Abstract
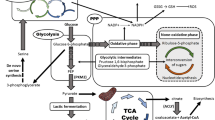
Pharmacological activation of the glycolytic enzyme PKM2 or expression of the constitutively active PKM1 isoform in cancer cells results in decreased lactate production, a phenomenon known as the PKM2 paradox in the Warburg effect. Here we show that oxaloacetate (OAA) is a competitive inhibitor of human lactate dehydrogenase A (LDHA) and that elevated PKM2 activity increases de novo synthesis of OAA through glutaminolysis, thereby inhibiting LDHA in cancer cells. We also show that replacement of human LDHA with rabbit LDHA, which is relatively resistant to OAA inhibition, eliminated the paradoxical correlation between the elevated PKM2 activity and the decreased lactate concentration in cancer cells treated with a PKM2 activator. Furthermore, rabbit LDHA-expressing tumours, compared to human LDHA-expressing tumours in mice, displayed resistance to the PKM2 activator. These findings describe a mechanistic explanation for the PKM2 paradox by showing that OAA accumulates and inhibits LDHA following PKM2 activation.







Similar content being viewed by others
Data availability
The accession codes used in this study are 6MV8 (hLDHA) and 5NQB (rbLDHA) from the PDB. Supplementary Information including Supplementary Fig. 1 exemplifying the gating strategy for Extended Data Fig. 4d are provided with this paper. Source data are provided with this paper.
References
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Lunt, S. Y. & Vander Heiden, M. G. Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu. Rev. Cell Dev. Biol. 27, 441–464 (2011).
Hitosugi, T. & Chen, J. Post-translational modifications and the Warburg effect. Oncogene 33, 4279–4285 (2014).
Dayton, T. L., Jacks, T. & Vander Heiden, M. G. PKM2, cancer metabolism, and the road ahead. EMBO Rep. 17, 1721–1730 (2016).
Wiese, E. K. & Hitosugi, T. Tyrosine kinase signaling in cancer metabolism: PKM2 paradox in the Warburg effect. Front Cell Dev. Biol. 6, 79 (2018).
Jurica, M. S. et al. The allosteric regulation of pyruvate kinase by fructose-1,6-bisphosphate. Structure 6, 195–210 (1998).
David, C. J., Chen, M., Assanah, M., Canoll, P. & Manley, J. L. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368 (2010).
Hitosugi, T. et al. Tyrosine phosphorylation inhibits PKM2 to promote the Warburg effect and tumor growth. Sci. Signal 2, ra73 (2009).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Anastasiou, D. et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science 334, 1278–1283 (2011).
Anastasiou, D. et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat. Chem. Biol. 8, 839–847 (2012).
Eigenbrodt, E. & Glossmann, H. Glycolysis—one of the keys to cancer? Trends Pharmacol. Sci. 1, 240–245 (1980).
Lv, L. et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol. Cell 42, 719–730 (2011).
Israelsen, W. J. et al. PKM2 isoform-specific deletion reveals a differential requirement for pyruvate kinase in tumor cells. Cell 155, 397–409 (2013).
Dayton, T. L. et al. Germline loss of PKM2 promotes metabolic distress and hepatocellular carcinoma. Genes Dev. 30, 1020–1033 (2016).
Wang, Y. H. et al. Cell-state-specific metabolic dependency in hematopoiesis and leukemogenesis. Cell 158, 1309–1323 (2014).
Dayton, T. L. et al. Isoform-specific deletion of PKM2 constrains tumor initiation in a mouse model of soft tissue sarcoma. Cancer Metab. 6, 6 (2018).
Kondoh, H. Cellular life span and the Warburg effect. Exp. Cell. Res. 314, 1923–1928 (2008).
Christofk, H. R., Vander Heiden, M. G., Wu, N., Asara, J. M. & Cantley, L. C. Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature 452, 181–186 (2008).
Miao, P., Sheng, S., Sun, X., Liu, J. & Huang, G. Lactate dehydrogenase A in cancer: a promising target for diagnosis and therapy. IUBMB Life 65, 904–910 (2013).
Shim, H. et al. c-Myc transactivation of LDH-A: implications for tumor metabolism and growth. Proc. Natl Acad. Sci. USA 94, 6658–6663 (1997).
Intlekofer, A. M. et al. Hypoxia induces production of L-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Alam, M. T. et al. The self-inhibitory nature of metabolic networks and its alleviation through compartmentalization. Nat. Commun. 8, 16018 (2017).
Yoshida, A. Enzymic properties of lactate dehydrogenase of Bacillus subtilis. Biochim. Biophys. Acta 99, 66–77 (1965).
Steinbuechel, A. & H., S. NAD-linked L(+)-lactate dehydrogenase from the strict aerobe Alcaligenes eutrophus. Eur. J. Biochem. 130, 329–334 (1983).
Fritz, P. J. Rabbit muscle lactate dehydrogenase 5: a regulatory enzyme. Science 150, 364–366 (1965).
Wiese, E. K. et al. Reductive amination of alpha-ketoglutarate in metabolite extracts results in glutamate overestimation. J. Chromatogr. A 1623, 461169 (2020).
Kurmi, K. et al. Carnitine palmitoyltransferase 1A has a lysine succinyltransferase activity. Cell Rep. 22, 1365–1373 (2018).
Zong, W. X., Rabinowitz, J. D. & White, E. Mitochondria and cancer. Mol. Cell 61, 667–676 (2016).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337 e1311 (2016).
Granchi, C., Paterni, I., Rani, R. & Minutolo, F. Small-molecule inhibitors of human LDH5. Future Med Chem. 5, 1967–1991 (2013).
Strambio-De-Castillia, C., Niepel, M. & Rout, M. P. The nuclear pore complex: bridging nuclear transport and gene regulation. Nat. Rev. Mol. Cell Biol. 11, 490–501 (2010).
Paine, P. L., Moore, L. C. & Horowitz, S. B. Nuclear envelope permeability. Nature 254, 109–114 (1975).
Lodish, H. et al. in Molecular Cell Biology 4th edn (ed. Tenney, S.) Ch. 5.4 (W.H. Freeman, 2000).
Rossignol, R. et al. Energy substrate modulates mitochondrial structure and oxidatie capacity in cancer cells. Cancer Res. 64, 985–993 (2004).
Wise, D. R. & Thompson, C. B. Glutamine addiction: a new therapeutic target in cancer. Trends Biochem. Sci. 35, 427–433 (2010).
Elia, I. et al. Breast cancer cells rely on environmental pyruvate to shape the metastatic niche. Nature 568, 117–121 (2019).
Smith, B. et al. Addiction to coupling of the Warburg effect with glutamine catabolism in cancer cells. Cell Rep. 17, 821–836 (2016).
Mazurek, S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int. J. Biochem. Cell Biol. 43, 969–980 (2011).
Chaneton, B. & Gottlieb, E. Rocking cell metabolism: revised functions of the key glycolytic regulator PKM2 in cancer. Trends Biochem. Sci. 37, 309–316 (2012).
Israelsen, W. J. & Vander Heiden, M. G. Pyruvate kinase: function, regulation and role in cancer. Semin. Cell Dev. Biol. 43, 43–51 (2015).
Kung, C. et al. Small molecule activation of PKM2 in cancer cells induces serine auxotrophy. Chem. Biol. 19, 1187–1198 (2012).
Chaneton, B. et al. Serine is a natural ligand and allosteric activator of pyruvate kinase M2. Nature 491, 458–462 (2012).
Gao, X., Wang, H., Yang, J. J., Liu, X. & Liu, Z. R. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol. Cell 45, 598–609 (2012).
Luo, W. et al. Pyruvate kinase M2 is a PHD3-stimulated coactivator for hypoxia-inducible factor 1. Cell 145, 732–744 (2011).
Wang, H. J. et al. JMJD5 regulates PKM2 nuclear translocation and reprograms HIF-1alpha-mediated glucose metabolism. Proc. Natl Acad. Sci. USA 111, 279–284 (2014).
Yang, W. et al. Nuclear PKM2 regulates beta-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Gruber, C. C. et al. An algorithm for the deconvolution of mass spectroscopic patterns in isotope labeling studies. evaluation for the hydrogen-deuterium exchange reaction in ketones. J. Org. Chem. 72, 5778–5783 (2007).
Mazlaghaninia, M., Atri, M. S. & Seyedalipour, B. Scopoletin and morin inhibit lactate dehydrogenase enzyme activity is critical for cancer metabolism. Hormozgan Med. J. 23, e88269 (2019).
Pang, Y. P. FF12MC: a revised AMBER forcefield and new protein simulation protocol. Proteins 84, 1490–1516 (2016).
Hitosugi, T. et al. Phosphoglycerate mutase 1 coordinates glycolysis and biosynthesis to promote tumor growth. Cancer Cell 22, 585–600 (2012).
Kurmi, K. et al. Tyrosine phosphorylation of mitochondrial creatine kinase 1 enhances a druggable tumor energy shuttle pathway. Cell Metab. 28, https://doi.org/10.1016/j.cmet.2018.08.008 (2018).
Cooper, G., Reed, C., Nguyen, D., Carter, M. & Wang, Y. Detection and formation scenario of citric acid, pyruvic acid, and other possible metabolism precursors in carbonaceous meteorites. Proc. Natl Acad. Sci. USA 108, 14015–14020 (2011).
Gonsalves, W. I. et al. Glutamine-derived 2-hydroxyglutarate is associated with disease progression in plasma cell malignancies. JCI Insight 3, https://doi.org/10.1172/jci.insight.94543 (2018).
Hua, Y., Laserstein, P. & Helmstaedter, M. Large-volume en-bloc staining for electron microscopy-based connectomics. Nat. Commun. 6, 7923 (2015).
Mereuta, O. M. et al. High-resolution scanning electron microscopy for the analysis of three-dimensional ultrastructure of clots in acute ischemic stroke. J. NeuroIntervent Surg. 0, 1–7 (2020).
Hurley, R. M. et al. 53BP1 as a potential predictor of response in PARP inhibitor-treated homologous recombination-deficient ovarian cancer. Gynecol. Oncol. 153, 127–134 (2019).
Acknowledgements
We thank the Mayo Microscopy and Cell Analysis Core and the Mayo Pathology Research Core at Mayo Clinic Rochester for experimental and technical support. We thank J. Maher and S. Kaufmann for their critical reading of the paper. This research was supported in part by National Institutes of Health (NIH) grant no. R01 CA225680 (T.H.), Research Scholar grant (no. RSG-19-076-01-TBE) from the American Cancer Society (T.H.), the Eagles Cancer Research Fund (T.H.), a Team Science Platform Award from the Mayo Clinic Center for Biomedical Discovery (T.H.), the Developmental Therapeutics Program from the Mayo Clinic Cancer Center (T.H.) and the Mayo Clinic Breast SPORE grant no. P50 CA116201 (T.H.). W.I.G. was supported by the National Cancer Institute of the NIH under award no. K23 CA218742. S.T.L. was supported by NIH grant no. R25 GM075148-14. E.K.W. was supported by NIH grant no. T32 GM072474 and a predoctoral fellowship from the Mayo Foundation for Education and Research.
Author information
Authors and Affiliations
Contributions
E.K.W., S.H., S.T.L., A.S. and T.H. performed the experiments and analysed the data with input from K.K. and L.M.K. L.G.A.-B. assisted with SBFSEM data analysis. Y.-P.P. performed the computational work. W.I.G. provided critical reagents and patient samples. E.K.W. and T.H. wrote the paper. All authors contributed to discussion about the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Chi Dang, Heather Christofk and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Caputa.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Increased PKM2 activity decreases LDH activity and lactate concentration. Related to Fig. 1.
a. Pyruvate kinase activity from lysates of PKM2 or PKM1-expressing H1299 cells. Corresponding western blots of H1299 vector, PKM2, and PKM1 cell lysates with antibodies against PKM, PKM2, PKM1, LDHA, and β-actin. b. Whole cell lactate concentrations of PKM2 and PKM1 expressing H1299 cells. c. Relative lactate production rate in PKM2 and PKM1 expressing H1299 cells. d. Relative lactate production rate in RPMI 8226 cells treated with vehicle or DASA. e. Pyruvate concentrations in PKM2 or PKM1 expressing H1299 cells. f-g. Relative glucose uptake of H1299 (f) and RPMI 8226 (g) cells with vehicle or DASA treatment. h. 13C lactate labelling rate of RPMI 8226 cells incubated with 13C6 glucose and vehicle or 40 μM DASA. i. 13C pyruvate labeling rate of RPMI 8226 cells incubated with 13C6 glucose and vehicle or 40 μM DASA. j. In situ LDH activity in RPMI 8226 cells with vehicle or DASA treatment. k. 13C lactate labeling rate of PKM2 and PKM1 expressing H1299 cells incubated with 13C6 glucose. l. 13C pyruvate labeling rate of PKM2 and PKM1 expressing H1299 cells incubated with 13C6 glucose. m. In situ LDH activity in PKM2 and PKM1 expressing H1299 cells. n. LDH activity from lysates of RPMI 8226 cells treated with 40 μM DASA or vehicle overnight. Below, the corresponding western blots of the RPMI 8226 lysates with antibodies against LDHA, LDHB, and β-actin. o. LDH activity from lysates of H1299 cells expressing PKM2 or PKM1. Below, the corresponding western blots of the PKM2 and PKM1 expressing H1299 lysates with antibodies against LDHA, LDHB, and β-actin. Data is represented as the mean and error bars represent the standard deviation from n = 3 for (a), (k), (l), (m), n = 4 for (b), (c), (d), (e), (f), (g), (h), (i), (j), n = 6 for (n), n = 7 for (o) of biologically independent replicates. Western blot results in (a), (n), (o) are representative experiments of 3 biologically independent replicates. P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 2 OAA is a competitive inhibitor of hLDHA. Related to Fig. 2.
a. Gas chromatogram of malate 3-TMS derivative (m/z 335) of pure malate standard (top). Gas chromatogram of malate 3-TMS derivative (m/z 335) of an in vitro LDH activity assay sample in which purified recombinant hLDHA was incubated with OAA (bottom). No malate was detected. b. Relative LDHA activity in the presence of various metabolites, determined using purified recombinant human LDHA and a fluorescence based LDH activity assay. c. Fluorescence intensity scan at ex. 280 nm of purified human LDHA incubated with increasing concentrations of OAA. Data is represented as the mean and error bars represent the standard deviation from n = 4 for (b) of independent replicates in vitro. Chromatograms in (a) and fluorescence curves in (c) are representative experiments of three independent replicates in vitro and the P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 3 GC–MS detection of OAA 3-TBDMS and analysis of glucose uptake.
a. (left) Scan of m/z 417 of pure 12C OAA standard derivatized with DMF-MTBSTFA. (middle) Mass spectrum of pure 12C OAA standard, derivatized with DMF-MTBSTFA. m/z 417 corresponds to OAA 3-TBDMS M-57 fragment. (right) Standard curve showing a positive, linear relationship between OAA amount and m/z 417 peak intensity of pure OAA standard. b. (left) Scan of m/z 417 of H1299 cellular metabolites labelled with 13C glucose, extracted with 1:1 methanol:water containing 5 mM ninhydrin, with or without 12C OAA spike. (middle) Isotopomer distribution of OAA (m/z 417) from H1299 cellular metabolites labeled with 13C glucose, extracted with 1:1 methanol:water containing 5 mM ninhydrin, with or without 12C OAA spike. (right) Ion abundance, normalized using IsoPat2, of isotopomer distribution data from (middle). c. Relative glucose uptake of H1299 (left) and RPMI 8226 (right) cells treated with vehicle or 1 mM diethyl-ester OAA. d. Relative glucose uptake of H1299 (left) and RPMI 8226 (right) cells treated with vehicle or 10 mM aspartate. Data is represented as the mean and error bars represent the standard deviation from n = 4 for (c), (d) of biologically independent replicates. Results in (a) are representative experiments of three independent replicates in vitro. Results in (b) are representative experiments of 3 biologically independent replicates. P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 4 Activation of PKM2 increases cellular OAA concentrations to inhibit LDH activity. Related to Fig. 4.
a. Relative whole cell OAA concentrations of RPMI 8226 cells treated with 40 μM DASA or vehicle overnight. Whole cell metabolites were extracted with 1:1 methanol:water containing 5 mM ninhydrin, with or without 12C OAA spike, derivatized, and analysed with GC-MS. b. Relative whole cell OAA concentrations of PKM2 and PKM1 expressing H1299 cells. c. Schematic of fractionation method. d. Relative mean signal intensity, determined by flow cytometry, of (left) ER-BODIPY red, (middle) Golgi tracker red, and (right) Mito tracker red stained H1299 cells, treated with 40 μM DASA or vehicle. e. Representative serial block face scanning electron microscopy (SBFSEM) image of an H1299 cell. Small organelles included in the small organelle fraction are colored blue. Representative golgi, endoplasmic reticulum (ER), and mitochondria are circled in white. Image is one slice from one representative cell. f. Whole cell volumes as well as fractionated volumes of 12 cells analysed using SBFSEM. g. Western blot of H1299 lysate and increasing concentrations of purified recombinant LDHA. Corresponding standard curve constructed from the average pixel density of each LDHA purified recombinant protein band and used to determine the amount of LDHA protein in H1299 lysate. h. In vitro LDHA activity assay, using purified recombinant LDHA and pyruvate and cytosolic OAA concentrations determined under 40 μM DASA or vehicle treatment in H1299 cells. LDHA activity was determined by analysing 13C1 lactate produced from 13C1 pyruvate by GC-MS. Western blot results are representative experiments of three independent replicates. Data is represented as the mean and error bars represent the standard deviation from n = 4 for (a), (b), (h), n = 3 for (d) of biologically independent replicates. Micrograph in (e) is a representative experiment of 12 biologically independent replicates. Standard curve in (g) is a representative experiment of 3 independent replicates in vitro. The western blot in (g) is a representative experiment of 3 biologically independent replicates. P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 5 PKM2 activity increases glutamine derived OAA. Related to Fig. 5.
a. Isotopomer analysis of 13C6 glucose-derived carbon incorporation to OAA in RPMI 8226 cells treated with 13C6 glucose and 40 μM DASA or vehicle overnight. b. Isotopomer analysis of 13C5 glutamine-derived carbon incorporation to OAA in RPMI 8226 cells treated with 13C5 glutamine and 40 μM DASA or vehicle overnight. c. Western blot analysis with antibodies against GPT2 and β-actin of H1299 lysates from cells treated with vehicle or 40 μM DASA overnight. d. Relative OAA concentrations in H1299 GPT2 knockdown cells treated with vehicle or 40 μM DASA overnight. e. Western blot analysis of H1299 vector and PHGDH knockdown cell lysates with antibodies against PHGDH and β-actin. f. 13C lactate labeling rate of H1299 vector and PHGDH knockdown cells incubated with 13C6 glucose and vehicle or 40 μM DASA. g. 13C pyruvate labeling rate of H1299 vector and PHGDH knockdown cells incubated with 13C6 glucose and vehicle or 40 μM DASA. h. In situ LDH activity in H1299 vector and PHGDH knockdown cells treated with vehicle or 40 μM DASA. i. Western blot analysis of H1299 vector and PC knockdown cell lysates with antibodies against PC and β-actin. j. 13C lactate labeling rate of H1299 vector and PC knockdown cells incubated with 13C6 glucose and vehicle or 40 μM DASA. k. 13C pyruvate labeling rate of H1299 vector and PC knockdown cells incubated with 13C6 glucose and vehicle or 40 μM DASA. l. In situ LDH activity in H1299 vector and PC knockdown cells treated with vehicle or 40 μM DASA. Data is represented as the mean and error bars represent the standard deviation from n = 4 for (a), (d), (f), (g), (h), (j), (k), (l), n = 3 for (b) of biologically independent replicates. Western blot results in (c), (e), (i) are representative experiments of 3 biologically independent replicates. P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 6 LDHB has a small effect on the total LDH activity in H1299 cells. Related to Fig. 6.
a. The IC50 of OAA against human LDHB determined using purified recombinant human LDHB and analysis of 13C1 lactate derived from 13C1 pyruvate. The IC50 was identified as 1,021 ± 86 μM. b. Western blot of H1299 lysate and increasing concentrations of purified recombinant LDHB. Corresponding standard curve constructed from the average pixel density of LDHB purified recombinant protein band and used to determine the amount of LDHB protein in H1299 lysate. c. LDH activity, determined using a fluorescence based LDH activity assay using diluted lysates from H1299 vector and LDHB knockdown cells. The corresponding western blots of the H1299 lysates used in the above activity assay with antibodies against LDHB and β-actin. Data is represented as the mean and error bars represent the standard deviation from n = 4 for (a), n = 3 for (c) of independent replicates in vitro. Standard curve in (b) is a representative experiment of 3 independent replicates in vitro. Western blots in (b) (c) are representative experiments of 3 biologically independent replicates. P values were determined by a two-tailed Student’s t-test.
Extended Data Fig. 7 OAA inhibition of LDHA increases the response to TEPP-46 in vivo.
a. Pyruvate kinase enzyme activity assay in lysates from H1299 cells treated with vehicle or 40 μM TEPP-46 overnight. The corresponding western blots of H1299 cell lysates with antibodies against PKM2 and β-actin. b. Isotopomer analysis of 13C5 glutamine-derived carbon incorporation to OAA in H1299 cells treated with 13C5 glutamine and 40 μM TEPP-46 or vehicle overnight. c. Ki67 staining of hLDHA- and rbLDHA-expressing H1299 tumours from mice treated with vehicle or TEPP-46. Ki67 staining results are representative experiments of 3 biologically independent replicates. Western blot results are representative experiments of 3 independent replicates. Data is represented as the mean and error bars represent the standard deviation from n = 3 for (a), (b) of biologically independent measurements. P values were determined by a two-tailed Student’s t-test.
Supplementary information
Supplementary Information
Supplementary Fig. 1
Source data
Source Data Fig. 1
Uncropped western blots for Fig. 1.
Source Data Fig. 3
Uncropped western blots for Fig. 3.
Source Data Fig. 4
Uncropped western blots for Fig. 4.
Source Data Fig. 5
Uncropped western blots for Fig. 5.
Source Data Fig. 6
Uncropped western blots for Fig. 6.
Source Data Fig. 7
Raw data for tumour measurements in Fig. 7a.
Source Data Extended Data Fig. 1
Uncropped western blots for Extended Data Fig. 1.
Source Data Extended Data Fig. 4
Uncropped western blots for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Uncropped western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Uncropped western blots for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Uncropped western blots for Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Wiese, E.K., Hitosugi, S., Loa, S.T. et al. Enzymatic activation of pyruvate kinase increases cytosolic oxaloacetate to inhibit the Warburg effect. Nat Metab 3, 954–968 (2021). https://doi.org/10.1038/s42255-021-00424-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00424-5
- Springer Nature Limited
This article is cited by
-
Chlorpromazine affects glioblastoma bioenergetics by interfering with pyruvate kinase M2
Cell Death & Disease (2023)
-
A novel protein encoded by ZCRB1-induced circHEATR5B suppresses aerobic glycolysis of GBM through phosphorylation of JMJD5
Journal of Experimental & Clinical Cancer Research (2022)