Abstract
In pancreatic ductal adenocarcinoma (PDAC), glutamine is a critical nutrient that drives a wide array of metabolic and biosynthetic processes that support tumor growth. Here, we elucidate how 6-diazo-5-oxo-l-norleucine (DON), a glutamine antagonist that broadly inhibits glutamine metabolism, blocks PDAC tumor growth and metastasis. We find that DON significantly reduces asparagine production by inhibiting asparagine synthetase (ASNS), and that the effects of DON are rescued by asparagine. As a metabolic adaptation, PDAC cells upregulate ASNS expression in response to DON, and we show that ASNS levels are inversely correlated with DON efficacy. We also show that l-asparaginase (ASNase) synergizes with DON to affect the viability of PDAC cells, and that DON and ASNase combination therapy has a significant impact on metastasis. These results shed light on the mechanisms that drive the effects of glutamine mimicry and point to the utility of cotargeting adaptive responses to control PDAC progression.






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Source data are provided with this paper.
References
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Lyssiotis, C. A., Son, J., Cantley, L. C. & Kimmelman, A. C. Pancreatic cancers rely on a novel glutamine metabolism pathway to maintain redox balance. Cell Cycle 12, 1987–1988 (2013).
Cluntun, A. A., Lukey, M. J., Cerione, R. A. & Locasale, J. W. Glutamine metabolism in cancer: understanding the heterogeneity. Trends Cancer 3, 169–180 (2017).
Zhang, J., Pavlova, N. N. & Thompson, C. B. Cancer cell metabolism: the essential role of the nonessential amino acid, glutamine. EMBO J. 36, 1302–1315 (2017).
Kamphorst, J. J. et al. Human pancreatic cancer tumors are nutrient poor and tumor cells actively scavenge extracellular protein. Cancer Res. 75, 544–553 (2015).
Recouvreux, M. V. et al. Glutamine depletion regulates Slug to promote EMT and metastasis in pancreatic cancer. J. Exp. Med. https://doi.org/10.1084/jem.20200388 (2020).
Lee, S. W. et al. EGFR-Pak signaling selectively regulates glutamine deprivation-induced macropinocytosis. Dev. Cell 50, 381–392.e385 (2019).
Luengo, A., Gui, D. Y. & Vander Heiden, M. G. Targeting metabolism for cancer therapy. Cell Chem. Biol. 24, 1161–1180 (2017).
Hensley, C. T., Wasti, A. T. & DeBerardinis, R. J. Glutamine and cancer: cell biology, physiology, and clinical opportunities. J. Clin. Invest. 123, 3678–3684 (2013).
Daye, D. & Wellen, K. E. Metabolic reprogramming in cancer: unraveling the role of glutamine in tumorigenesis. Semin. Cell Dev. Biol. 23, 362–369 (2012).
Wise, D. R. & Thompson, C. B. Glutamine addiction: a new therapeutic target in cancer. Trends Biochem. Sci. 35, 427–433 (2010).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Commisso, C. et al. Macropinocytosis of protein is an amino acid supply route in Ras-transformed cells. Nature 497, 633–637 (2013).
Hosios, A. M. et al. Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Dev. Cell 36, 540–549 (2016).
Shen, Y. A. et al. Inhibition of glutaminolysis in combination with other therapies to improve cancer treatment. Curr. Opin. Chem. Biol. 62, 64–81 (2021).
Biancur, D. E. et al. Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nat. Commun. 8, 15965 (2017).
Lemberg, K. M., Vornov, J. J., Rais, R. & Slusher, B. S. We’re not ‘DON’ yet: optimal dosing and prodrug delivery of 6-diazo-5-oxo-l-norleucine. Mol. Cancer Ther. 17, 1824–1832 (2018).
Tenora, L. et al. Tumor-targeted delivery of 6-diazo-5-oxo-l-norleucine (DON) using substituted acetylated lysine prodrugs. J. Med. Chem. 62, 3524–3538 (2019).
Rais, R. et al. Discovery of 6-diazo-5-oxo-l-norleucine (DON) prodrugs with enhanced CSF delivery in monkeys: a potential treatment for glioblastoma. J. Med. Chem. 59, 8621–8633 (2016).
Nedelcovych, M. T. et al. N-(pivaloyloxy)alkoxy-carbonyl prodrugs of the glutamine antagonist 6-diazo-5-oxo-l-norleucine (DON) as a potential treatment for HIV associated neurocognitive disorders. J. Med. Chem. 60, 7186–7198 (2017).
Hanaford, A. R. et al. Orally bioavailable glutamine antagonist prodrug JHU-083 penetrates mouse brain and suppresses the growth of MYC-driven medulloblastoma. Transl. Oncol. 12, 1314–1322 (2019).
Rais, R. et al. Discovery of DRP-104, a tumor-targeted metabolic inhibitor prodrug. Sci. Adv. 8, eabq5925 (2022).
Leone, R. D. et al. Glutamine blockade induces divergent metabolic programs to overcome tumor immune evasion. Science 366, 1013–1021 (2019).
Sharma, N. S. et al. Targeting tumor-intrinsic hexosamine biosynthesis sensitizes pancreatic cancer to anti-PD1 therapy. J. Clin. Invest. 130, 451–465 (2020).
Wang, Z. et al. Targeting glutaminolysis: new perspectives to understand cancer development and novel strategies for potential target therapies. Front. Oncol. 10, 589508 (2020).
Krall, A. S., Xu, S., Graeber, T. G., Braas, D. & Christofk, H. R. Asparagine promotes cancer cell proliferation through use as an amino acid exchange factor. Nat. Commun. 7, 11457 (2016).
Pavlova, N. N. et al. As extracellular glutamine levels decline, asparagine becomes an essential amino acid. Cell Metab. 27, 428–438.e425 (2018).
Pinkus, L. M. Glutamine binding sites. Methods Enzymol. 46, 414–427 (1977).
Rosenbluth, R. J., Cooney, D. A., Jayaram, H. N., Milman, H. A. & Homan, E. R. DON, CONV and DONV-II. Inhibition of l-asparagine synthetase in vivo. Biochem. Pharmacol. 25, 1851–1858 (1976).
Bayne, L. J. et al. Tumor-derived granulocyte-macrophage colony-stimulating factor regulates myeloid inflammation and T cell immunity in pancreatic cancer. Cancer Cell 21, 822–835 (2012).
Khanna, C. & Hunter, K. Modeling metastasis in vivo. Carcinogenesis 26, 513–523 (2005).
Bott, A. J. et al. Glutamine anabolism plays a critical role in pancreatic cancer by coupling carbon and nitrogen metabolism. Cell Rep. 29, 1287–1298.e1286 (2019).
Tsai, P. Y. et al. Adaptation of pancreatic cancer cells to nutrient deprivation is reversible and requires glutamine synthetase stabilization by mTORC1. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2003014118 (2021).
Balasubramanian, M. N., Butterworth, E. A. & Kilberg, M. S. Asparagine synthetase: regulation by cell stress and involvement in tumor biology. Am. J. Physiol. Endocrinol. Metab. 304, E789–E799 (2013).
Muller, H. J. & Boos, J. Use of l-asparaginase in childhood ALL. Crit. Rev. Oncol. Hematol. 28, 97–113 (1998).
Egler, R. A., Ahuja, S. P. & Matloub, Y. l-asparaginase in the treatment of patients with acute lymphoblastic leukemia. J. Pharmacol. Pharmacother. 7, 62–71 (2016).
Ponz-Sarvise, M. et al. Identification of resistance pathways specific to malignancy using organoid models of pancreatic cancer. Clin. Cancer Res. 25, 6742–6755 (2019).
Tiriac, H. et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 8, 1112–1129 (2018).
Villarino, N. et al. A screen for inducers of bHLH activity identifies pitavastatin as a regulator of p21, Rb phosphorylation and E2F target gene expression in pancreatic cancer. Oncotarget 8, 53154–53167 (2017).
Sullivan, M. R. et al. Quantification of microenvironmental metabolites in murine cancers reveals determinants of tumor nutrient availability. eLife https://doi.org/10.7554/eLife.44235 (2019).
Van Trimpont, M. et al. Novel insights on the use of l-asparaginase as an efficient and safe anti-cancer therapy. Cancers https://doi.org/10.3390/cancers14040902 (2022).
Krall, A. S. et al. Asparagine couples mitochondrial respiration to ATF4 activity and tumor growth. Cell Metab. 33, 1013–1026 e1016 (2021).
Pathria, G. et al. Translational reprogramming marks adaptation to asparagine restriction in cancer. Nat. Cell Biol. 21, 1590–1603 (2019).
Knott, S. R. V. et al. Asparagine bioavailability governs metastasis in a model of breast cancer. Nature 554, 378–381 (2018).
Mukherjee, P. et al. Therapeutic benefit of combining calorie-restricted ketogenic diet and glutamine targeting in late-stage experimental glioblastoma. Commun. Biol. 2, 200 (2019).
Johnson, M. L. et al. Phase 1 and phase 2a, first-in-human (FIH) study, of DRP-104, a broad glutamine antagonist, in adult patients with advanced solid tumors. J. Clin. Oncol. 39, TPS3149 (2021).
Wang, S. Z. et al. Unbiased metabolic profiling predicts sensitivity of high MYC-expressing atypical teratoid/rhabdoid tumors to glutamine inhibition with 6-diazo-5-oxo-l-norleucine. Clin. Cancer Res. 25, 5925–5936 (2019).
Pham, K. et al. Novel glutamine antagonist JHU395 suppresses MYC-driven medulloblastoma growth and induces apoptosis. J. Neuropathol. Exp. Neurol. 80, 336–344 (2021).
Wise, D. R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. USA 105, 18782–18787 (2008).
Qing, G. et al. ATF4 regulates MYC-mediated neuroblastoma cell death upon glutamine deprivation. Cancer Cell 22, 631–644 (2012).
Mihara, E. et al. Active and water-soluble form of lipidated Wnt protein is maintained by a serum glycoprotein afamin/alpha-albumin. eLife https://doi.org/10.7554/eLife.11621 (2016).
Seino, T. et al. Human pancreatic tumor organoids reveal loss of stem cell niche factor dependence during disease progression. Cell Stem Cell 22, 454–467.e456 (2018).
Acknowledgements
We thank members of the Commisso laboratory for their helpful comments and discussions. This work was supported by an American Cancer Society Discovery Boost Grant (no. DBG-22-172-01-TBE) and NIH grant nos R01CA254806 and R01CA207189 to C.C. SBP core services are supported by NCI Cancer Center Support grant no. P30CA030199.
Author information
Authors and Affiliations
Contributions
M.V.R. and C.C. designed the study, wrote the manuscript and prepared all figures. M.V.R., S.F.G., Y.Z., G.L., C.M.G., S.M., K.D.-P., D.B., R.N., D.A.S. and C.C. performed experiments and/or analyzed data. E.E., A.M.L. and H.T. designed and performed the PDO experiments and analyzed the results. C.C. supervised the study. All authors reviewed, edited or commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DON treatment minimally affects animal body weight and tumor cell apoptosis, but significantly impacts tumor cell proliferation.
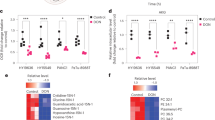
(a) Body weights of mice bearing subcutaneous or orthotopic PDAC tumors after treatment with vehicle (Ctrl) or DON. Data are presented as mean ± SEM. KPC heterotopic (n = 4, P = 0.0167), KPC orthotopic (n = 8 and n = 9, P = 0.6737), PaTu 8988 T orthotopic (n = 7, P = 0.0078), and 779E orthotopic (n = 8 and n = 7, P = 0.2302). (b) Immunohistochemical staining of the apoptosis marker Cleaved Caspase 3 (CC3) in KPC and 779E orthotopic tumors treated with vehicle (Ctrl) or DON (10 mg/kg). Representative images are shown of n = 3 independent experiments. Red scale bar 50 µm. (c) Immunohistochemical staining of the proliferation marker pHis-H3 in 779E orthotopic tumors treated with vehicle (Ctrl) or DON (10 mg/kg). Representative images are shown. Scale bar 100 µm. Quantification of pHis-H3-positive nuclei/ image field is shown as mean ± SEM of n = 4 tumors per group. P = 0.0057. Statistical significance was calculated using unpaired two-tailed Student’s t test (a, c). *P < 0.05, **P < 0.01, ns = not significant.
Extended Data Fig. 2 DON suppresses lung micrometastases in an i.v. injection mouse model.
(a) Immunohistochemical staining of p53 in lung tissue sections derived from animals i.v. injected with human HPAF-II cells. Representative images are shown. Black arrows indicate p53+ cells. Scale bar 50 µm. (b) Quantification of p53 staining in lung tissue sections. Data are presented as Min to Max box-and-whisker plot with median indicated at center line for n = 64 or n = 77 images per condition (Ctrl, DON) from a total of 4 or 5 mice per group (Ctrl, DON; P = 0.0002). Statistical significance was calculated using unpaired two-tailed Student’s t test. ***P < 0.001.
Extended Data Fig. 3 DON dose response curves and IC50 determinations for a panel of PDAC cells.
(a) DON dose response curves for the indicated human PDAC cell lines. Data are presented relative to the 0 μM DON condition and are representative of 3 independent experiments (MIA PaCa-2) or an average of 3 independent experiments (BxPC-3, Capan-2, CFPAC-1, PANC-1, PANC 10.05, PL45, SW-1990). Each independent experiment was performed with 3 replicates per condition. Data are presented as mean ± SEM. (b) Summary of the average IC50 from at least 3 independent experiments for each of the cell lines indicated. (c) Graphical representation of the average IC50 values presented in (b).
Extended Data Fig. 4 Effects of DON on cell fitness are selectively rescued by asparagine supplementation.
(a, b) Relative number of 779E (a) or PaTu 8988 T (b) cells treated with or without DON (1 mM or 10 mM, respectively) with supplementation of the indicated metabolites for 24 h. Cells were stained with crystal violet and stained area was quantified. Data are presented relative to the −DON condition and are representative of 3 independent experiments. Data are presented as mean ± SEM of triplicate wells. 779E cells: −DON vs +DON, P < 0.0001; +DON vs NEAAs, P < 0.0001. PaTu 8988 T cells: −DON vs +DON, P = 0.0002; +DON vs NEAAs, P = 0.0366. (c), (d) Relative number of 779E (c) or PaTu 8988 T (d) cells treated with or without DON (1 mM or 10 mM, respectively) supplemented with a cocktail of NEAAs or the indicated individual amino acids (0.1 mM) for 24 h. Data are presented relative to the −DON condition and are representative of 3 independent experiments. Data are presented as mean ± SEM of triplicate wells. 779E cells: +DON vs NEAAs, P < 0.0001; +DON vs Asn, P = 0.0001. PaTu 8988 T cells: For all statistical comparisons, P < 0.0001. (e), (f) Relative number of 779E cells (e) or MIA PaCa-2 cells (f) treated with DON at the indicated concentrations with or without 0.1 mM Asn supplementation. Data are presented relative to the no DON condition and are representative of 3 independent experiments. Data are presented as mean ± SEM of triplicate wells. 779E cells: 0.25 mM, P = 0.0004; 0.5 mM, P = 0.0014; 1 mM, P = 0.0004. MIA PaCa-2 cells: 5 mM, P = 0.0033; 25 mM, P = 0.0016; 125 mM, P = 0.0002. Statistical significance was calculated using one-way ANOVA followed by Dunnet’s multiple comparisons test (a–d) or unpaired two-tailed Student’s t test (e, f). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 5 DON treatment affects polar metabolite pools.
(a) Quantification of intracellular asparagine (Asn) levels in KPC cells treated with or without 10 μM DON and supplemented with or without 0.1 mM Asn for 24 h. Data are presented relative to vehicle control (Ctrl). Data are presented as mean ± SEM of at least n = 4 samples. Ctrl vs Asn, P = 0.0043. Ctrl vs DON, P < 0.0001. Asn vs DON+Asn, P = 0.0078. DON vs DON+Asn, P = 0.0074. (b) Quantification of intracellular aspartate (Asp) levels in PaTu 8988 T cells treated with or without DON at the indicated concentrations for 24 h. Data are presented relative to the 0 μM DON condition. Data are presented as mean ± SEM of n = 3 samples. 0 vs 2.5 μM, P = 0.1163. 2.5 μM vs 5 μM, P = 0.0011. 5 μM vs 10 μM, P = 0.0008. For all other statistical comparisons, P < 0.0001. (c–e) Quantification of intracellular levels of the indicated metabolites in KPC (c), 779E (d) or PaTu 8988 T (e) cells treated with vehicle control (Ctrl) or DON (10 μM, 10 μM, and 1 mM, respectively) for 24 h. Data are presented relative to Ctrl. Data are presented as mean ± SEM of n = 5 samples (c, d) and n = 3 samples (e). KPC cells: Glu, P = 0.0006. 779E cells: Succinate, P = 0.0003. PaTu 8988 T cells: Citrate, P = 0.0032. a-Ketoglutarate, P = 0.0009. Succinate, P = 0.0046. Fumarate, P = 0.0074. Malate, P = 0.013. For all other statistical comparisons, P < 0.0001. (f) Quantification of intracellular levels of the indicated metabolites in KPC cells treated with vehicle control (Ctrl), 10 μM DON and 10 μM DON supplemented with 0.1 mM Asn for 24 h. Data are presented relative to untreated control. Data are presented as mean ± SEM of at least n = 4 samples. Glu: Ctrl vs DON, P = 0.0018. DON vs DON+Asn, P = 0.0102. Citrate: Ctrl vs DON, P = 0.0060. DON vs DON+Asn, P = 0.0621. α-Ketoglutarate: Ctrl vs DON, P = 0.0029. DON vs DON+Asn, P = 0.0012. Succinate: Ctrl vs DON, P = 0.0166. DON vs DON+Asn, P = 0.4256. Fumarate: Ctrl vs DON, P = 0.0129. DON vs DON+Asn, P < 0.0001. Malate: Ctrl vs DON, P = 0.0024. DON vs DON+Asn, P = 0.1399. Statistical significance was calculated using unpaired two-tailed Student’s t test (a, c-f) or one-way ANOVA followed by Dunnet’s multiple comparisons test (b). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, ns=not significant.
Extended Data Fig. 6 Determinations of ASNS expression levels using qPCR, immunohistochemistry, and immunoblotting.
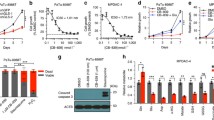
(a) Immunoblot assessing Asns protein levels in the indicated human PDAC cell lines treated with or without 10 µM DON for 24 h. Tubulin was used as a loading control. The results are representative of 2 independent experiments. Densitometry quantifications are presented relative to the −DON condition and values were normalized to the loading control. (b) Immunoblot assessing Asns protein expression in 779E cells treated with 0.5 mM DON with or without 0.1 mM Asn supplementation for 24 h. Tubulin was used as a loading control. (c) Relative ASNS mRNA expression levels as assessed by qPCR in 779E cells treated with 0.5 mM DON with or without 0.1 mM Asn supplementation for 24 h. Data are presented relative to Ctrl and are representative of 3 independent experiments. Data are presented as mean ± SEM of n = 3 replicates. For all statistical comparisons P < 0.0001. (d) Immunohistochemical staining of Asns protein in PaTu 8988 T orthotopic tumors treated with vehicle control (Ctrl) or DON (5 mg/kg). Representative images of n = 4 mice per group are shown. Scale bar, 100 µm. (e) Relative ASNS mRNA expression levels in KPC cells as assessed by qPCR after transfection with non-targeting negative control siRNA (siNC) or two different hairpins targeting ASNS (siASNS#1 and siASNS#2) for 24 h. Data are presented relative to siNC and are representative of 3 independent experiments. Data are presented as mean ± SEM of n = 3 replicates. For all statistical comparisons P < 0.0001. (f) Immunoblot assessing Asns protein levels in the indicated human PDAC cell lines treated with vehicle (Ctrl) or 10 µM DON for 24 h. Tubulin was used as a loading control. The results are representative of 2 independent experiments. Densitometry quantifications are presented relative to the values obtained for BxPC-3 cells and were normalized to the loading control. Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons test. ****P < 0.0001.
Extended Data Fig. 7 DON and ASNase co-treatment in 779E cells, glutamine quantification, and correlation analyses for DON target genes in PDOs.
(a, b) Relative number of 779E cells treated with the indicated doses of DON in combination with ASNase (0.5 U/ml; +ASNase) or without (−ASNase) for 24 h. Cells were stained with crystal violet and representative images are shown. Quantification of crystal violet staining is shown relative to −ASNase 0 μM DON condition and is representative of two independent experiments. The coefficient of drug interaction (CI) was calculated for each DON concentration (shown in graph). Data are presented as mean ± SEM of n = 3 replicate wells. 0.25 mM, P < 0.0001. 0.5 mM, P = 0.0004. 1 mM, P = 0.00017. (c) Determination of the glutamine concentration in cell culture media incubated with or without 0.5 U/mL ASNase for 24 hours. Data are presented as mean ± SEM of n = 3 replicate samples. (d–k) Correlation analysis between normalized gene expression of the indicated enzymes targeted by DON and DON viability ratio for the six PDOs assessed. Statistical significance was calculated using unpaired two-tailed Student’s t test (b, c) or two-tailed Spearman correlation coefficient (d–k). ***P < 0.001, ****P < 0.0001, ns = not significant.
Extended Data Fig. 8 Co-treatment with DON and ASNase does not further impact apoptosis or proliferation in tumors but does suppress liver metastases.
(a, b) Immunohistochemical staining of phospho-Histone H3 (pHis-H3; a) or cleaved Caspase 3 (CC3; b) in PaTu 8988 T orthotopic tumors treated with vehicle (Ctrl), ASNase (60U/mice), DON (5–10 mg/kg), or a combination of DON + ASNase for 3 weeks. Representative images are shown of n = 3 independent experiments. Scale bar 100 µm. (c) Hematoxylin and Eosin (H&E) staining of liver sections from PaTu 8988 T tumor-bearing mice treated with vehicle (Ctrl), DON (5–10 mg/kg), or a combination of DON + ASNase for 3 weeks. Metastases are indicated (M). Representative images are shown of n = 3 independent experiments. Scale bar 200 µm. (d) Determination of the glutamine plasma concentration in animals treated without or with ASNase (60U/mouse) for 3 weeks. Data are presented as mean ± SEM of n = 6 or n = 4 mice. Statistical significance was calculated using unpaired two-tailed Student’s t test. P = 0.2093. ns = not significant.
Extended Data Fig. 9 Suppression of metastasis by DON is enhanced by co-treatment with ASNase in a primary PDAC orthotopic mouse model.
(a) Tumor weights of 779E orthotopic tumors from animals treated with vehicle (Ctrl), ASNase (3U/g), DON (5 mg/kg), or combination of DON + ASNase for 2 weeks. Data are presented as mean ± SEM of n = 9, n = 7, n = 8, or n = 9 mice per group. Ctrl vs ASNase, P = 0.5836. Ctrl vs DON, P = 0.042. Ctrl vs DON+ASNase, P = 0.0491. (b) Characterization of macrometastases in animals with 779E orthotopic tumors treated as indicated. Representative images of macrometastases in the different tissues are shown. The white arrows point to macrometastases. The number of mice with macrometastases in each organ site was quantified as indicated in the table. n = 9, n = 7, n = 8, or n = 9 mice per group. (c) Quantification of the percentage of mice with 779E-derived primary orthotopic tumors that presented with macrometastases to any organ (all sites), the liver, intestines, or diaphragm. n = 9, n = 7, n = 8, or n = 9 mice per group. All Sites: Ctrl vs DON, P = 0.029; Ctrl vs DON+ASNase, P = 0.009. Liver: Ctrl vs DON, P = 0.0498; Ctrl vs DON+ASNase, P = 0.0034. Diaphragm: Ctrl vs DON, P = 0.0498; Ctrl vs DON+ASNase, P = 0.347. (d) Quantification of the metastatic tumors in the liver using image-based computation. The total area of tumor tissue per area of liver tissue was determined. Data are presented as Min to Max box-and-whisker plot with median indicated at center line for n = 16, n = 14, n = 12, or n = 18 images per condition from a total of 9, 7, 6 or 9 mice per group. Ctrl vs ASNase, P = 0.347. Ctrl vs DON, P = 0.016. Ctrl vs DON+ASNase, P < 0.0001. DON vs DON+ASNase, P = 0.013. Statistical significance was calculated using one-way ANOVA followed by Tukey’s multiple comparisons (a), two-tailed chi-square test (c), or unpaired two-tailed Student’s t test (d). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, ns = not significant.
Supplementary information
Source data
Figs. 1–6 and Extended Data Figs. 1–9
Statistical source data for graphs.
Fig. 4 and Extended Data Fig. 6
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Recouvreux, M.V., Grenier, S.F., Zhang, Y. et al. Glutamine mimicry suppresses tumor progression through asparagine metabolism in pancreatic ductal adenocarcinoma. Nat Cancer 5, 100–113 (2024). https://doi.org/10.1038/s43018-023-00649-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-023-00649-1
- Springer Nature America, Inc.
This article is cited by
-
Long-acting Erwinia chrysanthemi, Pegcrisantaspase, induces alternate amino acid biosynthetic pathways in a preclinical model of pancreatic ductal adenocarcinoma
Cancer & Metabolism (2024)
-
Emerging mechanisms and promising approaches in pancreatic cancer metabolism
Cell Death & Disease (2024)
-
Glutamine analogs for pancreatic cancer therapy
Nature Cancer (2024)
-
Mechanism insights and therapeutic intervention of tumor metastasis: latest developments and perspectives
Signal Transduction and Targeted Therapy (2024)
-
Oncometabolites in cancer: from cancer cells to the tumor microenvironment
Holistic Integrative Oncology (2024)