Abstract
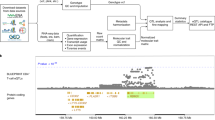
Detecting genetic variants associated with traits (quantitative trait loci, QTL) requires genotyped study individuals. Here we describe BaseQTL, a Bayesian method that exploits allele-specific expression to map molecular QTL from sequencing reads (eQTL for gene expression) even when no genotypes are available. When used with genotypes to map eQTL, BaseQTL has lower error rates and increased power compared with existing QTL mapping methods. Running without genotypes limits how many tests can be performed, but due to the proximity of QTL variants to gene bodies, the 2.8% of variants within a 100 kB window that could be tested contained 26% of eQTL detectable with genotypes. eQTL effect estimates were invariably consistent between analyses performed with and without genotypes. Often, sequencing data may be generated in the absence of genotypes on patients and controls in differential expression studies, and we identified an apparent psoriasis-specific eQTL for GSTP1 in one such dataset, providing new insights into disease-dependent gene regulation.






Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Data availability
Geuvadis samples were accessed from E-GEUV-1, ftp://ftp.sra.ebi.ac.uk/vol1/fastq, on 16 April 2017 or 23 January 2018 as indicated in Supplementary Table 1. Psoriasis and normal skin samples were accessed from E-GEOD-54456, ftp://ftp.sra.ebi.ac.uk/vol1/fastq, on 2 November 2018. GTEx associations for skin, blood and lymphoblastic cell lines corresponding to Analysis V7 were downloaded from https://gtexportal.org/home/datasets on 21 June 2019. Differentially regulated genes between psoriasis and normal skin were downloaded from https://ars.els-cdn.com/content/image/1-s2.0-S0022202X15368834-mmc2.xls on the 21 November 2018. We downloaded RNA-seq data from 86 Geuvadis samples with EUR ancestry (GBR code) from ArrayExpress (E-GEUV-1, Supplementary Table 1). We also analyzed 94 and 90 RNA-seq normal and psoriasis skin samples13 obtained from ArrayExpress (E-GEOD-54456). For the analysis of psoriasis eQTL we selected 51 upregulated genes in psoriasis versus normal skin (P ≤ 10−6 corresponding to family-wise error rate <0.025) and with a median expression of at least 500 RPKM in psoriasis samples (data extracted from https://ars.els-cdn.com/content/image/1-s2.0-S0022202X15368834-mmc2.xls13, and/or within 100 kB of a psoriasis GWAS hit 25 (380 genes). Datasets to reproduce figures in this paper were uploaded into Zenodo 41.
Code availability
The source code and documentation for BaseQTL are available at https://gitlab.com/evigorito/baseqtl. We also provide a pipeline to process RNA fastq files and genotypes, if available, to prepare for running BaseQTL at https://gitlab.com/evigorito/baseqtl_pipeline (Supplementary Fig. 12 and Supplementary Section 3). The code to reproduce the figures is available at https://gitlab.com/evigorito/baseqtl_paper. The three repositories have been uploaded to Zenodo41.
References
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Chun, S. et al. Limited statistical evidence for shared genetic effects of eQTLs and autoimmune-disease-associated loci in three major immune-cell types. Nat. Genet. 49, 600–605 (2017).
Guo, H. et al. Integration of disease association and eQTL data using a Bayesian colocalisation approach highlights six candidate causal genes in immune-mediated diseases. Hum. Mol. Genet. 24, 3305–3313 (2015).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Zhernakova, D. V. et al. Identification of context-dependent expression quantitative trait loci in whole blood. Nat. Genet. 49, 139–145 (2017).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014).
Gamazon, E. R. et al. Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation. Nat. Genet. 50, 956–967 (2018).
Wall, J. D. et al. Estimating genotype error rates from high-coverage next-generation sequence data. Genome Res. 24, 1734–1739 (2014).
Peters, J. E. et al. Insight into genotype-phenotype associations through eQTL mapping in multiple cell types in health and immune-mediated disease. PLoS Genet. 12, e1005908 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Carpenter, B. et al. Stan: a probabilistic programming language. J. Stat. Softw. 76, 1–32 (2017).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Li, B. et al. Transcriptome analysis of psoriasis in a large case-control sample: RNA-seq provides insights into disease mechanisms. J. Invest. Dermatol. 134, 1828–1838 (2014).
Kumasaka, N., Knights, A. J. & Gaffney, D. J. Fine-mapping cellular QTLs with RASQUAL and ATAC-seq. Nat. Genet. 48, 206–213 (2016).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Sun, W. A statistical framework for eQTL mapping using RNA-seq data. Biometrics 68, 1–11 (2012).
Hu, Y.-J., Sun, W., Tzeng, J.-Y. & Perou, C. M. Proper use of allele-specific expression improves statistical power for cis-eQTL mapping with RNA-seq data. J. Am. Stat. Assoc. 110, 962–974 (2015).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 1, 457–470 (2011).
Degner, J. F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Liu, Z. et al. Comparing computational methods for identification of allele-specific expression based on next generation sequencing data. Genet. Epidemiol. 38, 591–598 (2014).
Castel, S. E., Levy-Moonshine, A., Mohammadi, P., Banks, E. & Lappalainen, T. Tools and best practices for data processing in allelic expression analysis. Genome Biol. 16, 195 (2015).
Stranger, B. E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Brown, A. A. et al. Predicting causal variants affecting expression by using whole-genome sequencing and RNA-seq from multiple human tissues. Nat. Genet. 49, 1747+ (2017).
Võsa, U. et al. Unraveling the polygenic architecture of complex traits using blood eQTL meta-analysis. Preprint at bioRxiv https://doi.org/10.1101/447367 (2018).
Tsoi, L. C. et al. Large scale meta-analysis characterizes genetic architecture for common psoriasis associated variants. Nat. Commun. 8, 15382 (2017).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Ding, J. et al. Gene expression in skin and lymphoblastoid cells: refined statistical method reveals extensive overlap in cis-eQTL signals. Am. J. Hum. Genet. 87, 779–789 (2010).
Gudjonsson, J. E. et al. Assessment of the psoriatic transcriptome in a large sample: additional regulated genes and comparisons with in vitro models. J. Invest. Dermatol. 130, 1829–1840 (2010).
Schalkwijk, J., Chang, A., Janssen, P., De Jongh, G. J. & Mier, P. D. Skin-derived antileucoproteases (SKALPs): characterization of two new elastase inhibitors from psoriatic epidermis. Br. J. Dermatol. 122, 631–641 (1990).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414 (2016).
Joehanes, R. et al. Integrated genome-wide analysis of expression quantitative trait loci aids interpretation of genomic association studies. Genome Biol. 18, 16 (2017).
Nestle, F. O., Kaplan, D. H. & Barker, J. Psoriasis. N. Engl. J. Med. 361, 496–509 (2009).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Dobin, A. & Gingeras, T. R. Mapping RNA-seq reads with STAR. Curr. Protoc. Bioinformatics 51, 11.14.1–11.14.19 (2015).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Castel, S. E., Mohammadi, P., Chung, W. K., Shen, Y. & Lappalainen, T. Rare variant phasing and haplotypic expression from RNA sequencing with phASER. Nat. Commun. 7, 12817 (2016).
Delaneau, O., Marchini, J. & Zagury, J.-F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2011).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Muller, P., Parmigiani, G. & Rice, K. FDR and Bayesian Multiple Comparisons Rules Working Paper (Johns Hopkins University, Department of Biostatistics, 2006).
Vigorito, E. et al. Dataset to reproduce BaseQTL figures. Zenodo https://doi.org/10.5281/zenodo.4759202 (2021).
Acknowledgements
This work was co-funded by the Wellcome Trust (WT107881), the MRC (MC_UU_00002/2, MC_UU_00002/4, MC_UU_00002/13, MR/R013926/1 (to the CLUSTER Consortium)) and the NIHR Cambridge Biomedical Research Centre (BRC-1215-20014). S.R.W. was supported by the NIHR Cambridge Biomedical Research Centre. The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care. This research was funded in whole, or in part, by the Wellcome Trust (WT107881). For the purpose of open access, the author has applied a CC BY public copyright licence to any author accepted manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
C.W. conceived of the project. E.V., C.W. and S.R.W. developed the model. E.V. wrote the software and performed analyses. W.-Y.L. and C.S. performed analyses and implemented the software. P.D.W.K. and S.R.W. contributed to the design of statistical analysis. E.V. and C.W. wrote the manuscript with input from all authors. C.W. directed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Computational Science thanks Eric Gamazon, Wei Sun and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Handling editor: Ananya Rastogi, in collaboration with the Nature Computational Science team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–17, Figs. 1–21, Sections 1–4 and References.
Supplementary Data 1
Geuvadis samples used in this study.
Supplementary Data 2
eQTL estimates for psoriasis and normal skin running individual models.
Supplementary Data 3
eQTL estimates for psoriasis and normal skin running a joint model.
Rights and permissions
About this article
Cite this article
Vigorito, E., Lin, WY., Starr, C. et al. Detection of quantitative trait loci from RNA-seq data with or without genotypes using BaseQTL. Nat Comput Sci 1, 421–432 (2021). https://doi.org/10.1038/s43588-021-00087-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-021-00087-y
- Springer Nature America, Inc.
This article is cited by
-
DeCAF: a novel method to identify cell-type specific regulatory variants and their role in cancer risk
Genome Biology (2022)
-
Detecting context-dependent gene regulation
Nature Computational Science (2021)