Abstract
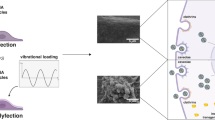
Gene therapies and cellular programming rely on effective cell transfection. Despite continuous advancements in carrier development and transfection techniques to enhance efficiency, the biophysical parameter of extracellular fluid viscosity has been largely overlooked. Here we report a substantial impact of culture media viscosity on transfection efficiency of several delivery vehicles, including lipid nanoparticles, polyplexes, adeno-associated vectors and lentiviral vectors across a range of cell types. We observed substantially increased transfection efficiencies for lipid nanoparticles and polyplexes when the media viscosity matched that of biological fluids (2.0–4.0 centipoise (cP)). This enhancement correlates with higher levels of cellular uptake and improved endosomal escape. Moreover, cells cultured in optimized viscosity conditions exhibit a different profile of uptake pathways compared with those cultured at the standard viscosity of 0.8 cP. This discovery highlights the critical role of media viscosity in the transfection process and provides an additional method to optimize gene delivery and cell programming processes, potentially reducing production costs and increasing the accessibility of gene and cell therapies.







Similar content being viewed by others
Data availability
The main data supporting the results of this study are available within the Article and its Supplementary Information. The raw and analyzed datasets generated during the study are available for research purposes from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Zu, H. & Gao, D. Non-viral vectors in gene therapy: recent development, challenges, and prospects. AAPS J. 23, 78 (2021).
Butt, M. H. et al. Appraisal for the potential of viral and nonviral vectors in gene therapy: a review. Genes 13, 1370 (2022).
Fan, G. et al. Bio-inspired polymer envelopes around adenoviral vectors to reduce immunogenicity and improve in vivo kinetics. Acta Biomater. 30, 94–105 (2016).
Zhu, Y. et al. Optimization of lipid nanoparticles for gene editing of the liver via intraduodenal delivery. Biomaterials 308, 122559 (2024).
Gonçalves, G. A. R. & Paiva, R. d. M. A. Gene therapy: advances, challenges and perspectives. Einstein (São Paulo) 15, 369–375 (2017).
Tian, Y., Hu, D., Li, Y. & Yang, L. Development of therapeutic vaccines for the treatment of diseases. Mol. Biomed. 3, 40 (2022).
Kacherovsky, N. et al. Traceless aptamer-mediated isolation of CD8+ T cells for chimeric antigen receptor T-cell therapy. Nat. Biomed. Eng. 3, 783–795 (2019).
Zhu, Y. et al. Screening for lipid nanoparticles that modulate the immune activity of helper T cells towards enhanced antitumour activity. Nat. Biomed. Eng. 8, 544–560 (2024).
Zhang, Y. et al. Close the cancer-immunity cycle by integrating lipid nanoparticle-mRNA formulations and dendritic cell therapy. Nat. Nanotechnol. 18, 1364–1374 (2023).
Billingsley, M. M. et al. In vivo mRNA CAR T cell engineering via targeted ionizable lipid nanoparticles with extrahepatic tropism. Small 20, e2304378 (2024).
Zhao, Y. et al. Polymetformin combines carrier and anticancer activities for in vivo siRNA delivery. Nat. Comm. 7, 11822 (2016).
Xiong, M. P. et al. Poly(aspartate-g-PEI800), a polyethylenimine analogue of low toxicity and high transfection efficiency for gene delivery. Biomaterials 28, 4889–4900 (2007).
Wang, G. P. et al. Analysis of lentiviral vector integration in HIV+ study subjects receiving autologous infusions of gene modified CD4+ T cells. Mol. Ther. 17, 844–850 (2009).
Ghassemi, S. et al. Rapid manufacturing of non-activated potent CAR T cells. Nat. Biomed. Eng. 6, 118–128 (2022).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Pekrun, K. et al. Using a barcoded AAV capsid library to select for clinically relevant gene therapy vectors. JCI Insight 4, e131610 (2019).
Metzloff, A. E. et al. Antigen presenting cell mimetic lipid nanoparticles for rapid mRNA CAR T cell cancer immunotherapy. Adv. Mater. 36, e2313226 (2024).
Nemir, S. & West, J. L. Synthetic materials in the study of cell response to substrate rigidity. Ann. Biomed. Eng. 38, 2–20 (2010).
Yankaskas, C. L. et al. The fluid shear stress sensor TRPM7 regulates tumor cell intravasation. Sci. Adv. 7, eabh3457 (2021).
Zhao, R. et al. Cell sensing and decision-making in confinement: the role of TRPM7 in a tug of war between hydraulic pressure and cross-sectional area. Sci. Adv. 5, eaaw7243 (2019).
Maity, D. et al. Extracellular hydraulic resistance enhances cell migration. Adv. Sci. 9, e2200927 (2022).
Bera, K. et al. Extracellular fluid viscosity enhances cell migration and cancer dissemination. Nature 611, 365–373 (2022).
Zhang, Y. et al. Polarized NHE1 and SWELL1 regulate migration direction, efficiency and metastasis. Nat. Comm. 13, 6128 (2022).
Goult, B. T., von Essen, M. & Hytönen, V. P. The mechanical cell—the role of force dependencies in synchronising protein interaction networks. J. Cell Sci. 135, jcs259769 (2022).
Sun, L., Suo, C., Li, S.-T., Zhang, H. & Gao, P. Metabolic reprogramming for cancer cells and their microenvironment: beyond the Warburg effect. Biochim. Biophys. Acta, Rev. Cancer 1870, 51–66 (2018).
Mayor, S. & Pagano, R. E. Pathways of clathrin-independent endocytosis. Nat. Rev. Mol. Cell Biol. 8, 603–612 (2007).
Parton, R. G. et al. Caveolae: the FAQs. Traffic 21, 181–185 (2020).
Donahue, N. D., Acar, H. & Wilhelm, S. Concepts of nanoparticle cellular uptake, intracellular trafficking, and kinetics in nanomedicine. Adv. Drug Deliv. Rev. 143, 68–96 (2019).
Pegu, A. et al. Durability of mRNA-1273 vaccine-induced antibodies against SARS-CoV-2 variants. Science 373, 1372–1377 (2021).
Zhang, L. et al. Effect of mRNA-LNP components of two globally-marketed COVID-19 vaccines on efficacy and stability. npj Vaccines 8, 156 (2023).
Pittman, M. et al. Membrane ruffling is a mechanosensor of extracellular fluid viscosity. Nat. Phys. 18, 1112–1121 (2022).
Ryter, A. Relationship between ultrastructure and specific functions of macrophages. Comp. Immunol. Microbiol. Infect. Dis. 8, 119–133 (1985).
Hu, Y. et al. Size-controlled and shelf-stable DNA particles for production of lentiviral vectors. Nano Lett. 21, 5697–5705 (2021).
Rui, Y. et al. High-throughput and high-content bioassay enables tuning of polyester nanoparticles for cellular uptake, endosomal escape, and systemic in vivo delivery of mRNA. Sci. Adv. 8, eabk2855 (2022).
Rennick, J. J., Johnston, A. P. R. & Parton, R. G. Key principles and methods for studying the endocytosis of biological and nanoparticle therapeutics. Nat. Nanotechnol. 16, 266–276 (2021).
Zhu, Y. et al. Albumin-biomineralized nanoparticles to synergize phototherapy and immunotherapy against melanoma. J. Control. Rel. 322, 300–311 (2020).
Nyamay’Antu, A., Kédinger, V. & Erbacher, P. Simplifying the efficient clinical-grade production of viruses. Genet. Eng. Biotechnol. News 37, 24–25 (2017).
Meng, M. & Wu, Y.-C. Combination of AAV-CCL19 and GPC3 CAR-T cells in the treatment of hepatocellular carcinoma. J. Immunol. Res. 2021, 1782728 (2021).
Moço, P. D., Farnós, O., Sharon, D. & Kamen, A. A. Targeted delivery of chimeric antigen receptor into T cells via CRISPR-mediated homology-directed repair with a dual-AAV6 transduction system. Curr. Issues Mol. Biol. 45, 7705–7720 (2023).
Sawaisorn, P. et al. Comparison of the efficacy of second and third generation lentiviral vector transduced CAR CD19 T cells for use in the treatment of acute lymphoblastic leukemia both in vitro and in vivo models. PLoS ONE 18, e0281735 (2023).
Yang, P. et al. CD24 is a novel target of chimeric antigen receptor T cells for the treatment of triple negative breast cancer. Cancer Immunol. Immunother. 72, 3191–3202 (2023).
Zhou, J.-E. et al. Lipid nanoparticles produce chimeric antigen receptor T cells with interleukin-6 knockdown in vivo. J. Control. Rel. 350, 298–307 (2022).
Billingsley, M. M. et al. Ionizable lipid nanoparticle-mediated mRNA delivery for human CAR T cell engineering. Nano Lett. 20, 1578–1589 (2020).
Gray, S. J. et al. Production of recombinant adeno-associated viral vectors and use in in vitro and in vivo administration. Curr. Protoc. Neurosci. 57, 4.17.1–4.17.30 (2011).
Edwards, D. A., Gooch, K. J., Zhang, I., McKinley, G. H. & Langer, R. The nucleation of receptor-mediated endocytosis. Proc. Natl Acad. Sci. USA 93, 1786–1791 (1996).
Muto, S., Ohtaka, A., Nemoto, J., Kawakami, K. & Asano, Y. Effects of hyperosmolality on Na, K-ATPase gene expression in vascular smooth muscle cells. J. Membr. Biol. 162, 233–245 (1998).
Yue, Y. & Wu, C. Progress and perspectives in developing polymeric vectors for in vitro gene delivery. Biomater. Sci. 1, 152–170 (2013).
Kanasty, R., Dorkin, J. R., Vegas, A. & Anderson, D. Delivery materials for siRNA therapeutics. Nat. Mater. 12, 967–977 (2013).
Dahlman, J. E. et al. Barcoded nanoparticles for high throughput in vivo discovery of targeted therapeutics. Proc. Natl Acad. Sci. USA 114, 2060–2065 (2017).
Da Silva Sanchez, A. J. et al. Universal barcoding predicts in vivo ApoE-independent lipid nanoparticle delivery. Nano Lett. 22, 4822–4830 (2022).
Zhu, Y. et al. Multi-step screening of DNA/lipid nanoparticles and co-delivery with siRNA to enhance and prolong gene expression. Nat. Comm. 13, 4282 (2022).
Acknowledgements
The authors disclose support for the research described in this study from the National Institutes of Health (Grant numbers: R01 GM134542 to S.X.S. and K.K.; R01 CA257647 to K.K.). We thank J. Schneck from the Department of Pathology at the Johns Hopkins University School of Medicine for providing the human PBMCs as a gift and H. Bui from the Johns Hopkins University Integrated Imaging Center (IIC) for assistance with the flow cytometry assessments. Portions of the schematics in Figs. 3–6 and Supplementary Figs. 1, 2 and 13 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
H.-Q.M., S.X.S., Y.Z. and J.M. conceived and designed this study. J.M., Y.Z., J.K., D.Y., W.H.T., M.J. and J.L. performed the experiments. J.M., Y.Z., J.K., D.Y., W.H.T., M.J., Q.N., Z.G., J.C., K.K., M.F.K., S.X.S. and H.-Q.M. contributed to the data analysis and interpretation. J.M., Y.Z. and H.-Q.M. wrote the manuscript with input from all other authors. H.-Q.M. and S.X.S. secured the funding and supervised this study.
Corresponding authors
Ethics declarations
Competing interests
H.-Q.M., S.X.S., J.M., Y.Z., Q.N. and Z.G. are co-inventors of a patent application (PCT/US2024/039036, filed in July 2024) covering the compositions and transfection methods described in this paper, filed through and managed by Johns Hopkins Office of Technology Ventures. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Engineering thanks Michael Mitchell, Suzie Pun and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–20 and Tables 1–3.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, J., Zhu, Y., Kong, J. et al. Tuning extracellular fluid viscosity to enhance transfection efficiency. Nat Chem Eng (2024). https://doi.org/10.1038/s44286-024-00116-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44286-024-00116-3
- Springer Nature America, Inc.