myeloid leukemia factor acts with DREF to activate the JNK signaling pathway. Oncogenesis 3, e98, 10.1038/oncsis.2014.13 (2014).
Valadez-Graham, V. et al. XNP/dATRX interacts with DREF in the chromatin to regulate gene expression. Nucleic Acids Res. 40, 1460–1474, 10.1093/Nar/Gkr865 (2012).
Killip, L. E. & Grewal, S. S. DREF is required for cell and organismal growth in Drosophila and functions downstream of the nutrition/TOR pathway. Dev. Biol. 371, 191–202, 10.1016/J.Ydbio.2012.08.020 (2012).
Gurudatta, B. V., Yang, J., Van Bortle, K., Donlin-Asp, P. G. & Corces, V. G. Dynamic changes in the genomic localization of DNA replication-related element binding factor during the cell cycle. Cell Cycle 12, 1605–1615, 10.4161/cc.24742 (2013).
Hochheimer, A. & Tjian, R. Diversified transcription initiation complexes expand promoter selectivity and tissue-specific gene expression. Genes Dev. 17, 1309–1320, 10.1101/gad.1099903 (2003).
Takacs, S., Biessmann, H., Reddy, H. M., Mason, J. M. & Torok, T. Protein interactions on telomericretrotransposons in Drosophila. Int. J. Biol. Sci. 8, 1055–1061, 10.7150/ijbs.4460 (2012).
Fujiwara, S., Ida, H., Yoshioka, Y., Yoshida, H. & Yamaguchi, M. The warts gene as a novel target of the Drosophila DRE/DREF transcription pathway. Am. J. Cancer Res. 2, 36–44 (2012).
Justice, R. W., Zilian, O., Woods, D. F., Noll, M. & Bryant, P. J. The Drosophila tumor-suppressor gene warts encodes a homolog of human Myotonic-Dystrophy Kinase and is required for the control of cell-shape and Proliferation. Genes Dev. 9, 534–546, 10.1101/Gad.9.5.534 (1995).
Xu, T., Wang, W., Zhang, S., Stewart, R. A. & Yu, W. Identifying tumor suppressors in genetic mosaics: the Drosophila lats gene encodes a putative protein kinase. Development 121, 1053–1063 (1995).
Tapon, N. et al. Salvador promotes both cell cycle exit and apoptosis in Drosophila and is mutated in human cancer cell lines. Cell 110, 467–478 (2002).
Harvey, K. F., Pfleger, C. M. & Hariharan, I. K. The Drosophila Mst ortholog, hippo, restricts growth and cell proliferation and promotes apoptosis. Cell 114, 457–467 (2003).
Pantalacci, S., Tapon, N. & Leopold, P. The Salvador partner Hippo promotes apoptosis and cell-cycle exit in Drosophila. Nat. Cell Biol. 5, 921–927, 10.1038/ncb1051 (2003).
Udan, R. S., Kango-Singh, M., Nolo, R., Tao, C. & Halder, G. Hippo promotes proliferation arrest and apoptosis in the Salvador/Warts pathway. Nat. Cell Biol. 5, 914–920, 10.1038/ncb1050 (2003).
Wu, S., Huang, J., Dong, J. & Pan, D. Hippo encodes a Ste-20 family protein kinase that restricts cell proliferation and promotes apoptosis in conjunction with salvador and warts. Cell 114, 445–456 (2003).
Huang, J. B., Wu, S., Barrera, J., Matthews, K. & Pan, D. J. The Hippo signaling pathway coordinately regulates cell proliferation and apoptosis by inactivating Yorkie, the Drosophila homolog of YAP. Cell 122, 421–434, 10.1016/J.Cell.2005.06.007 (2005).
Oh, H. & Irvine, K. D. In vivo regulation of Yorkie phosphorylation and localization. Development 135, 1081–1088, 10.1242/Dev.015255 (2008).
Dong, J. X. et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell 130, 1120–1133, 10.1016/J.Cell.2007.07.019 (2007).
Edgar, B. A. From cell structure to transcription: Hippo forges a new path. Cell 124, 267–273, 10.1016/j.cell.2006.01.005 (2006).
Bandura, J. L. & Edgar, B. A. Yorkie and Scalloped: partners in growth activation. Dev. Cell 14, 315–316, 10.1016/j.devcel.2008.02.010 (2008).
Goulev, Y. et al. SCALLOPED interacts with YORKIE, the nuclear effector of the hippo tumor-suppressor pathway in Drosophila. Curr. Biol. 18, 435–441, 10.1016/j.cub.2008.02.034 (2008).
Wu, S., Liu, Y., Zheng, Y. G., Dong, J. X. & Pan, D. J. The TEAD/TEF family protein scalloped mediates transcriptional output of the hippo growth-regulatory pathway. Dev. Cell 14, 388–398, 10.1016/J.Devcel.2008.01.007 (2008).
Zhang, L. et al. The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev. Cell 14, 377–387, 10.1016/j.devcel.2008.01.006 (2008).
Pan, D. Hippo signaling in organ size control. Genes Dev. 21, 886–897, 10.1101/gad.1536007 (2007).
Reddy, B. V. V. G. & Irvine, K. D. The Fat and Warts signaling pathways: new insights into their regulation, mechanism and conservation. Development 135, 2827–2838, 10.1242/Dev.020974 (2008).
Saucedo, L. J. & Edgar, B. A. Filling out the Hippo pathway. Nat Rev Mol. Cell. Biol. 8, 613–621, 10.1038/nrm2221 (2007).
Zeng, Q. & Hong, W. The emerging role of the hippo pathway in cell contact inhibition, organ size control and cancer development in mammals. Cancer Cell 13, 188–192, 10.1016/j.ccr.2008.02.011 (2008).
Badouel, C. et al. The FERM-Domain Protein Expanded Regulates Hippo Pathway Activity via Direct Interactions with the Transcriptional Activator Yorkie. Dev.Cell 16, 411–420, 10.1016/J.Devcel.2009.01.010 (2009).
Peng, H. W., Slattery, M. & Mann, R. S. Transcription factor choice in the Hippo signaling pathway: homothorax and yorkie regulation of the microRNA bantam in the progenitor domain of the Drosophila eye imaginal disc. Genes Dev. 23, 2307–2319, 10.1101/Gad.1820009 (2009).
Ren, F. et al. Hippo signaling regulates Drosophila intestine stem cell proliferation through multiple pathways. Proc. Natl. Acad. Sci. U. S. A. 107, 21064–21069, 10.1073/pnas.1012759107 (2010).
Hirose, F. et al. Ectopic expression of DREF induces DNA synthesis, apoptosis and unusual morphogenesis in the Drosophila eye imaginal disc: possible interaction with Polycomb and trithorax group proteins. Mol. Cell Biol. 21, 7231–7242, 10.1128/MCB.21.21.7231-7242.2001 (2001).
Kango-Singh, M. et al. Shar-pei mediates cell proliferation arrest during imaginal disc growth in Drosophila. Development 129, 5719–5730, 10.1242/Dev.00168 (2002).
Jia, J., Zhang, W., Wang, B., Trinko, R. & Jiang, J. The Drosophila Ste20 family kinase dMST functions as a tumor suppressor by restricting cell proliferation and promoting apoptosis. Genes Dev. 17, 2514–2519, 10.1101/gad.1134003 (2003).
Lai, Z. C. et al. Control of cell proliferation and apoptosis by mob as tumor suppressor, mats. Cell 120, 675–685, 10.1016/j.cell.2004.12.036 (2005).
Hay, B. A., Wassarman, D. A. & Rubin, G. M. Drosophila homologs of baculovirus inhibitor of apoptosis proteins function to block cell death. Cell 83, 1253–1262 (1995).
Ryoo, H. D., Bergmann, A., Gonen, H., Ciechanover, A. & Steller, H. Regulation of Drosophila IAP1 degradation and apoptosis by reaper and ubcD1. Nat. Cell. Biol. 4, 432–438, 10.1038/ncb795 (2002).
Hirose, F., Yamaguchi, M. & Matsukage, A. Targeted expression of the DNA binding domain of DRE-binding factor, a Drosophila transcription factor, attenuates DNA replication of the salivary gland and eye imaginal disc. Mol. Cell. Biol. 19, 6020–6028 (1999).
Yoshida, H. et al. Over-expression of DREF in the Drosophila wing imaginal disc induces apoptosis and a notching wing phenotype. Genes Cells 6, 877–886 (2001).
Sun, J. & Tower, J. FLP recombinase-mediated induction of Cu/Zn-superoxide dismutase transgene expression can extend the life span of adult Drosophila melanogaster flies. Mol. Cell.Biol. 19, 216–228 (1999).
Hirose, F. et al. Isolation and characterization of cDNA for DREF, a promoter-activating factor for Drosophila DNA replication-related genes. J. Biol. Chem. 271, 3930–3937 (1996).
Acknowledgements
This study was supported by Grants-in-Aid from JSPS, Core-to-Core program, B Asia-Africa Science Platforms and the Ministry of Education, Science, Sports and Culture of Japan. We are grateful to Dr. Helen McNeil and Dr. Duojia Pan for sharing high quality of antibodies; Dr. Malcolm Moore and Dr. Sue Cotterill for comments on the English usage.
Author information
Authors and Affiliations
Contributions
N.V. and M.Y. conceived the project. N.V., T.H. and H. Ya. carried out the experiments. N.V. prepared figures 1–8 and 11. T.H. prepared figures 9–10. N.V., H. Yo. and M.Y. analyzed the data. N.V. wrote the main manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figures
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Vo, N., Horii, T., Yanai, H. et al. The Hippo pathway as a target of the Drosophila DRE/DREF transcriptional regulatory pathway. Sci Rep 4, 7196 (2014). https://doi.org/10.1038/srep07196
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07196
- Springer Nature Limited
This article is cited by
-
The Drosophila histone methyltransferase NSD is positively regulated by the DRE/DREF system
Genes & Genomics (2018)
-
Phosphine inhibits transcription of the catalase gene through the DRE/DREF system in Drosophila melanogaster
Scientific Reports (2017)