Abstract
Probiotics have been characterized as useful for maintaining the balance of host gut flora and conferring health effects, but few studies have focused on their potential for delaying aging in the host. Here we show that Lacticaseibacillus rhamnosus Probio-M9 (Probio-M9), a healthy breast milk probiotic, enhances the locomotor ability and slows the decline in muscle function of the model organism Caenorhabditis elegans. Live Probio-M9 significantly extends the lifespan of C. elegans in a dietary restriction-independent manner. By screening various aging-related mutants of C. elegans, we find that Probio-M9 extends lifespan via p38 cascade and daf-2 signaling pathways, independent on daf-16 but dependent on skn-1. Probio-M9 protects and repairs damaged mitochondria by activating mitochondrial unfolded protein response. The significant increase of amino acids, sphingolipid, galactose and fatty acids in bacterial metabolites might be involved in extending the lifespan of C. elegans. We reveal that Probio-M9 as a dietary supplementation had the potential to delay aging in C. elegans and also provide new methods and insights for further analyzing probiotics in improving host health and delaying the occurrence of age-related chronic diseases.
Similar content being viewed by others

Introduction
The morbidity and mortality of cardiovascular diseases, neurodegenerative diseases, cancer and other illnesses, are significantly and positively correlated with age1. Hence, efforts to extend the lifespan and delay the aging process have received wide attention. Many studies have focused on the key effects of specific diets and nutrient resources in anti-aging, particularly probiotics supplementation2,3,4,5. Living microorganism probiotics have functions that are beneficial to human health when consumed in sufficient amounts6. It has been demonstrated that probiotics confer health-promoting benefits to their host by improving the intestinal microbial balance7,8,9, enhancing immune modulation10,11, and/or competing with pathogens12. However, the mechanisms by which probiotics extend host lifespan remain largely elusive.
The nematode Caenorhabditis elegans is a widely used and successful model organism in anti-aging studies because of its obvious advantages, including its short and easily monitored lifespan13,14, and the absence of ethical issues15. Genes that modulate aging processes in C. elegans include some of the classical and conserved signaling pathways also found in vertebrates. The p38 mitogen-activated protein kinase (MAPK) pathway is one of the most ancient evolutionarily conserved pathways in C. elegans16. In human, the p38 cascade is typically activated by inflammatory cytokines and pathogen invasion17,18. In C. elegans, the p38 cascade is activated by NSY-1 MAPK kinase, SEK-1 MAPK kinase and PMK-115,16. Some conserved components of the insulin/insulin-like growth factor 1 (IGF-1) signaling (IIS) pathway also regulate aging. The sole insulin/IGF-1 receptor encoded by the gerontogene daf-219 is a key upstream component of IIS, regulating various physiological processes, such as aging and adult lifespan in C. elegans20,21. The lifespan extending effects of daf-2 are mediated via the daf-16/FOXO transcription factor in reduced insulin signaling mutants such as daf-2 (e1370)20. SKN-1, a stress-responsive nuclear transcription factor, contributes to the reduction of IIS-associated longevity22. Additionally, certain substances secreted by bacterial metabolites, such as colanic acid, agmatine and methylglyoxal have been identified as key factors that have great impacts on longevity in their hosts2,23,24.
Mitochondrial function has a far-reaching effect on the process of aging25. It has been shown that mitochondrial dysfunction has beneficial effects on the lifespan of C. elegans26,27. Mutations that cause electron transport chain (ETC) dysfunction extend the worm lifespan by as much as 50%27. The mitochondrial unfolded protein response (UPRmt) is also associated with health-promoting and mitochondrial homeostasis28. UPRmt is activated when the transcription factor ATFS-1 shuttles from mitochondria to the nucleus in response to mitochondrial stress29.
In this study, we explored the effects of the probiotic Lacticaseibacillus rhamnosus Probio-M9, which is isolated from healthy breast milk, on physiological functions including lifespan, locomotor ability and lipofuscin accumulation in C. elegans. Feeding conditioned with Probio-M9 significantly extended the worm lifespan. Using various aging-related mutants of C. elegans involving signal transduction, including mutations in the p38 MAPK, nutrient-sensing signaling pathways and UPRmt, we investigated the molecular mechanism of lifespan extension by Probio-M9. Combined with the differential expression of metabolites by Probio-M9, our study revealed the possible metabolic pathway of Probio-M9 to delay host aging. These data reveal the vital link between probiotics and host longevity, provide new insights for the continued development of probiotics, and indicate that dietary supplementation with probiotics might have the potential to delay host aging.
Results
Probio-M9 extends the lifespan of C. elegans
Probio-M9 and E. coli OP50 belong to different bacterial genera, it has been reported that nematodes exhibited a preference when the normal food Escherichia coli OP50 (E. coli OP50) is replaced with other bacteria30,31. Therefore, Probio-M9 precipitate was resuspended in E. coli OP50 solution to exclude the preference of worms. In this study, we used an approximately 1:4 ratio of E. coli OP50 to Probio-M9 (viable plate count: 6.8 × 108 CFU/mL and 2.82 × 109 CFU/mL, respectively) as the experimental group (OP50 + Probio-M9), and E. coli OP50 alone as the control group (OP50). First, we used choice assays to explore whether the worms had a preference for OP50 over OP50 + Probio-M9. Worm eggs were cultured to late L4 stage on OP50 (Supplementary Fig. 1a) or OP50 + Probio-M9 (Supplementary Fig. 1c), then observed following transfer to OP50 or OP50 + Probio-M9 for 1 h or 2 h. The results showed that the number of worms on each plate was similar (Supplementary Fig. 1b, d), suggesting that C. elegans did not display a preference between OP50 and OP50 + Probio-M9. We also performed a binary selection analysis. Worm eggs were cultured to late L4 stage on OP50 (Supplementary Fig. 1e) or OP50 + Probio-M9 (Supplementary Fig. 1g), then transferred to plate with two bacterial lawns with OP50 and OP50 + Probio-M9. Similarly, there was no significant difference in the number of worms in each lawn after 1 h or 2 h feeding (Supplementary Fig. 1f, h). These results indicated that the worms did not exhibit any avoidance behavior or preference regarding OP50 + Probio-M9. Taken together, these results indicated that Probio-M9 is a suitable dietary supplementation for C. elegans.
Next, we evaluated the effect of Probio-M9 on the lifespan of C. elegans. Synchronized late L4 stage worms were shifted from OP50 to OP50 + Probio-M9 or continued to culture on OP50, and survival was measured daily until all of the worms had died. The results showed that OP50 + Probio-M9 significantly extended worm lifespan by about 30% (Fig. 1a and Supplementary Table 1). When the proportion of Probio-M9 in the bacterial mixture was changed (either diluted or increased), extension of lifespan was still observed (Fig. 1b and Supplementary Table 1), but was optimal in the 1:4 ratio of OP50: Probio-M9 (OP50 + Probio-M9), suggesting that the beneficial effect of Probio-M9 is dose dependent.
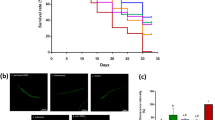
a Probio-M9 precipitate was resuspended in OP50 significantly extended the lifespan of wild-type N2 worms (p < 0.0001, Log rank test). b The most suitable dose of Probio-M9 (OP50 + 4 Probio-M9) significantly extended the lifespan of wild-type N2 worms (p < 0.0001, Log rank test). The lower (OP50 + 0.04 Probio-M9 and OP50 + 0.4 Probio-M9) or higher (OP50 + 40 Probio-M9) amount of Probio-M9 exhibits less extension on lifespan (p < 0.05, Log rank test). In a and b, lifespans were determined at least in three biologically independent experiments, and 90 worms of OP50 or OP50 + Probio-M9 feeding were tested in the single experiment, result from a single representative experiment is shown. The amount of OP50 and Probio-M9 which were used is 6.8 × 108 CFU/mL and 2.82 × 109 CFU/mL, respectively.
Probio-M9 exhibits health-promoting effects in C. elegans
Dietary restriction (DR) is well known for delaying the development of C. elegans, thereby prolonging its lifespan32. To investigate the effects of Probio-M9 feeding on the development of C. elegans, we examined life cycle, brood size and body size. Culturing on OP50 + Probio-M9 did not affect the time required to reach reproductive age, nor did it affect the reproductive capacity, or cycle of the worms (Fig. 2a–c). Body size of worms was also unaffected by OP50 + Probio-M9 (Fig. 2d). In addition, we investigated the effect of DR signaling pathway in lifespan extension of C. elegans. We found that OP50 + Probio-M9 still extended the lifespans of eat-2 (ad1116) and aak-2 (ok524) mutant worms (Fig. 2e, f and Supplementary Table 2). Together, these data showed that feeding worms with Probio-M9 had no effect on their growth and development, indicating that Probio-M9 extends the lifespan of C. elegans in a DR-independent manner.
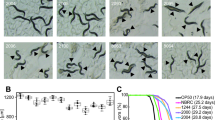
The life cycle (from egg to its first egg laid) (a), total number of progeny (b), reproductive cycle (c) and body size (d) from wild-type N2 worms grown on OP50 and OP50 + Probio-M9 are not significantly different (N = 20 worms, p > 0.05, Student’s t test). Survival curves of mutants of DR signaling pathways, eat-2 (ad1116) (e) and aak-2 (ok524) (f), fed with Probio-M9 (N = 90 worms, p < 0.0001, Log rank test). g The pharyngeal pumping rate from wild-type N2 worms grown on OP50 + Probio-M9 was enhanced on days 2–14 compared with those fed with OP50 (N = 20 worms, p < 0.01, two-way ANOVA). h The normal locomotor activity from wild-type N2 worms grown on OP50 and OP50 + Probio-M9 was measured on days 8–16. According to their locomotion, worms were divided into four classes: Class A, normal locomotion, spontaneous and/or rhythmical sinusoidal movement (blue bars); Class B, cannot move body, irregular and/or uncoordinated movement (orange bars); Class C, uncoordinated/sluggish, moved merely their head and/or tail in respond to a moderate touch with a platinum wire picker (green bars); and Class D, dead, dead worms (red bars) (N ≥ 90 worms, p < 0.05, Chi-squared test). i, j The lipofuscin accumulation of wild-type N2 worms was weakened followed OP50 + Probio-M9 feeding. Accumulation of lipofuscin was detected by autofluorescence under FV1200 confocal microscope on day 10 or 15 worms, scale bar, 20 μm (i), and fluorescence intensity was measured using ImageJ software (j). Results are shown by arbitrary units (a.u.) (N = 34 worms, p < 0.001, Student’s t test). k The thermotolerance of wild-type N2 worms grown on OP50 + Probio-M9 increased the survival rate compared those feds with OP50. Day 5 of adulthood worms cultured on OP50 or OP50 + Probio-M9 were transferred from 20 °C to 34 °C, and the survival rate was counted at 12 h, 18 h and 24 h, respectively (N = 3 biologically experiments, p < 0.05, Student’s t test). In a–d, g, h, j and k, values are presented as the mean ± SEM.
In C. elegans, the decline in muscle function and accumulation of lipofuscin are closely correlated with aging. To investigate whether feeding with Probio-M9 might improve these physiological functions, we measured three classical phenotypes associated with aging. The pharyngeal pumping rate declined progressively with aging in both groups of worms, and feeding with OP50 + Probio-M9 had no significant difference in pharyngeal pumping rate on days 2 and 4 of young adulthood, but significantly delayed the decrease in pharyngeal pumping rate on days 6–14 of adulthood (Fig. 2g). Consistent with the changes in pharyngeal pumping rate, the decline in normal locomotor ability of the worms was also delayed by OP50 + Probio-M9 feeding (Fig. 2h). Furthermore, we found that the accumulation of lipofuscin, an autofluorescent compound that accumulates in aging cells, was significantly reduced in worms fed with OP50 + Probio-M9 compared with those fed with OP50 on days 10 and 15 of adulthood (Fig. 2i, j). These findings indicated that Probio-M9 feeding supports the maintenance of higher muscle mass and enhances physical function in older worms.
Because longevity and stress resistance are interrelated, we also examined the effect of Probio-M9 on resistance to physical (temperature) stress in C. elegans. At day 5 of adulthood, worms fed with OP50 + Probio-M9 displayed significantly higher survival rates, than those fed with OP50, following 18 or 24 h of exposure to 34 °C heat (Fig. 2k). These results suggested that Probio-M9 has a positive effect on the thermostability of C. elegans. Altogether, our data indicated that Probio-M9 feeding improves the health of worms and extends their lifespan.
The health-promoting effects of Probio-M9 depends on its adhesion to the host gut
Several different standards have been used to screen potential bacterial strains for probiotics usefulness. The adhesion of probiotics to intestinal epithelial cells, and the intestinal accumulation of probiotics are vital prerequisites for maintaining homeostasis in the host gut microbiota33. To evaluate the effects of Probio-M9 on the intestinal health of the C. elegans, we examined the effects of feeding intervention with OP50 + Probio-M9 at specific developmental stages on the lifespan of worms, and tested the adhesion of Probio-M9 in the worm intestine. Worms derived from eggs that were cultured to either the L4 stage or adulthood (day 2 or day 4 after L4) on OP50 + Probio-M9 exhibited a significantly longer lifespan extension compared with those fed with OP50 (Fig. 3a–e and Supplementary Table 3). We also found that worms cultured from eggs to adulthood (day 4 after L4) on OP50 + Probio-M9 had a significantly longer lifespan extension than those cultured from eggs to the L4 stage on OP50 + Probio-M9 (Fig. 3f and Supplementary Table 3). These results indicated that the extent of lifespan extension is dependent on timing of on Probio-M9 feeding in the developmental cycle of C. elegans.
a, c Schematics illustrate the method and bacteria used in each assay. After bleaching, eggs were transferred to bacterial lawn (OP50 or OP50 + Probio-M9) and allowed to develop to late L4 stage or adulthood (day 2 or day 4 after L4 stage). b, d, e Survival curves of OP50 or OP50 + Probio-M9-fed to late L4 stage (b), day 2 after L4 stage (d) and day 4 after L4 stage (e) (N = 90 worms, p < 0.05, Log rank test). f OP50 + Probio-M9 feeding at day 4 after L4 stage significantly lengthened lifespan, but not at day 2 after L4 stage, compared with those fed with OP50 + Probio-M9 to L4 stage (N = 3 biologically experiments, p < 0.05, Student’s t test). g Colony forming indicated that Probio-M9 was attached to the C. elegans gut similar to LGG (N = 3 biologically experiments, p > 0.05, Student’s t test). Day 5 or day 10 worms were collected from the NGM plates lawned with the test bacterial and processed to count the number of bacterial CFU, expressed as CFU per worm. OP50 and LGG were used as control. h Accumulation of the C. elegans gut by Probio-M9. Worms were fed with Probio-M9 or control (OP50 and LGG) for 10 days, and then transferred them to new plates lawn with OP50. After 3 days, the number of CFU in the worm gut was calculated, expressed as CFU per worm, and the results indicated that Probio-M9 was still attached to C. elegans gut (N = 3 biologically experiments, p > 0.05, Student’s t test). In f–h, values are presented as the mean ± SEM. i Heat-inactivated Probio-M9 (95 °C for 30 min) failed to extend the lifespan of wild-type N2 worms (N = 90 worms, p > 0.05, Log rank test).
To verify the presence of Probio-M9 in the intestine of C. elegans, we assessed the ability of Probio-M9 to adhere to the worm intestinal tract on days 5 and 10 of adulthood. After exposure to OP50 + Probio-M9 for 5 or 10 days, the total number of colonies forming units (CFU) was used to determine the number of bacteria accumulated in the intestine. The number of Probio-M9 on days 5 and 10 (217 CFU/mL/worm and 793 CFU/mL/worm, respectively), were comparable to those in the positive control Lacticaseibacillus rhamnosus GG (LGG, 166 CFU/mL/worm and 872 CFU/mL/worm, respectively) (Fig. 3g). To confirm that Probio-M9 remained in the intestine, day 10 worms fed with OP50 + Probio-M9 or OP50 + LGG were transferred to OP50 for 3 days. The total number of Probio-M9 colonies was 667 CFU/mL/worm in those previously fed with OP50 + Probio-M9, similar to the 747 CFU/mL/worm in the LGG group (Fig. 3h). These results demonstrated that Probio-M9 is maintained in the intestinal tract of C. elegans and contributes to the healthspan of the host.
In addition, we examined the impact of feeding with heat-inactivated Probio-M9 (95 °C for 30 min) on the lifespan of C. elegans. We found that OP50 + heat-inactivated Probio-M9 was unable to extend the lifespan of worms (Fig. 3i and Supplementary Table 3), suggesting that Probio-M9 must be in the live state to exert beneficial effects on the host’s healthspan. Taken together, these results indicated that the health-promoting effects of Probio-M9 involve adherence to the intestinal tract of C. elegans.
Probio-M9 acts through the p38 MAPK signaling pathway
To understand the molecular mechanisms of longevity associated with Probio-M9 feeding, we explored the p38 signaling pathway, a conserved MAPK subfamily signaling pathway that plays a pivotal role in host lifespan regulation16. Using C. elegans mutants, we first investigated whether NSY-1, SEK-1 and PMK-1, three key factors in the p38 signaling pathway, were associated with host longevity induced by Probio-M9 feeding. Interestingly, we found that nsy-1 (ag3) and pmk-1 (km25) mutation almost completely suppressed the lifespan extension associated with Probio-M9 feeding, sek-1 (km4) mutation strongly but incompletely suppressed the longevity (~30% lifespan extension in N2 vs ~12% lifespan extension in the sek-1 mutant) (Fig. 4a–c and Supplementary Table 4), suggesting that the C. elegans p38 cascade is involved in the lifespan extension mediated by Probio-M9.
To further verify the involvement of the p38 signaling pathway, we evaluated the effect of Probio-M9 on the lifespan of two mutants of the Toll-interleukin-1 resistance (TIR-1) domain protein tir-1 (tm3036) and tir-1 (ok1052), which functions upstream of PMK-1 in C. elegans. Similarly, OP50 + Probio-M9 was unable to extend the lifespan of either mutant (Fig. 4d, e and Supplementary Table 4). As further confirmation, we explored the effect of Probio-M9 on ATF-7, a transcription factor that acts downstream of PMK-1 in the p38 signaling pathway. Again, OP50 + Probio-M9 was unable to extend the lifespan of atf-7 (gk715) mutant worms (Fig. 4f and Supplementary Table 4), suggesting that Probio-M9 might activate ATF-7 through PMK-1 to extend the lifespan of C. elegans. Altogether, our data indicated that the effects of Probio-M9 on lifespan extension in C. elegans require the p38 signaling pathway.
Probio-M9 downregulates the insulin-like signaling pathway
The interactions between host and probiotic in the promotion of longevity are complex and multifactorial34. We wondered whether Probio-M9 might work through one of the well-known nutrient-sensing mechanisms in the host. Using C. elegans with loss-of-function mutants in the IIS and target of rapamycin (TOR) signaling pathways, we investigated whether these host factors were associated with host longevity through Probio-M9 feeding. Of the four mutants tested, only the daf-2 (e1370) mutant failed to exhibit an extended lifespan in response to feeding with OP50 + Probio-M9 (Fig. 5a, b, Supplementary Fig. 2a, b and Supplementary Tables 5, 6). This result suggested that the mechanism of Probio-M9 lifespan extension involves the IIS signaling pathway, but not the TOR signaling pathway in C. elegans.
Lifespans of insulin-like signaling pathway mutants, daf-2 (e1370) (a), age-1 (hx546) (b), daf-16 (mgDf50) (c), skn-1 (mg570) (d) and hsf-1 (sy441) (e) fed with Probio-M9 (N = 90 worms, p > 0.05, p < 0.0001, Log rank test). The stress response reporters skn-1::GFP (f) and hsp-16.2p::GFP (h) were used to measure SKN-1 and HSF-1, respectively. Day 5 worms were heated for 20 min at 34 °C, which induced hsp-16.2p::GFP reporter expression. g, i Fluorescence intensity of GFP was quantified using ImageJ software. The skn-1::GFP (g) worms grown on OP50 + Probio-M9 showed increased fluorescence intensity in the intestine (N = 30 worms, p < 0.01, Student’s t test). Probio-M9 had no influence on HSF-1 (i) responding stress induction (N = 30 worms, p > 0.05, Student’s t test). j, k The expression of GST-4, an indicator of SKN-1 activity, increased after OP50 + Probio-M9 feeding. Expression of gst-4p::gfp was detected on day 5 worms (j), and fluorescence intensity was measured using ImageJ software (k) (N = 30 worms, p < 0.01, Student’s t test). In f, h and j, scale bar, 20 μm. In g, i and k, results are shown by arbitrary units (a.u.), values are presented as the mean ± SEM.
DAF-2 is an upstream component of IIS and regulates a variety of physiological processes35. To determine how Probio-M9 interacts with the IIS signaling pathway in the regulation of longevity, we examined the potential involvement of DAF-2 downstream transcription factors HSF-1 (heat shock transcription factor 1), SKN-1/nuclear factor erythroid 2 (NRF2) and DAF-16/FOXO in lifespan extension. We found that Probio-M9-mediated extension of the C. elegans lifespan was independent of DAF-16, but dependent on SKN-1 and HSF-1 (Fig. 5c–e and Supplementary Table 5), suggesting that SKN-1 and HSF-1 might be required for the longevity effect of Probio-M9.
SKN-1 and HSF-1 function as stress resistance transcription factors22,36. To confirm whether Probio-M9 feeding improves stress resistance and extends the lifespan of C. elegans, we investigated two GFP reporters skn-1::gfp (transcriptional target of skn-1) and hsp-16.2p::gfp (a direct transcriptional target of hsf-1) to reflect the stress resistance SKN-1 and HSF-1, respectively. Our results showed that the expression of skn-1::gfp increased in the intestine of worms fed with OP50 + Probio-M9, but had no effect on hsp-16.2p::gfp (Fig. 5f–i). Furthermore, we detected the expression of GST-4, an indicator of SKN-1 activity37, and the results indicated that the expression of gst-4p::gfp increased in C. elegans fed with OP50 + Probio-M9, compared with those fed with OP50 (Fig. 5j, k). Taken together, these results suggested that Probio-M9 influences lifespan extension and stress resistance by a skn-1-dependent but hsf-1-independent mechanism.
Probio-M9 acts on host mitochondria to promote longevity
To further probe the molecular mechanism underlying the stress resistance effect of Probio-M9, we investigated two other organelle-specific stress responses endoplasmic reticulum unfolded protein response (UPRER) and UPRmt. Probio-M9 did not affect the induction of GFP reporter of stress gene in the UPRER (hsp-4p::gfp) (Supplementary Fig. 3a, b), it did have an impact on mitochondrial stress. Particularly, the expression of hsp-6p::gfp increased in the intestine and tail of worms fed with OP50 + Probio-M9 (Fig. 6a, b), suggesting that feeding with Probio-M9 induces the UPRmt.
a, b UPRmt was detected by mitochondrial chaperone reporter hsp-6p::gfp. The representative photomicrographs of hsp-6p::GFP were shown in a, and the fluorescence intensity of hsp-6p::GFP was quantified using ImageJ software (b). The hsp-6p::GFP worms grown on OP50 + Probio-M9 showed increased ability to respond to stress in the intestine and tail (N = 40 worms, p < 0.05, Student’s t test). c In the mutant of UPRmt transcription factor atfs-1 (gk3094), the lifespan extension conferred by Probio-M9 is fully abolished (N = 90 worms, p > 0.05, Log rank test). d, e Probio-M9 is not able to extend the lifespan in the mutants of C. elegans ETC components, isp-1 (qm150) (d) and nuo-6 (qm200) (e) (N = 90 worms, p > 0.05, Log rank test). f, g The OP50 or OP50 + Probio-M9 fed worms were stained with TMRE to indicate the mitochondrial membrane potential. The pharyngeal bulbs were taken (f) and statistical analysis of TMRE fluorescence intensity in arbitrary units was shown in g (N = 30 worms, p < 0.001, Student’s t test). In a and f, scale bar, 20 μm. In b and g, results are shown by arbitrary units (a.u.), values are presented as the mean ± SEM.
To confirm the involvement of molecular components known to activate the UPRmt, we first checked ATFS-1, a pivotal transcription factor in UPRmt regulation. As expected, OP50 + Probio-M9 was unable to extend the lifespan of the loss-of-function atfs-1 (gk3094) mutant (Fig. 6c and Supplementary Table 7). We also examined the effects of Probio-M9 on loss-of-function mutants isp-1 (qm150) and nuo-6 (qm200) to evaluate whether ISP-1 and NUO-6, components of the ETC, were associated with Probio-M9-mediated promotion of host longevity. Again, OP50 + Probio-M9 was unable to extend the lifespan of the isp-1 (qm150) or nuo-6 (qm200) mutants (Fig. 6d, e and Supplementary Table 7), suggesting that the lifespan extending effects of Probio-M9 converge with the pro-longevity effects of reduced mitochondrial function in C. elegans. These results showed that Probio-M9 extends its host’s lifespan by enhancing the UPRmt and thus maintaining mitochondrial homeostasis in intestinal cells.
To evaluate the effect of Probio-M9 on mitochondrial membrane potential, we used a mitochondria-specific fluorescent dye, tetramethylrhodamine ethyl ester (TMRE), to stain worms. The TMRE fluorescence intensity was significantly increased in the pharyngeal bulbs of worms fed with OP50 + Probio-M9 compared with OP50 (Fig. 6f, g), suggesting that Probio-M9 increases the mitochondrial membrane potential of the host. Because the membrane potential across the inner membrane is the major driving force for mitochondria to produce ATP, Probio-M9 feeding might contribute to ATP production and storage in the cells and body of the C. elegans.
Identification and analysis of differential metabolites between OP50 and Probio-M9
To elucidate the mechanism underlying Probio-M9-mediated longevity, we performed metabolomics analysis using ultra-performance liquid chromatography-mass spectrometry to identify differentially regulated metabolites between OP50 and Probio-M9. Total ion chromatography (TIC) and principal component analysis (PCA) were used to analyze and evaluate the quality control (QC) bacterial data collected in the experiment. The TIC for samples was tested in positive and negative modes. These analyses revealed a high level of superposition when comparing the retention time and intensity across each chromatographic peak, reflecting high instrument stability (Supplementary Fig. 4a, b). PCA scores showed that the mixed quality control bacteria (mix01, mix02 and mix03) were well-gathered, but were separated from OP50 and OP50 + Probio-M9, indicating that the repeatability of the experimental conditions and data obtained were stable and reliable (Supplementary Fig. 4c).
Totally 1,253 metabolites were identified and quantified. PCA, orthogonal partial least squares discriminant analysis (OPLS-DA) models, and a heat map were used to evaluate the presence of metabolic variations between OP50 and OP50 + Probio-M9. PCA scores, OPLS-DA score plots and the heat map indicated that OP50 and OP50 + Probio-M9 were distributed in different regions, without overlap or crossover, instead, with obvious separation (Fig. 7a–c). Differentially regulated metabolites between OP50 and OP50 + Probio-M9 were considered to be statistically significant when the change in metabolite level simultaneously met the criteria of a fold change (FC) ≤ 0.5 or ≥ 2 and a variable importance in projection (VIP) score threshold ≥ 1. These metabolites were then represented using a volcano plot (Fig. 7d), with the details summarized in Supplementary Data 1. Metabolites showing significant differences were divided into 14 main classes, including amino acids and their metabolites, carboxylic acids and its derivatives, organic acids and its derivatives, sphingolipid, nucleotides and their metabolites, benzene and its derivatives, and heterocyclic compounds. The top ten up-regulated and top ten down-regulated metabolites with Log2FC values between OP50 and OP50 + Probio-M9 are shown in Fig. 7e, reflecting the levels of variation in metabolites in these pathways. To correlate the Probio-M9-associated changes with metabolic pathways, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway database was used to identify pathway enrichment patterns for metabolites with significant differences. Eighty-two metabolic pathways were identified as being affected by the metabolic alterations, (Supplementary Fig. 4d), with twenty of the significantly changed metabolites contributing to enrichment in pathways related to tryptophan metabolism, fatty acid biosynthesis, alanine, aspartate and glutamate metabolism, ferroptosis, galactose metabolism and sphingolipid metabolism (Fig. 7f), indicating that these pathways might contribute to the host lifespan extension mediated by Probio-M9.
PCA scores (a), OPLS-DA score plots (b) and heat map (c) of metabolites showed significant differences between OP50 and OP50 + Probio-M9, without overlap or crossover, with obvious separation. d Volcano plot displaying significantly metabolites between OP50 and OP50 + Probio-M9. Compared with OP50, purple dots represent metabolites that were increased with OP50 + Probio-M9 (up-regulated), blue dots represent metabolites that were decreased with OP50 + Probio-M9 (down-regulated), and red dots represent metabolites that were not changed with OP50 + Probio-M9 (insignificant). e The top ten changed metabolites between OP50 and OP50 + Probio-M9. Compared with OP50, red bars represent metabolites that were up-regulated by OP50 + Probio-M9, green bars represent metabolites that were down-regulated by OP50 + Probio-M9. f KEGG enrichment analysis of metabolites that were significantly changed between OP50 and OP50 + Probio-M9 (the color of the point represents p value, and the color is the redder, the enrichment degree is the more significant; the size of the point represents the number of enriched metabolites within this pathway).
Discussion
This study revealed that feeding with Lacticaseibacillus rhamnosus Probio-M9, which is isolated from healthy breast milk, can extend the lifespan of the model organism C. elegans (Fig. 8). We found that feeding with live Probio-M9, but not heat-inactivated Probio-M9, improved the health of worms. Probio-M9 extended the lifespan of C. elegans by regulating the p38 cascade and daf-2 signaling pathways, independent on daf-16 but dependent on skn-1. By activating the UPRmt, it also appears to protect and repair damaged mitochondria to maintain metabolic stability. Meanwhile, amino acid metabolism, sphingolipid metabolism, galactose metabolism and fatty acid metabolism may be involved in the Probio-M9-mediated longevity of C. elegans.
Live Probio-M9 feeding increases the lifespan of worms via the Insulin/IGF-1 signaling (DAF-2/SKN-1) and p38 MAPK signaling (NSY-1–SEK-1–PMK-1) pathways. It also triggers mitochondrial unfolded protein response (UPRmt) to maintain mitochondrial homeostasis. Therefore, Probio-M9 feeding extends the lifespan of C. elegans by improving the stress resistance, activating defensive system, and enhancing UPRmt.
It has been reported that probiotics benefit their hosts by improving the intestinal microecological balance38,39,40, regulating the immune system10,41,42 and attaching to the intestinal tract43. We previously reported that Probio-M9 has the ability to survive in high bile salts or at a low pH in vitro44. Additionally, we found that Probio-M9 inhibits the formation of colorectal tumors in mice by regulating the intestinal environment and improving the inflammatory response45.
DR has been shown to extend lifespan and delay aging in a variety of species32,46,47. Although DR is known to extend the lifespan of C. elegans, the worms under DR are smaller in size than those fed with a normal diet48. In this study, the body size and development of worms fed with OP50 + Probio-M9 were normal, and the developmental cycle was similar to those fed with OP50. Furthermore, OP50 + Probio-M9 feeding extended the lifespan of loss-of-function eat-2 (ad1116) and aak-2 (ok524) mutants of the DR signaling pathway, indicating that Probio-M9 extends the lifespan of C. elegans in a DR-independent manner. Consistent with these findings, Lactobacillus gasseri SBT2055 was shown to extend the lifespan of C. elegans by 37% in a DR-independent manner49.
Decreased muscle function and lipofuscin accumulation are closely related to aging2,50,51. Our study showed that feeding with Probio-M9 enhanced locomotor ability and reduced lipofuscin accumulation in older worms, indicating that Probio-M9 improves the healthspan of the host and extends their lifespan. This is consistent with a previous report that Weissella also increased locomotor ability and lowered the accumulation of lipofuscin in older worms52. Furthermore, at day 5 of adulthood, the survival rate of worms exposed to high temperature (34 °C) was obviously higher in those fed with OP50 + Probio-M9 than those fed with OP50, demonstrating that Probio-M9 feeding confers worms with an increased ability to endure the environmental stress. Consistent with our findings, Bacillus subtilis NCIB3610 was shown to extend the lifespan of C. elegans by enhancing resistance to heat stress, osmotic stress, metal stress and H2O2 oxidative stress53. The present study, which demonstrates that Probio-M9 feeding can enhance the stress response and extend the lifespan of C. elegans, provides a theoretical basis for further study of Probio-M9 in delaying aging processes in its host.
Because the intestinal cells of C. elegans are structurally similar to those of humans, they are considered an ideal tool to explore bacteria-host communications in the intestines54. The ability to adhere to the intestinal lining, a prerequisite for the colonization and physiological or pathophysiological effects of both detrimental and salutary bacteria in the body, is a key indicator for screening probiotic strains and reducing the accumulation of harmful bacteria. Several studies have shown that beneficial bacteria help to maintain the intestinal microbiome homeostasis and thereby promote host health42,55,56. However, when E. coli was used as a sole dietary supplementation in C. elegans, the increased numbers of E. coli in the worm intestine contributed to increased pathogenicity and accelerated aging in the worms57. Our study also showed that the extent of lifespan extension conferred by Probio-M9 was dependent on the timing of Probio-M9 feeding. Furthermore, our quantitative CFU assay showed the continued presence of Probio-M9 in worms after switching from OP50 + Probio-M9 to OP50 feeding, similar to the positive control LGG. These results are also consistent with a previous report in which plate counting and transmission electron microscopy analysis were used to show that Limosilactobacillus fermentum JDFM216 adhered to the intestinal tract of C. elegans41. However, OP50 + heat-inactivated Probio-M9 was unable to extend the lifespan of worms, suggesting that Probio-M9 in an active state has beneficial effects on host health that extend the lifespan of worms, consistent with the definition of probiotics6. By contrast, a previous report showed that Bifidobacterium longeum subsp. longeum BB68 that was heat-inactivated at 95°C for 30 min significantly extended the lifespan of worms by 28%58, indicating that probiotics might have strain-specific properties. Altogether, our results indicate that Probio-M9 has beneficial effects on health-promoting and longevity in C. elegans, and these effects are dependent on adhesion of Probio-M9 to the worm intestine.
It is well known that aging is a complex process involving interactions among multiple signaling pathways16,59,60, including the p38 signaling pathway, a key signaling pathway that regulates host lifespan61. Our results showed that OP50 + Probio-M9 was unable to extend the lifespan of loss-of-function pmk-1 mutant, indicating that the p38 cascade might be involved in the longevity related to Probio-M9 feeding, consistent with a previous report of Lacticaseibacillus rhamnosus GG extending the lifespan of C. elegans by activating the pmk-1 (p38 MAPK) signaling pathway62. The TIR-1 effector protein can activate the downstream PMK-1 signaling pathway63,64. Our data demonstrated that OP50 + Probio-M9 extended the lifespan of worms by activating TIR-1, upstream of the p38 signaling pathway. Related studies have also confirmed that Lactobacillus acidophilus NCFM activates the C. elegans immune response against Enterococcus faecalis and Staphylococcus aureus through TIR-1 and PMK-1 immune signaling pathways, and delays host aging as well34. Additionally, ATF-7, which acts downstream of the p38 signaling pathway, has a dual immunomodulatory effect on C. elegans. ATF-7 suppresses the transcription of immune-related genes. However, this inhibition is eliminated when p38 (PMK-1) phosphorylation occurs, after which ATF-7 acts as an activator, promotes the activation of immune genes65. We also observed that OP50 + Probio-M9 was unable to extend the lifespan of loss-of-function atf-7 mutant, suggesting that the innate immune pattern recognition system might be involved in the Probio-M9-activated p38 cascade. Usually E. coli is used as the sole dietary supplementation for C. elegans culture; however, proliferating bacteria might be pathogenic to older worms66. Therefore, the ingested Probio-M9 might activate the host defense systems through PMK-1 to contribute its health-promoting effects. In a future investigation, we plan to compare the transcriptomes and metabolomes of pmk-1 mutant and wild-type N2 worms fed with OP50 + Probio-M9, which may reveal genes and relevant metabolites regulated by pmk-1 in contributing to retarding aging.
SKN-1 controls specific pathways of stress resistance and is also widely involved in immune response, metabolism, and detoxification67. Kato et al. showed that Clostridium butyricum MIYAIRI 588 could improve stress resistance and extend the lifespan of C. elegans via skn-168. Our results illustrated that OP50 + Probio-M9 was unable to extend the lifespan of loss-of-function skn-1 mutant, indicating that SKN-1 might be required for the Probio-M9-mediated lifespan extension. SKN-1 has been shown to be directly suppressed by DAF-222, and DAF-2 signaling pathway via SKN-1 activity might regulate the lifespan of worms fed with Probio-M9. Bishop and Guarente revealed that the stress resistance function of SKN-1 was mediated by its expression in the intestine48. Our results showed that the intensity of fluorescence for SKN-1 in the intestine of worms was significantly stronger in OP50 + Probio-M9-fed compared with OP50. Besides, OP50 + Probio-M9 feeding increased the expression of gst-4p::gfp in C. elegans, the downstream target of skn-169, suggesting that Probio-M9 feeding activates SKN-1, which in turn contributes to improving stress resistance and extending the lifespan of C. elegans. Consistent with our findings, Komura et al. found that Bifidobacterium infantis feeding improves stress resistance and extends the lifespan of C. elegans via skn-170 signaling pathway.
Cellular organelles mitochondria were likely originated from bacteria; indeed, they share a number of metabolic pathways71. Mitochondria are involved in aging processes, as a signaling hub for both metabolism and the vital signaling pathways72. Our results demonstrated that Probio-M9 enhanced intestinal tract UPRmt levels and extended lifespan of worms, probably through the mitochondrial ETC components ISP-1 and NUO-6. The reduced function of mitochondrial ETC complexes contributes to the activation of the UPRmt to maintain mitochondrial homeostasis73,74. UPRmt-activated ATFS-1 in turn inhibits ETC gene expression in the mitochondrial genome75. Consistent with our findings, Han et al. found that E. coli mutants with increased secretion of the metabolite colanic acid (CA) extended the lifespan of C. elegans by regulating host mitochondrial dynamics and the UPRmt, and that purified CA polymers promoted the longevity of worms through ATFS-12. Our results suggest that ATFS-1 is also a crucial mediator of the Probio-M9 health-promoting effect.
Bacterial metabolites play crucial roles in host aging57,76,77. Metabolomics analysis in this study found that the differentially expressed metabolites were mainly enriched in pathways involving amino acid metabolism, galactose metabolism, sphingolipid metabolism, fatty acid biosynthesis and cAMP signaling. Amino acids affect the longevity of the host, tryptophan and proline contribute to delaying the aging of C. elegans, while phenylalanine accelerates aging;78 aspartate, glutamate and methionine extend the lifespan of yeast79. Our results suggested that the significant enrichment of bacterial tryptophan, aspartate and glutamate metabolites could be related with the extension of the lifespan of C. elegans by Probio-M9. Bacterial glucose metabolites, such as glucose, galactose and lactose, may also be associated with host longevity participate in anti-aging activities57. Our metabolomics data indicated that the significant enrichment of bacterial galactose metabolites may contribute to the extension of the lifespan of C. elegans by Probio-M9. Consistent with our findings, Zanni et al. found that galactose secreted by Lactobacillus delbrueckii subsp. bulgaricus might be associated with its beneficial effects on C. elegans80. The accumulation of bacterial metabolites such as sphingolipid and ceramide are closely related to aging and age-related diseases81,82. It has been shown that sphingolipid metabolism might be vital for enhancing locomotor ability83 and improvement of the health status of C. elegans, which were partially dependent on the insulin receptor daf-2 signaling pathway84. Goya et al. observed that Bacillus subtilis mediated host sphingolipid metabolism and inhibited a-syn aggregation in C. elegans85. In light of our results, we revealed that Probio-M9 might regulate sphingolipid metabolism through nutrient sensing to extend lifespan of C. elegans. Besides, our data showed that 2-aminoethanesulfonic acid derived from taurine metabolism was significantly accumulated upon OP50 + Probio-M9 feeding. Taurine is of great significance in the scavenging of lipid metabolism, immune function, and other physiological functions, making it an important promoter of anti-inflammatory activity86,87. However, such inference needs further experimental verification. It will be of interest to extract the most key metabolites in Probio-M9 and perform further lifespan verification experiments to demonstrate the health-promoting and longevity extension effects of probiotic metabolites on host. It is necessary to identify the major anti-aging components of probiotics and explore signaling pathways that probiotic utilizes to delay host aging. Although challenging, future study should focus on the effects of probiotics on longevity in mammals and elucidate the molecular mechanisms of action.
Methods
Caenorhabditis elegans strains and culture
The nematodes C. elegans and bacterial strains used in this study can be found in Supplementary Table 8. Worms were cultured at 20 °C on nematode growth medium (NGM) agar88 plates seeded with pre-cultured bacterial strains according to standard techniques89. Hermaphrodites of C. elegans were age-synchronized by isolating eggs, and then transferred to NGM plates seeded with E. coli OP5090.
Culture conditions of bacterial strains
In this study, Probio-M9 was isolated from healthy breast milk. Lacticaseibacillus rhamnosus GG (LGG) was acquired from Guangdong Microbial Culture Preservation Center. Probio-M9 and LGG were anaerobic cultured in MRS broth at 37 °C for 24 h and continuously cultured to three generations as test bacterial solution. E. coli OP50 was incubated in Luria-Bertani (LB) broth at 37 °C for 12 h with a shaking at 225 rpm.
For preparing bacterial plates for the worms feeding, Probio-M9 and LGG were collected through centrifugation at 4000 × g for 10 min, and washed with M9 buffer (3 g KH2PO4, 6 g Na2HPO4, 5 g NaCl and 1 mL 1 M MgSO4) twice, and then resuspended in E. coli OP50 (6.8 × 108 CFU/mL) to a final dose (2.82 × 109 CFU/mL and 3.02 × 109 CFU/mL, respectively), which were used as the bacteria supply. Before resuspending in E. coli OP50, the concentrated Probio-M9 was heated at 95°C for 30 min and used as heat-inactived Probio-M9. 40 μL of the bacteria solution was inoculated on 60 mm NGM plate.
Lifespan assay
As described previously, all lifespan analyses were performed at 20 °C88. Late L4 stage worms were used as t = 0 for lifespan assay and transferred to fresh plates every two days until all worms died, unless otherwise noted. For each experiment, 90 worms were tested on 3 plates (30 worms per plate) for bacterial strain. Survival of the worms was determined on a daily basis. Ceasing pharyngeal pumping and not responding to gentle mechanical stimulation by platinum wire picker were scored death2,53. Worms, which accidentally lost, extruded organs or displayed matricidal hatching, were censored and excluded from lifespan analysis.
Bacterial choice assay
Worm eggs were cultured on OP50 or OP50 + Probio-M9 until they reached late L4 stage. And then these worms were washed with M9 buffer three times. For the first paradigm, OP50 or OP50 + Probio-M9 was seeded on the center of plate, and 50 worms were placed at the edges of each end of the plate, being equidistant from OP50 or OP50 + Probio-M9. For the second paradigm, OP50 or OP50 + Probio-M9 was seeded on the edges of each end of the plate, and 100 worms were placed in the center of the plate, being equidistant from OP50 and OP50 + Probio-M9. The worms were allowed to move freely for 1 h or 2 h, and the number of worms entering each bacterial lawn was counted.
Life cycle, brood size and body size assays
The assays were determined according to the previous method55,91.
To determine the life cycle of worms, 20 eggs were placed on 20 plates and used as t = 0 for life cycle analysis. When the worm lays its first egg, which is the time to complete the life cycle.
To evaluate the brood size of worms, 20 synchronized hermaphrodites at late L4 stage were transferred to fresh plates daily (one worm per plate) until reproductive termination, the total number of offspring per worm was counted.
To measure the body size of worms, 20 worms were selected from day 1–5 of adulthood. The anaesthetized with 25 mM levamisole worms were taken photos using a stereomicroscope (ZEISS Axio Imager M2), and analyzed using ImageJ software (National Institutes of Health, Bethesda, MD, USA). The projected area of worm was computed and used as an indicator of body size.
Pharyngeal pumping rate assay
The pharyngeal pumping rate of worms at different ages was examined according to the previous method2. Worms were selected from days 2–14 of adulthood and the pharyngeal pumping rate was measured using SMZ-168 stereo microscope (Motic). The times of pharyngeal constrictions in the terminal bulb of the pharynx for 30 s interval were counted.
Locomotor scoring
The motility of worms at different ages was measured according to the previous method50. Worms were selected from day 8–16 of adulthood and their locomotor scoring was measured. Classification of worms was based on difference in the movement activity. When worms show spontaneous and/or rhythmical sinusoidal movement (normal locomotion), they were classified as class “A”; worms with irregular and/or uncoordinated movement (uncoordinated/sluggish), they were classified as class “B”; worms moved merely their head and/or tail in respond to a moderate touch with a platinum wire picker (cannot move body), they were classified as class “C”; dead worms were classified as class “D”. At least 90 worms were scored for each experiment.
Lipofuscin accumulation assay
The autofluorescence of intestinal lipofuscin was detected according to the previous method52. Day 10 or 15 worms were washed with M9 buffer three times, and then anaesthetized with 25 mM levamisole. Autofluorescence images of lipofuscin were obtained using blue excitation light (405 nm) in confocal laser scanning microscope (Olympus FV1200). The accumulation level of lipofuscin was detected by fluorescence intensity quantification using ImageJ software.
Thermotolerance assay
To assess thermotolerance, the survival rate of worms was measured according to the previous method53. Day 5 worms were exposed at 34 °C and then the survival rate was measured after 12 h, 18 h and 24 h, respectively.
Determination of bacterial CFU within worm gut
The number of CFU in the worm gut was determined according to the previous method55. Day 5 or 10 worms were washed three times with M9 buffer and transferred them on empty NGM plates for 1 h to remove surface-attached bacteria. 5 worms were placed into M9 buffer and ground with a glass grinder. The worm lysate was continuously diluted with M9 buffer, and anaerobic cultured at 37 °C for 48 h on MRS plates, the number of CFU was counted.
The number of bacteria pulse-chase in the worm intestine was determined according to the previous method2. Day 10 of adulthood worms were selected from the NGM plates lawned with OP50 + LGG (as a positive control) or OP50 + Probio-M9, washed with M9 buffer three times, and transferred them to the plate lawned with OP50. After 3 days, the number of CFU was tested as described above.
Analysis of the fluorescence intensity of stress responses
The fluorescence expression was measured according to the previous method92. Fluorescence intensity images were obtained using green excitation light (473 nm) of laser confocal scanning microscope (Olympus FV1200). Fluorescence intensity of GFP was analyzed using ImageJ software.
To measure hsp-4p::gfp fluorescence expression, day 5 worms were induced for 4 h in M9 buffer containing 50 ng/mL tunicamycin or tantamount DMSO. To measure hsp-16.2p::gfp fluorescence expression, day 5 worms were heated at 34 °C for 20 min, and then recovery for 6 h. At last, the fluorescence photos were taken and analyzed as described above.
Estimation of mitochondrial membrane potential
To evaluate the effect of mitochondrial membrane potential, a specific fluorescent dye tetramethylrhodamine ethyl ester (TMRE) was used to stain worms. Worms were cultured in OP50 or OP50 + Probio-M9 containing 10 μm TMRE for 5 days, and then placed on 2% agarose pads with 25 mM levamisole to take photos using red excitation light (556 nm) of laser confocal scanning microscope (Olympus FV1200). Fluorescence intensity of pictures was analyzed using ImageJ software.
Metabolomics analysis
The bacteria of control group (OP50) and experimental group (OP50 + Probio-M9) prepared for worms feeding were collected from NGM plates with M9 buffer to 15 mL centrifuge tube, washed with M9 buffer for three times, taken 50 ± 2 mg, added cold steel balls and homogenized at 30 Hz for 3 min. The homogenized solution was added 1 mL 70% methanol with internal standard. Samples were vortexed for 2 min, incubated on ice for 15 min and then centrifuged (13,400 × g, 4 °C, 10 min). Samples were drawed 400 μL of supernatant into the EP tube and stored at −20 °C overnight. Samples were thawed on ice, centrifuged (13,400 × g, 4 °C, 3 min), and taken 200 μL of supernatant in the liner of the injection bottle for on-board analysis.
Metabolomics samples were analyzed using UPLC system (ExionLC AD, https://sciex.com.cn/) and Quadrupole-Time of Flight (TripleTOF 6600, AB SCIEX).
Samples were separated using a ACQUITY HSS T3 (2.1 × 100 mm, 1.8 μm) at 40 °C. Mobile phases were consisting of mobile phase A (water with 0.1% formic acid, v/v) and mobile phase B (acetonitrile with 0.1% acetic acid, v/v). The elution gradient was as follows: 95% A to 5% B for 0.0 min; 10% A to 90% B for 10.0 min; 95% A to 5% B for 14.0 min. The flow rate was 0.35 mL/min with an injection volume of 5 μL.
Software Analyst 1.6.3 was used to process the mass spectrum data. The repeatability of metabolite extraction and detection were estimated using TIC and multi peak multiple reaction monitoring (MRM). According to home-made metadata database (MWDB) and other databases, qualitative analysis of information and secondary general data was carried out based on retention time (RT) and mass-to-charge ratio. The metabolite structure analysis referred to several public mass spectrometry databases, including Human Metabolome Database (HMDB, http://www.hmdb.ca/), Metlin (http://metlin.scripps.edu/index.php), massbank (http://www.massbank.jp/) and knapsack (http://kanaya.naist.jp/knapsack/).
Ten microliters per sample was used for quality control (QC) of metabolomics analysis. When running sample sets on column, one QC sample was injected after 10 samples. Metabolite quantification was analyzed using MRM of triple quadrupole mass spectrometry. The mass spectrum file under the sample machine was opened with multi-quantitative software, and the chromatographic peaks were integrated and calibrated. The peak area of each chromatographic peak represented the relative content of the corresponding substance. The integral data of chromatographic peak area was extracted and saved, and the positive and negative ions of metabolites were removed using self-built software package.
The coefficient of variation (CV) values of metabolites in QC samples were calculated, and metabolites with CV values greater than 0.5 were removed. When metabolites were detected in both positive and negative ionization modes, those with larger CVs were removed in either positive or negative mode. Orthogonal partial least squares discriminant analysis (OPLS-DA) model was used to identify differences in metabolic profiles between groups.
Statistics and reproducibility
All results were derived from at least three biologically replicates. Prism8 software was used to analyze the lifespan curves, and Log rank (Mantel–Cox) method was used to analyze the significance of the difference. For statistical data, Student’s t test, Chi-squared test, one-way ANOVA and two-way ANOVA analysis coupled with multiple comparisons were used for significance analysis. In all cases, p < 0.05 was considered significant. In the figures, the asterisk indicates the statistical significance of the Log rank test, Student’s t test, Chi-squared test, one-way ANOVA and two-way ANOVA analysis as compared to control. For metabolomics data, the OPLS-DA model was applied using R package “MetaboAnalyst”. Permutation test repeated 200 times were used to verify this model, and p < 0.05 indicated the available OPLS-DA model. The significance of each metabolite was measured by Student’s t test and fold change, and one-tailed Student’s t test or Fisher’s exact test were used to analyze the statistical significance, p < 0.05 was considered to be statistically significant. Using the Benjamini–Hochberg method, p value was corrected for multiple testing by false-discovery rate (FDR).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The source data used to create the box plots in the figures were deposited in Supplementary Data 2. The source data used to create the survival curves in the figures were deposited in Supplementary Data 3. The source data used to create the differentially metabolites between OP50 and Probio-M9 in the figures were deposited in Supplementary Data 4.
References
Singh, T. & Newman, A. B. Inflammatory markers in population studies of aging. Ageing Res. Rev. 10, 319–329 (2011).
Han, B. et al. Microbial genetic composition tunes host longevity. Cell 169, 1249–1262 (2017).
Kasozi, K. I. et al. Low concentrations of Lactobacillus rhamnosus GG (Yoba) are safe in male Drosophila melanogaster. BMC Res. Notes 12, 269–273 (2019).
Pan, F. W. et al. Predominant gut Lactobacillus murinus strain mediates anti-inflammaging effects in calorie-restricted mice. Microbiome 6, 54–70 (2018).
So, S., Tokumaru, T., Miyahara, K. & Ohshima, Y. Control of lifespan by food bacteria, nutrient limitation and pathogenicity of food in C. elegans. Mech. Ageing Dev. 132, 210–212 (2011).
Hill, C. et al. The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 11, 506–514 (2014).
Zhang, J. C. et al. Probiotics maintain the intestinal microbiome homeostasis of the sailors during a long sea voyage. Gut Microbes 11, 930–943 (2020).
Ma, T. et al. Probiotic consumption relieved human stress and anxiety symptoms possibly via modulating the neuroactive potential of the gut microbiota. Neurobiol. Stress 14, 100294 (2021).
Hou, Q. C. et al. Probiotic-directed modulation of gut microbiota is basal microbiome dependent. Gut Microbes 12, 1736974 (2020).
Komura, T., Ikeda, T., Hoshino, K., Shibamura, A. & Nishikawa, Y. Caenorhabditis elegans as an alternative model to study senescence of host defense and the prevention by immunonutrition. Adv. Exp. Med. Biol. 710, 19–27 (2012).
Landete, J. M. et al. Probiotic bacteria for healthier aging: immunomodulation and metabolism of phytoestrogens. BioMed. Res. Int 2017, 1–10 (2017).
Zhou, Y. M. et al. An efficient and novel screening model for assessing the bioactivity of extracts against multidrug-resistant Pseudomonas aeruginosa using Caenorhabditis elegans. Biosci. Biotechnol. Biochem. 75, 1746–1751 (2011).
Finch, C. E. & Ruvkun, G. The genetics of aging. Annu. Rev. Genomics Hum. Genet. 2, 435–462 (2001).
Heintz, C. & Mair, W. You are what you host: microbiome modulation of the aging process. Cell 156, 408–411 (2014).
Irazoqui, J. E., Urbach, J. M. & Ausubel, F. M. Evolution of host innate defence: insights from Caenorhabditis elegans and primitive invertebrates. Nat. Rev. Immunol. 10, 47–58 (2010).
Kim, D. H. et al. A conserved p38 MAP kinase pathway in Caenorhabditis elegans innate immunity. Science 297, 623–626 (2002).
Arthur, J. S. C. & Ley, S. C. Mitogen-activated protein kinases in innate immunity. Nat. Rev. Immunol. 13, 679–692 (2013).
Newton, K. & Dixit, V. M. Signaling in innate immunity and inflammation. Cold Spring Harb. Perspect. Biol. 4, a006049 (2012).
Kimura, K. D., Tissenbaum, H. A., Liu, Y. X. & Ruvkun, G. daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science 277, 942–946 (1997).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993).
Altintas, O., Park, S. & Lee, S. J. V. The role of insulin/IGF-1 signaling in the longevity of model invertebrates, C. elegans and D. melanogaster. BMB Rep. 49, 81–92 (2016).
Tullet, J. M. A. et al. Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132, 1025–1038 (2008).
Shin, M. G. et al. Bacteria-derived metabolite, methylglyoxal, modulates the longevity of C. elegans through TORC2/SGK-1/DAF-16 signaling. Proc. Natl Acad. Sci. USA 117, 17142–17150 (2020).
Pryor, R. et al. Host-microbe-drug-nutrient screen identifies bacterial effectors of metformin therapy. Cell 178, 1299–1312 (2019).
Balaban, R. S., Nemoto, S. & Finkel, K. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005).
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002).
Feng, J. L., Bussière, F. & Hekimi, S. Mitochondrial electron transport is a key determinant of life span in Caenorhabditis elegans. Dev. Cell 1, 633–644 (2001).
Pellegrino, M. W., Nargund, A. M. & Haynes, C. M. Signaling the mitochondrial unfolded protein response. Biochim. Biophys. Acta 1833, 410–416 (2013).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Abada, E. A. E. et al. C. elegans behavior of preference choice on bacterial food. Mol. Cells 28, 209–213 (2009).
Zhang, Y., Lu, H. & Bargmann, C. I. Pathogenic bacteria induce aversive olfactory learning in Caenorhabditis elegans. Nature 438, 179–184 (2005).
Klass, M. R. Aging in the nematode Caenorhabditis elegans: major biological and environmental factors influencing life span. Mech. Ageing Dev. 6, 413–429 (1977).
Ouwehand, A. C., Kirjavainen, P. V., GroKnlund, M. M., Isolauri, E. & Salminen, S. J. Adhesion of probiotic micro-organisms to intestinal mucus. Int. Dairy J. 9, 623–630 (1999).
Kim, Y. & Mylonakis, E. Caenorhabditis elegans immune conditioning with the probiotic bacterium Lactobacillus acidophilus strain NCFM enhances gram-positive immune responses. Infect. Immun. 80, 2500–2508 (2012).
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512 (2010).
Hsu, A. L., Murphy, C. T. & Kenyon, C. Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science 300, 1142–1145 (2003).
Leiers, B. et al. A stress-responsive glutathione S-transferase confers resistance to oxidative stress in Caenorhabditis elegans. Free Radic. Biol. Med. 34, 1405–1415 (2003).
Malaguarnera, G. et al. Probiotics in the gastrointestinal diseases of the elderly. J. Nutr. Health Aging 16, 402–410 (2012).
Rauch, M. & Lynch, S. V. The potential for probiotic manipulation of the gastrointestinal microbiome. Curr. Opin. Biotechnol. 23, 192–201 (2012).
Azad, M. A. K., Sarker, M., Li, T. J. & Yin, J. Probiotic species in the modulation of gut microbiota: an overview. BioMed. Res. Int. 2018, 1–8 (2018).
Park, M. R. et al. Lactobacillus fermentum strain JDFM216 stimulates the longevity and immune response of Caenorhabditis elegans through a nuclear hormone receptor. Sci. Rep. 8, 7441–7450 (2018).
Lee, J. et al. Evaluation of probiotic characteristics of newly isolated Lactobacillus spp.: immune modulation and longevity. Int. J. Food Microbiol 148, 80–86 (2011).
Oelschlaeger, T. A. Mechanisms of probiotic actions-a review. Int. J. Med. Microbiol. 300, 57–62 (2009).
Liu, W. J. et al. Characterization of potentially probiotic lactic acid bacteria and bifidobacteria isolated from human colostrum. J. Dairy Sci. 103, 4013–4025 (2020).
Xu, H. Y. et al. Inhibitory effects of breast milk-derived Lactobacillus rhamnosus Probio-M9 on colitis-associated carcinogenesis by restoration of the gut microbiota in a mouse model. Nutrients 13, 1143–1157 (2021).
Tain, L. S., Lozano, E., Sáez, A. G. & Leroi, A. M. Dietary regulation of hypodermal polyploidization in C. elegans. BMC Dev. Biol. 8, 28–36 (2008).
Ingram, D. K. et al. Calorie restriction mimetics: an emerging research field. Aging Cell 5, 97–108 (2006).
Bishop, N. A. & Guarente, L. Two neurons mediate diet-restriction-induced longevity in C. elegans. Nature 447, 545–549 (2007).
Nakagawa, H. et al. Effects and mechanisms of prolongevity induced by Lactobacillus gasseri SBT2055 in Caenorhabditis elegans. Aging Cell 15, 227–236 (2016).
Hosono, R., Sato, Y., Aizawa, S. I. & Mitsui, Y. Age-dependent changes in mobility and separation of the nematode Caenorhabditis elegan. Exp. Gerontol. 15, 285–289 (1980).
Pincus, Z. & Slack, F. J. Developmental biomarkers of aging in Caenorhabditis elegans. Dev. Dyn. 239, 1306–1314 (2010).
Lee, J. Y., Kwon, G. & Lim, Y. H. Elucidating the mechanism of Weissella-dependent lifespan extension in Caenorhabditis elegans. Sci. Rep. 5, 17128–17141 (2015).
Donato, V. et al. Bacillus subtilis biofilm extends Caenorhabditis elegans longevity through downregulation of the insulin-like signalling pathway. Nat. Commun. 8, 14332–14346 (2017).
Park, M. R., Yun, H. S., Son, S. J., Oh, S. & Kim, Y. Short communication: development of a direct in vivo screening model to identify potential probiotic bacteria using Caenorhabditis elegans. J. Dairy Sci. 97, 6828–6834 (2014).
Ikeda, T., Yasui, C., Hoshino, K., Arikawa, K. & Nishikawa, Y. Influence of lactic acid bacteria on longevity of Caenorhabditis elegans and host defense against Salmonella enterica serovar enteritidis. Appl. Environ. Microbiol. 73, 6404–6409 (2007).
Grompone, G. et al. Anti-inflammatory Lactobacillus rhamnosus CNCM I-3690 strain protects against oxidative stress and increases lifespan in Caenorhabditis elegans. PLoS One 7, e52493 (2012).
Brokate-Llanos, A. M., Garzón, A. & Muñoz, M. J. Escherichia coli carbon source metabolism affects longevity of its predator Caenorhabditis elegans. Mech. Ageing Dev. 141-142, 22–25 (2014).
Zhao, L. et al. The transcription factor DAF-16 is essential for increased longevity in C. elegans exposed to Bifidobacterium longum BB68. Sci. Rep. 7, 7408–7414 (2017).
Kapahi, P. et al. With TOR, less is more: a key role for the conserved nutrient-sensing TOR pathway in aging. Cell Metab. 11, 453–465 (2010).
Ma, S. et al. Caloric restriction reprograms the single-cell transcriptional landscape of rattus norvegicus aging. Cell 180, 984–101 (2020).
Aw, D., Silva, A. B. & Palmer, D. B. Immunosenescence: emerging challenges for an ageing population. Immunology 120, 435–446 (2007).
Yun, B. et al. Probiotic Lacticaseibacillus rhamnosus GG increased longevity and resistance against foodborne pathogens in Caenorhabditis elegans by regulating microRNA miR-34. Front Cell Infect. Microbiol 11, 819328 (2022).
Couillault, C. et al. TLR-independent control of innate immunity in Caenorhabditis elegans by the TIR domain adaptor protein TIR-1, an ortholog of human SARM. Nat. Immunol. 5, 488–494 (2004).
Liberati, N. T. et al. Requirement for a conserved Toll/interleukin-1 resistance domain protein in the Caenorhabditis elegans immune response. Proc. Natl Acad. Sci. USA 101, 6593–6598 (2004).
Shivers, R. P. et al. Phosphorylation of the conserved transcription factor ATF-7 by PMK-1 p38 MAPK regulates innate immunity in Caenorhabditis elegans. PLoS Genet 6, e1000892 (2010).
Garigan, D. et al. Genetic analysis of tissue aging in Caenorhabditis elegans: a role for heat-shock factor and bacterial proliferation. Genetics 161, 1101–1112 (2002).
Blackwell, T. K., Steinbaugh, M. J., Hourihan, J. M., Ewald, C. Y. & Isik, M. SKN-1/Nrf, stress responses, and aging in Caenorhabditis elegans. Free Radic. Biol. Med. 88, 290–301 (2015).
Kato, M., Hamazaki, Y., Sun, S., Nishikawa, Y. & Kage-Nakadai, E. Clostridium butyricum MIYAIRI 588 increases the lifespan and multiple-stress resistance of Caenorhabditis elegans. Nutrients 10, 1291–1303 (2018).
Ransome, V. D. H., Mccallum, K. C., Cruz, M. R. & Garsin, D. A. Ce-Duox1/BLI-3 generated reactive oxygen species trigger protective SKN-1 activity via p38 MAPK signaling during infection in C. elegans. PLoS Pathog. 7, e1002453 (2011).
Komura, T., Ikeda, T., Yasui, C., Saeki, S. & Nishikawa, Y. Mechanism underlying prolongevity induced by bifidobacteria in Caenorhabditis elegans. Biogerontology 14, 73–87 (2013).
Andersson, S. G. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–140 (1998).
Riera, C. E. & Dillin, A. Tipping the metabolic scales towards increased longevity in mammals. Nat. Cell Biol. 17, 196–203 (2015).
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011).
Houtkooper, H. R. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Nargund, A. M., Fiorese, C. J., Pellegrino, M. W., Deng, P. & Haynes, C. M. Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPR(mt). Mol. Cell 58, 123–133 (2015).
Nguyen, T. P. & Clarke, C. F. Folate status of gut microbiome affects Caenorhabditis elegans lifespan. BMC Biol. 10, 66–69 (2012).
Yu, L. et al. Bacterial respiration and growth rates affect the feeding preferences, brood size and lifespan of Caenorhabditis elegans. PloS One 10, e0134401 (2015).
Edwards, C. et al. Mechanisms of amino acid-mediated lifespan extension in Caenorhabditis elegans. BMC Genet 16, 8–31 (2015).
Wu, Z. Y., Song, L. X., Liu, S. Q. & Huang, D. J. Independent and additive effects of glutamic acid and methionine on yeast longevity. PloS One 8, e79319 (2013).
Zanni, E. et al. Combination of metabolomic and proteomic analysis revealed different features among Lactobacillus delbrueckii Subspecies bulgaricus and lactis strains while in vivo testing in the model organism Caenorhabditis elegans highlighted probiotic properties. Front. Microbiol. 8, 1206–1217 (2017).
Perez, G. I. et al. A central role for ceramide in the age-related acceleration of apoptosis in the female germline. FASEB J. 19, 860–862 (2005).
Venable, M. E., Webb-Froehlich, L. M., Sloan, E. F. & Thomley, J. E. Shift in sphingolipid metabolism leads to an accumulation of ceramide in senescence. Mech. Ageing Dev. 127, 473–480 (2006).
Chan, J. P. et al. Loss of sphingosine kinase alters life history traits and locomotor function in Caenorhabditis elegans. Front. Genet. 8, 132–147 (2017).
Mosbech, M. B. et al. Functional loss of two ceramide synthases elicits autophagy-dependent lifespan extension in C. elegans. PloS One 8, e70087 (2013).
Goya, M. E. et al. Probiotic Bacillus subtilis protects against α-synuclein aggregation in C. elegans. Cell Rep. 30, 367–380 (2020).
Aydn, A. F. et al. Carnosine and taurine treatments diminished brain oxidative stress and apoptosis in D-galactose aging model. Metab. Brain Dis. 31, 337–345 (2016).
Suliman, M. E., Bárány, P., Filho, J. C. D., Lindholm, B. & Bergström, J. Accumulation of taurine in patients with renal failure. Nephrol. Dial. Transpl. 17, 528–529 (2002).
Gusarov, L. et al. Bacterial nitric oxide extends the lifespan of C. elegans. Cell 152, 818–830 (2013).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Sulston, J. E. & Brenner, S. The DNA of C. elegans. Genetics 77, 95–104 (1974).
Keith, S. A., Amrit, F. R. G., Ratnappan, R. & Ghazi, A. The C. elegans healthspan and stress-resistance assay toolkit. Methods 68, 476–486 (2014).
Zhang, Q. et al. The mitochondrial unfolded protein response is mediated cell-non-autonomously by retromer-dependent wnt signaling. Cell 174, 870–883 (2018).
Acknowledgements
We are greatly thankful to professor Long Miao for his helpful comments and manuscript editing. We are grateful to professors Ye Tian and Xiaoyun Xu for their assistance with the worm strains. This work was financially supported by the Inner Mongolia Science and Technology Major Projects (2021ZD0014) and China Agriculture Research System of MOF and MARA and Natural Science Foundation of China (32070694, 31872822 to Y.Z.). Some worm strains were obtained from the Caenorhabditis Genetics Center (CGC, https://cgc.umn.edu/), which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440), USA.
Author information
Authors and Affiliations
Contributions
Experiments were designed by J.Z., Y.Z., Z.S., and T.S. All experiments were performed by J.Z. J.Z. wrote the manuscript with significant contributions from Y.Z., Z.S., and T.S. All authors made comments and suggestions on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Christopher Hine and Zhijuan Qiu. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Zhao, Y., Sun, Z. et al. Lacticaseibacillus rhamnosus Probio-M9 extends the lifespan of Caenorhabditis elegans. Commun Biol 5, 1139 (2022). https://doi.org/10.1038/s42003-022-04031-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-04031-2
- Springer Nature Limited
This article is cited by
-
Heat-killed probiotic Levilactobacillus brevis MKAK9 and its exopolysaccharide promote longevity by modulating aging hallmarks and enhancing immune responses in Caenorhabditis elegans
Immunity & Ageing (2024)
-
Geroprotective potential of microbiome modulators in the Caenorhabditis elegans model
GeroScience (2023)