Abstract
Well-quantified laboratory studies can provide a fundamental understanding of animal behavior in ecology, ethology and ecotoxicology research. These types of studies require observation and tracking of each animal in well-controlled and defined arenas, often for long timescales. Thus, these experiments produce long time series and a vast amount of data that require the use of software applications to automate the analysis and reduce manual annotation. In this review, we examine 28 free software applications for animal tracking to guide researchers in selecting the software that might best suit a particular experiment. We also review the algorithms in the tracking pipeline of the applications, explain how specific techniques can fit different experiments, and finally, expose each approach’s weaknesses and strengths. Our in-depth review includes last update, type of platform, user-friendliness, off- or online video acquisition, calibration method, background subtraction and segmentation method, species, multiple arenas, multiple animals, identity preservation, manual identity correction, data analysis and extra features. We found, for example, that out of 28 programs, only 3 include a calibration algorithm to reduce image distortion and perspective problems that affect accuracy and can result in substantial errors when analyzing trajectories and extracting mobility or explored distance. In addition, only 4 programs can directly export in-depth tracking and analysis metrics, only 5 are suited for tracking multiple unmarked animals for more than a few seconds and only 11 have been updated in the period 2019–2021.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Main
Animal behavior studies are fundamental in ecology, ethology, ecotoxicology, neuroscience and many other fields1,2. These studies can be performed in a wide variety of ways, ranging from observational tests in natural conditions to experimental trials in a laboratory environment. To compare experiments in laboratory conditions, performed by different research groups and with different organisms, it is important that these types of experiments are implemented in carefully controlled conditions and that they use standardized and repeatable protocols. Therefore, these experiments often use model organisms that have been widely studied, such as zebrafish or rodents, and take place in well-defined environments, so-called ‘arenas’. The size of the arena and the number of animals in an arena should also be carefully considered and controlled not to bias the study’s outcome.
Standardized tests can measure an organism’s activity in different arenas (Fig. 1a). For example, open arenas and plus-mazes are common in anxiety or motivation studies3, T-maze or Y-maze test arenas are commonly used for memory and spatial learning studies4 and three-chambered arenas are used to measure social approach or fear responses5. Other standard tests include water mazes6, elevated mazes7, arenas with light-dark transitions8 and arenas with thermal gradients or hot plates9. Stimuli or distractors can also be added to these arenas to measure behavioral changes in different conditions10. Although these arenas have different layouts, the way the data (video files) are acquired is similar (Fig. 1b).
a, Examples of different arena configurations commonly used for insects, rodents and fish. b, Experiments in laboratory conditions should be carefully designed for front or back illumination depending on the type of organism under study and the arena used. In addition, consider using diffuse illumination if details in the appearance of the animals are important, or backlight illumination to achieve high contrast for motion tracking. Make sure that no reflections from the surrounding environment or the lamp are seen.
Regardless of the arena design or organism used, most behavioral studies use video recording and analysis techniques in which the video frame rate, field of view and resolution are adjusted to capture the organism’s motion with the required temporal and spatial resolution11. These experiments can produce a vast amount of image data, which can be very time consuming to analyze, especially if manual annotation is used. In addition, manual annotation can introduce human errors and biases, which can reduce the accuracy of the results12. Therefore, objective and automatic approaches for animal analysis are needed. Software and algorithms that can track and analyze the organism’s position under study are critical for efficient research.
In general, these software packages can help to solve two main issues: detecting the position of animals via tracking algorithms or detecting the positions of the animals’ parts (so-called ‘pose estimation’). Pose estimation typically requires a previous tracking analysis. Therefore, because tracking is the main focus of this work, we refer only to the tracking software and the relevant tracking stages of pose estimation software. We explicitly exclude pose estimation software such as DeepPoseKit and DeepLabCut that are based on the extraction of images with distinct postures and the manual annotation of body parts to train machine learning models13,14,15. We think that these techniques are more oriented toward the detection of the behavior of a single animal at a fine scale and are not as directly relevant for general tracking applications. In addition, they are sufficiently different in their theoretical approach to grant a separate analysis in another review.
Tracking applications are available as free or commercial tools. Commercial software usually offer more features and flexibility, especially regarding input video formats and statistical outputs. However, many researchers cannot afford these expensive tools, and the algorithms used by these software are often not transparent to the user. Thus, we focus this analysis on free tracking software. Several new types and versions are published each year. The continuous release of new software makes it challenging to select the appropriate software for a particular experiment or understand each option’s limitations and differences. For example, many applications use the same pipeline and the same processing techniques, but there might be steps in the algorithm that are a limiting factor for some types of experiments. In addition, some programs are limited and specific in what they can do and what they can analyze.
With these considerations, we conducted our analysis using a systematic search of tracking applications on Google Scholar and Research Gate, with software published from 2008 until 2020 as the inclusion criterion. We based our analysis primarily on the descriptions of the software in their respective papers and the published results from their authors. In addition, we tested some of the analyzed software to gather information that was not available from these sources.
In this review, we compare 28 tracking applications in a comprehensive and accessible way to help guide researchers within the field of behavior studies. In contrast to previous reviews that focused on specific research fields and organisms16,17, explored only specific features such as social interactions18, did not include an in-depth discussion19 or included only a handful of software18,19, we point out the weaknesses of the applications in every step of the pipeline, identifying the problems that might face behavioral researchers when choosing a specific software. In addition, we provide a detailed taxonomy of the main algorithms and their limitations and show the emerging opportunities in animal-tracking software development.
General pipeline of a tracking program
To understand how a tracking application operates, it is essential to decompose the pipeline involved in the tracking process. Starting from video recording and ending in the analysis of animal trajectories, all the tracking applications that we review here use a similar sequence of algorithms. We show the general workflow of these tracking applications (Fig. 2) and describe each step of the pipeline below. It is important to recall that some applications do not include all the steps shown in the figure. For example, some applications lack the calibration procedure (Fig. 2b) and the possibility to analyze trajectories (Fig. 2d).
Image acquisition
This step defines how video images are obtained. There are two main approaches: offline, in which a sequence of images feeds the algorithm from a video file, or online, in which a live camera streams each image. An advantage of using the offline mode is that complex algorithms can use extensive computations and access future and past frames to process a given image. The online method uses real-time computations, which requires less storing capacity but increases the computational requirements, because each frame needs to be analyzed with speed similar to the camera frame rate to avoid losing data. As a result, the online method is currently suitable for use only with relatively simple algorithms.
Calibration
This step determines the relationship between the camera’s natural units (pixels) and the real-world units. A calibration algorithm that can correct distortions in the images caused by misalignment, projection errors and the camera lens is valuable.
Detection
This step consists of finding the spatial position of the animals in each image of a video sequence. Detection algorithms can include different steps, such as background subtraction to remove stationary features from the environment, segmentation to separate the objects that represent potential animals and a filtering step to remove false detections.
Trajectory generation
This step associates the detected animals in each image to the tracked individuals. In this step, a set of trajectories are generated, each defined by an animal’s positions in the subsequent images of the video. Trajectory generation is a trivial task for a single animal in an open arena without occlusions or reflections. However, in arenas with uneven illumination conditions, with multiple targets, possible occlusions or complex backgrounds, trajectory generation requires identity (id.) preservation algorithms to track animals when an occlusion or a crossing occurs. In general, trajectory generation is a very complex task that usually requires manual correction, especially with a large number of animals.
Data analysis
Analysis involves extracting information from each trajectory to obtain the behavioral data required by the experiment. This extracted data consists of statistics related to moving velocity, distance, animal orientation, rate of explored territory or activity rate. Data analysis is usually a post-processing step done after all other tracking tasks are completed.
Results and discussion
To compare how the 28 selected free programs handle the different steps in the tracking pipeline, we analyze and evaluate their main characteristics and functionalities for each step. Because there are no public datasets and standard metrics, we cannot make a direct quantitative comparison of their performance. In addition, these programs have been developed to use different video files, are optimized for specific arenas and are aimed at tracking different animals. Therefore, we point out their most important limitations, study the benefits and drawbacks of the techniques and algorithms, evaluate the areas that require further research and, finally, comment on the most important missing features of the software. We list these features in two tables that follow the tracking pipeline structure (Figs. 3 and 4) and discuss each step below.
Red: functionality or characteristic not present or lacking when compared to the best software for the evaluated characteristic. Light gray: limited functionality or characteristic when compared to the best software for the evaluated characteristic. *User-friendliness: +, bad (programming or high-level domain-specific knowledge is required, or the software needs to be compiled, has complex installation requirements or is difficult to use); ++, OK (requires relevant domain-specific knowledge to install and run; it is suitable for laboratories or academia); +++, good (the software is easy to install and run, and it is suitable for domestic use). CNN, convolutional neural network; GPU, graphics processing unit; MCR, MATLAB Compiler Runtime; NA, not available; OS, operating system.
Red: functionality or characteristic not present or lacking when compared to the best software for the evaluated characteristic. Light gray: limited functionality or characteristic when compared to the best software for the evaluated characteristic. *Multiple animals: Minimum–maximum number of animals that can be tracked; in parentheses, the time the software is able to preserve their identity according to the results published by their authors.
Video acquisition
Behavioral experiments are often long and require a relatively large sample size; thus, they usually generate a vast amount of video data. The quality of the video data and, therefore, the results extracted from it are heavily dependent on experimental conditions such as illumination, camera position, camera resolution, optical quality and background color. The influence of these parameters in the behavioral results has rarely been studied, and most researchers are not familiar with the variables that need to be considered or how to adjust them20.
It is also important to note that parameters such as resolution, codec configuration or frame rate can affect the computational cost of tracking algorithms exponentially. It is recommended that each animal is composed of at least 50 pixels in a video21,22 and that the frame rate is high enough that the animal position overlaps in two consecutive frames. However, most tracking applications are limited by the maximum resolution that they can handle. For example, EthoWatcher23,24 is restricted to a maximum video resolution of 320 × 240 pixels, and MouseMove25 can handle resolutions only up to 640 × 480 pixels22. Thus, these applications are somewhat limited in the field of view and the number of organisms that they can track if one wants to fulfill the requirements presented above. To our knowledge, only anTraX26, Automated Planar Tracking27 and ToxTrac22,28,29 have been successfully tested with resolutions higher than 1,920 × 1,080. Other programs such as Idtracker21,30 and Idtracker.ai31,32 or Ctrax33,34 are, in our experience, also able to handle videos with such resolutions.
The frame rate or the number of frames in a video file is also a computational limiting factor. Because Idtracker21,30 and Idtracker.ai31,32 require high computation times for each frame, even on moderate resolutions, analyzing large data sets using these software programs can be very time consuming. Finally, ABC tracker35,36 can handle only videos shorter than 10 min, restricting the use of this software to short-time experiments37.
The most important limitation of video acquisition is the processing speed and memory required to run tracking algorithms. This limitation is currently the bottleneck in behavioral experiments. The ability to process high-resolution, high-frame-rate videos in real time (online analysis) would revolutionize behavioral experiments by markedly decreasing the analysis timescale and reducing the need for video data storage. Currently, real-time tracking is possible only by using simple algorithms that do not work with complex backgrounds and multiple targets. Of the 28 applications, BioSense38,39, EthoWatcher23,24, MARGO40,41, Multi-Worm Tracker42,43 and SwisTrack44,45 offer both online and offline video acquisition modes, whereas the other programs only operate in offline mode.
Calibration
Camera calibration is a process that allows the user to obtain measurements in real-world coordinates. In addition, calibration can also include removing image distortion and perspective errors, which occur when imaging a 2D surface with a fixed camera; it is important to recall that the distance from the center to the edges is not linearly increasing. Although calibration is a critical feature of animal-tracking software to obtain reliable data, only 9 of the 28 analyzed programs offer a calibration function. See Fig. 3, ‘Calibration’ column.
Calibration techniques are commonly based on the use of the pinhole mathematical camera model46 to solve the equations that describe the projection of a point in the real world to the image plane through the lens of an ideal camera. This model takes into account not only the pixel scale but also the rotation of the camera with respect to the arena, also allowing the estimation and removal of lens distortion. This technique requires solving a complex equation system and using a calibration pattern. Only SwisTrack44,45, MARGO40,41 and ToxTrac22,28,29 implement this technique.
Most calibration techniques do not take advantage of the pinhole model and use a simple scale transformation. A scale transformation converts image coordinates to world coordinates by multiplying them by a constant factor. The programs Animapp47, BEMOVI48,49, BioSense38,39, EthoWatcher23,24, MARGO40,41, MouseMove25, SwisTrack44,45 and ToxTrac22,28,29 use this approach. This technique is more straightforward for the user but far less flexible and accurate becaue image distortion is not taken into account. Only SwisTrack44,45, ToxTrac22,28,29 and MARGO40,41 have both calibration systems.
The lack of calibration options in available animal-tracking software is surprising. We believe that this issue illustrates the deep gap between the considerations of the academic community that develops tracking software and the actual laboratory needs of software users.
Detection
The detection step consists of finding the animals of interest in the images. In Fig. 3, we divide the detection step into three different sections.
Background subtraction
Background subtraction algorithms aim to remove features of the environment that can interfere with animal detection. Background subtraction is a key feature when recording animals in a natural setting with dynamic lighting conditions or in aquatic environments where images are changed by reflections, shadows and other artifacts. The two most common types of background subtraction techniques are those based on static images and those based on dynamic models.
Static techniques commonly use a reference frame or video of the background without animals, such as in EthoWatcher23,24, Pathtrackr50,51, MARGO40,41, MouseMove25, SpectralTL52 and SwisTrack44,45, or estimate a background by averaging the frames of the video, such as in anTraX26, Biotracker53, Idtracker21,30 and Idtracker.ai31,32. Static background techniques are easy to implement and are effective when detecting stationary or moving animals, if the background objects and illumination do not change during the experiment. Otherwise, these techniques should not be used.
Dynamic techniques use moving or decaying average models, such as in Multi-Worm Tracker42,43 and SwisTrack44,45, or Gaussian mixture models, such as in BioSense38,39, Ctrax33,34, ToxTrac22,28,29 and UMATracker54,55. Dynamic techniques can account for illumination changes or other gradual changes in the background. This factor is important when running long-time experiments in which the sun is used as a light source. However, these techniques are not reliable when detecting animals that remain static during a substantial part of the experiment.
The only program that approaches background subtraction with an innovative technique is Mouse Tracking56,57. Mouse Tracking56,57 uses a deep learning algorithm to separate the pixels from the background and foreground. This strategy is more robust than other techniques but is also complex and requires a massive amount of training data, which makes the method very computationally heavy.
From our experience, there is room for improvement in background subtraction algorithms given that only a few studies have addressed situations with low and/or changing contrast within the background.
Segmentation
Segmentation is performed immediately after background subtraction and usually consists of a technique aimed at separating potential animals in the image and a filtering step that removes possible false positives. The most common segmentation technique is based on so-called ‘thresholding’. Thresholding is a simple segmentation method that uses a reference value to separate pixel regions of the image of different brightness. Thresholding is based on contrast and requires the animals to appear as bright objects in a dark background or as dark objects in a bright background. The main advantage of thresholding is that it is a very computationally efficient technique. However, thresholding is very sensitive to false-positives or false-negatives in non-uniform images. Most of the analyzed software use a variation of this technique.
To increase the sensitivity and robustness of detection, a few techniques locate specific animal features on the image. This method limits the thresholding step because it can be applied only to specific animals with certain body shapes and therefore can not be used for general tracking. Examples of software using this strategy are Ctrax33,34 and Mouse Tracking56,57, which use an ellipse fitting strategy to search for circular shaped objects. Automated Planar Tracking27 and Fish CnnTracker58, on the other hand, rely on locating the specific shape of the fish head to improve detection.
The only applications that provide some innovations in animal detection are ABC Tracker35,36, Multi-Animal Tracker59,60 and Pathtrackr50,51. These programs use a system that requires the user to mark each animal’s location in a few frames on the video and use a machine learning technique to locate each animal in the remaining frames. This approach provides a more robust detection system than other algorithms. ABC Tracker35,36 successfully uses this approach by applying support vector machines and obtains robust results with a user-friendly experience.
Species
Most of the software studied are versatile and can be used for different species. Of the 28 tested applications, 16 can be used for tracking any type of animal, whereas 11 are designed for a specific animal type, and 1 requires expanding the software with specific animal models (see column ‘Species’ in Fig. 3).
Trajectory generation
The challenge of trajectory generation is to associate potential targets with previous trajectories (i.e., to associate a set of detections to a group of animals, where we know the trajectories of these animals before the current frame). The most common technique to solve this problem uses the Kalman filter, which is a prediction-correction technique61. With this technique, one can estimate an animal’s position in the next frame on the basis of its previous known positions by assuming a constant speed or a constant acceleration model. Then, in the next frame, the predicted positions are compared with the actual detections using a Hungarian optimization technique62. The Kalman filter is very efficient computationally, and most tracking programs that we are aware of have implemented a variation of this algorithm or use similar techniques (e.g., Particle Filters63).
However, the Kalman filter and other similar techniques are not reliable in animal-tracking scenarios where occlusions or multiple interacting animals exist. The reason for this limitation is that these techniques are not able to keep the identity of the objects and use only spatial information to match the trajectories. A typical example of a situation where these techniques fail is when two animals cross paths and change direction after the collision. When this situation occurs, the algorithm will lose track of the animals for a brief moment. It will then search for the animals’ new positions, assuming they continued moving in the same direction. As a consequence, animals’ identities will be switched.
Preserving the identity of multiple individuals (id. preservation) after an occlusion is currently the main limitation in the trajectory-generation step. The complexity of this problem is illustrated in a 2014 study21, in which Pérez-Escudero and colleagues analyzed a scenario with multiple interacting animals. In this scenario, when correctly solving 99% of all crossings, only 11% of the animals were correctly identified after 2 min of tracking, owing to error propagation22. In summary, preserving the identity is complex and computationally expensive, and only a few offline tracking applications offer major contributions to this field.
Motr64,65 uses an id. preservation that relies on marking the individuals with visually distinctive markers that can be easily identified automatically. This technique is reliable and allows the tracking of animals for long periods of time and in multiple sessions. However, many modern techniques try to avoid placing markers or sensors on the animals because the markers can be impractical and sometimes interfere with the experiment by affecting animal behavior66,67. Programs such as Automated Planar Tracking27, Tracktor19,68 and UMATracker54,55 offer id. preservation algorithms that, in our opinion, have only marginally improved the basic Kalman strategy and are not robust and reliable for some specific situations (Fig. 4).
Idtracker21,30 and ToxTrac22,28,29 use a strategy based on a probabilistic texture analysis to analyze animals’ similarity between collisions. Idtracker21,30 was one of the first applications that seriously approached this issue when tracking multiple targets and uses a complex algorithm based on a Bayesian analysis with a similarity metric to compare the objects’ texture. ToxTrac22,28,29, on the other hand, uses a combination of a similarity analysis with a Hungarian algorithm to manage the identity preservation of multiple targets. This technique builds on top of a very fast tracking algorithm that can handle simultaneous tracking in multiple arenas, resulting in one of the most flexible free tracking tools for trajectory generation.
Traditional probabilistic texture analysis is not capable of tracking many targets or for doing so for very long times, but the approach is very useful in short experiments with small groups of animals, for which body shape and appearance do not change much in comparison to their position or posture. On the basis of the results reported by ToxTrac22,28,29 and Idtracker21,30, we recommend using these programs with groups of up to five animals in videos no longer than 20 min. Between these two techniques, especially for users with limited computational speed, we think that ToxTrac22,28,29 is a better alternative because it requires substantially less processing time19,28.
Idtracker.ai31,32 and Fish CnnTracker58 use an approach based on deep learning models called ‘convolutional neural networks’ (CNNs). CNNs are optimized for image classification tasks and are among the most powerful image classification techniques nowadays, outperforming those based on traditional probabilistic texture analysis. The current drawbacks of these models are that they require specific training data and cannot be used in real-time applications even with optimized hardware, such as graphics processing units (GPUs). Although some techniques such as transfer learning can mitigate some of these issues, none of the software in the list use this approach.
Idtracker.ai31,32, which is an extension of Idtracker21,30, combines two different deep learning algorithms: one to detect occlusions and one to identify targets with classification analysis. Idtracker21,30 and Idtracker.ai31,32 present the most solid identity-preservation techniques on the market. However, the computational time required to analyze a standard experiment can be as high as 1 h per frame if not using expensive GPU computing hardware, making it impractical to run on a standard laboratory computer. Fish CnnTracker58 offers a less flexible approach only suitable for fish tracking and is also less accurate28 than Idtracker.ai31,32. Idtracker.ai31,32 achieves the best accuracy in id. preservation to date. In a set of experiments, Romero-Ferrero et al.31,32 manually reviewed a significant sample of the crossings for individual animals, and Idtracker.ai was able to track up to 100 zebrafish and fruit flies for 10 min or 4 mice for 1 h31. However, in our opinion, the computation times this technique requires make Idtracker.ai31,32 not suitable for most common scenarios, given that many behavioral laboratories do not have a high-end computer with a state of the art GPU and that processing a 1-h video with Idtracker.ai31,32, with a modern GPU, can take >1 d of computation in extreme situations32. Despite this limitation, it is important to note that Idtracker.ai31,32 is the best of the reviewed software to analyze experiments with large groups of unmarked animals.
ABC Tracker35,36 offers a novel approach that uses forward and backward particle filtering. A particle filter is a selection-prediction-measurement solution of similar complexity to the Kalman filter. The particle filter uses a set of samples called particles to estimate the internal states in dynamical systems from partial observations and with random perturbations. This filter is also easy to parallelize and can be more accurate than a Kalman filter69. In our opinion, the main novelty of ABC Tracker35,36 is that it combines with this prediction scheme an algorithm to track stationary objects on the basis of a local search strategy that, in practice, is able to solve most animal interactions when the scenario does not involve complex occlusions in a 3D space. ABC Tracker35,36 can currently track up to 30 animals in a video for <10 min37, obtaining very good results in these situations. In our opinion, ABC Tracker35,36 also has the most intuitive interface for analyzing experiments with multiple individuals.
Finally, anTraX26 combines the use of color tags to mark individuals, CNNs and a graph-based approach. According to the authors, these features should allow the tracking of dozens of marked individuals for hours, if not days. However, given the use of tags and the lack of validation results for specific times, it is unclear how robust the algorithm is compared to other techniques.
In summary, id. preservation algorithms for multiple interacting animals are still insufficient when looking at the computational performance and accuracy that are required for behavioral tests and analysis. So far, id. preservation has not been accomplished when running multiple arenas simultaneously, with animal sizes smaller than 50 pixels or with online image-acquisition techniques.
Data analysis
Nowadays, automatic behavior recognition can be achieved using annotated video datasets to train machine-learning classifiers70. Kabra et al.71 proposed an automatic animal behavior annotator that led to the creation of individual and social behavior classifiers for organisms, such as mice and larval flies. Robie et al.18 proposed a similar strategy to detect patterns of social interactions. Using these techniques, some researchers studied complex behaviors such as mating and feeding in mice72 and behavioral responses of larval fish to chemicals73. Despite this development, only a minority of the analyzed software include functions for complex behavior recognition, and they are always limited to specific behaviors in particular scenarios. For example, Multi-Animal Tracker59,60 detects pirouette movements in worms, Ctrax33,34 detects touch and chase social behaviors in flies, QTrack12,74 also detects specific courtship behaviors in flies and MouseMove25 can quantify unilateral locomotor deficits in mice.
In general, tracking applications provide only movement metrics. Thus, most tracking applications are limited to movement, orientation and zone exploration metrics. Furthermore, only a fraction of the assessed software in this review provides a useful array of these metrics (i.e., BioSense38,39, MouseMove25, Multi-Animal Tracker59,60, Multi-Worm Tracker42,43 and ToxTrac22,28,29). These software provide advanced toolkits that allow non-programmers to analyze parameters such as movement, time spent in selected areas, changes in direction and time spent moving. An important note is that these software packages can also provide individual or population metrics.
We conclude that more work is needed to integrate behavior recognition in tracking software, and we believe that algorithms for automatic detection of stress and other complex behaviors would represent a true innovation if included in tracking tools.
Extra features
Some tracking applications implement extra features to facilitate user experience or to add versatility. The most useful extra feature, from our point of view, is the ability to analyze video files that have been split into multiple files. This feature is explicitly supported by Idtracker21,30, Idtracker.ai31,32 and ToxTrac22,28,29. Another important feature is the possibility of processing a batch of video files by using the same camera configuration, allowing the user to adjust the parameters only once for a set of experiments. Only Animapp47, Ctrax33,34, idTracker21,30, MARGO40,41, MouseMove25 and ToxTrac22,28,29 implement this functionality.
The possibility of controlling hardware peripherals such as external sensors, lights or temperature sources is a nice feature implemented in MARGO40,41 and RAT75,76. This feature allows measuring parameters such as reaction times to stimuli, planning long-term tests without supervision and modifying the stimuli according to behaviors, thereby creating a closed control loop.
UMATracker54,55 and BioTracker53 implement a modular approach to facilitate the development and integration of new processing modules. We think that this addition can be useful in collaboration with the community to integrate new features into the software. However, taking full advantage of this feature requires a constant level of support and commitment that may not be realistic.
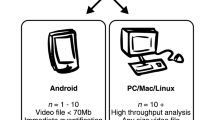
Finally, we highlight that Animapp47 includes an Android application that directly analyzes images using a smartphone camera. With the increased computational power of smartphones, we believe that this type of program can be useful for simple studies performed in a field environment.
Conclusions
Current tracking software need to balance robustness, accuracy and processing speed. As a rule of thumb, higher robustness and accuracy require complex algorithms that reduce processing speed. Therefore, online processing programs use simple algorithms that increase efficiency to reach real-time performance, but with the trade-off of less robustness and accuracy. Offline software can take advantage of more complex processing algorithms but require a computational capacity that is not achievable for all users or suitable for every experiment. We tried to highlight these differences and the contributions of each application to the animal-tracking field. (BioTrack77,78 and FIMTrack79,80 are not discussed specifically in the text but are included in Figs. 3 and 4.)
Our assessment shows that all applications share more or less the same pipeline and that very few of them offer a unique or revolutionary approach, with the use of CNN networks for id. preservation being the most relevant new contribution. However, our biggest concern is the lack of usability of recent software. Of the 28 tested programs, only four (IdTracker21,30, IdTracker.ai31,32, ABC Tracker35,36 and Toxtrac22,28,29) provide innovative algorithms, useful features and user-friendly interfaces. We believe that the main reason for this lack of usability is the existence of a gap between software design and their intended use in a laboratory. Most software packages are not easy to use, require tuning of several complex parameters for each experiment and do not include important features such as calibration options. In addition, most programs do not offer data analysis tools beyond the most basic ones and cannot extract valuable behavioral metrics. In our opinion, developers of tracking tools must change their paradigm from creating programs that can be published to creating programs that are useful and easy to use.
Finally, we want to draw attention to the lack of complete, open and well-labeled datasets that provide a standard reference for validation and accuracy testing. Such datasets would give researchers an objective tool for a quantitative comparison of tracking programs.
References
Dell, A. I. et al. Automated image-based tracking and its application in ecology. Trends Ecol. Evol. 29, 417–428 (2014).
Hajar, R. Animal testing and medicine. Heart. Views 12, 42 (2011).
Carola, V., D’Olimpio, F., Brunamonti, E., Mangia, F. & Renzi, P. Evaluation of the elevated plus-maze and open-field tests for the assessment of anxiety-related behaviour in inbred mice. Behav. Brain Res. 134, 49–57 (2002).
Olton, D. S. Mazes, maps, and memory. Am. Psychol. 34, 583–596 (1979).
Silverman, J. L., Babineau, B. A., Oliver, C. F., Karras, M. N. & Crawley, J. N. Influence of stimulant-induced hyperactivity on social approach in the BTBR mouse model of autism. Neuropharmacology 68, 210–222 (2013).
Cirulli, F., Berry, A. & Alleva, E. Intracerebroventricular administration of brain-derived neurotrophic factor in adult rats affects analgesia and spontaneous behaviour but not memory retention in a Morris Water Maze task. Neurosci. Lett. 287, 207–210 (2000).
Borta, A. & Schwarting, R. K. W. Inhibitory avoidance, pain reactivity, and plus-maze behavior in Wistar rats with high versus low rearing activity. Physiol. Behav. 84, 387–396 (2005).
Kulesskaya, N. & Voikar, V. Assessment of mouse anxiety-like behavior in the light–dark box and open-field arena: role of equipment and procedure. Physiol. Behav. 133, 30–38 (2014).
Lee, H., Iida, T., Mizuno, A., Suzuki, M. & Caterina, M. J. Altered thermal selection behavior in mice lacking transient receptor potential vanilloid 4. J. Neurosci. 25, 1304–1310 (2005).
Woodley, C. M., Urbanczyk, A. C., Smith, D. L. & Lemasson, B. H. Integrating visual psychophysical assays within a Y-Maze to isolate the role that visual features play in navigational decisions. J. Vis. Exp. 147, e59281 (2019).
Jonsson, M. et al. High-speed imaging reveals how antihistamine exposure affects escape behaviours in aquatic insect prey. Sci. Total Environ. 648, 1257–1262 (2019).
Dankert, H., Wang, L., Hoopfer, E. D., Anderson, D. J. & Perona, P. Automated monitoring and analysis of social behavior in Drosophila. Nat. Methods 6, 297–303 (2009).
Pereira, T. D. et al. Fast animal pose estimation using deep neural networks. Nat. Methods 16, 117–125 (2019).
Graving, J. M. et al. DeepPoseKit, a software toolkit for fast and robust animal pose estimation using deep learning. eLife 8, e47994 (2019).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Franco-Restrepo, J. E., Forero, D. A. & Vargas, R. A. A review of freely available, open-source software for the automated analysis of the behavior of adult zebrafish. Zebrafish 16, 223–232 (2019).
Husson, S. J., Costa, W. S., Schmitt, C. & Gottschalk, A. Keeping track of worm trackers. in WormBook: The Online Review of C. elegans Biology (WormBook, 2018).
Robie, A. A., Seagraves, K. M., Egnor, S. E. R. & Branson, K. Machine vision methods for analyzing social interactions. J. Exp. Biol. 220, 25–34 (2017).
Sridhar, V. H., Roche, D. G. & Gingins, S. Tracktor: image-based automated tracking of animal movement and behaviour. Methods Ecol. Evol. 10, 815–820 (2018).
Henry, J., Rodriguez, A. & Wlodkowic, D. Impact of digital video analytics on accuracy of chemobehavioural phenotyping in aquatic toxicology. PeerJ 7, e7367 (2019).
Pérez-Escudero, A., Vicente-Page, J., Hinz, R. C., Arganda, S. & de Polavieja, G. G. idTracker: tracking individuals in a group by automatic identification of unmarked animals. Nat. Methods 11, 743–748 (2014).
Rodriguez, A. et al. ToxTrac: a fast and robust software for tracking organisms. Methods Ecol. Evol. 9, 460–464 (2018).
Junior, C. F. C. et al. ETHOWATCHER: validation of a tool for behavioral and video-tracking analysis in laboratory animals. Comput. Biol. Med. 42, 257–264 (2012).
Crispim Junior, C. F. et al. EthoWatcher. http://ethowatcher.paginas.ufsc.br/ (2019).
Samson, A. L. et al. MouseMove: an open source program for semi-automated analysis of movement and cognitive testing in rodents. Sci. Rep. 5, 16171 (2015).
Gal, A., Saragosti, J. & Kronauer, D. J. C. anTraX, a software package for high-throughput video tracking of color-tagged insects. eLife 9, e58145 (2020).
Wang, S. H., Cheng, X. E., Qian, Z.-M., Liu, Y. & Chen, Y. Q. Automated planar tracking the waving bodies of multiple zebrafish swimming in shallow water. PLoS ONE 11, e0154714 (2016).
Rodriguez, A., Zhang, H., Klaminder, J., Brodin, T. & Andersson, M. ToxId: an efficient algorithm to solve occlusions when tracking multiple animals. Sci. Rep. 7, 14774 (2017).
Rodriguez, A. et al. ToxTrac. https://sourceforge.net/projects/toxtrac/ (2019).
Pérez-Escudero, A., Vicente-Page, J., Hinz, R. C., Arganda, S. & de Polavieja, G. G. idTracker. http://www.idtracker.es/ (2019).
Romero-Ferrero, F., Bergomi, M. G., Hinz, R., Heras, F. J. H. & de Polavieja, G. G. idtracker.ai: tracking all individuals in large collectives of unmarked animals. Nat. Methods 16, 179–182 (2019).
Romero-Ferrero, F., Bergomi, M. G., Hinz, R., Heras, F. J. H. & de Polavieja, G. G. idtracker.ai. https://idtracker.ai/ (2019).
Branson, K., Robie, A. A., Bender, J., Perona, P. & Dickinson, M. H. High-throughput ethomics in large groups of Drosophila. Nat. Methods 6, 451–457 (2009).
Branson, K., Robie, A. A., Bender, J., Perona, P. & Dickinson, M. H. Ctrax: the Caltech multiple walking fly tracker. http://ctrax.sourceforge.net/ (2019).
Shin, M. C. ABCTracker. http://abctracker.org/ (2018).
Rice, L. A. A beginning-to-end system for efficiently gathering tracking data on multiple targets. Thesis, University of North Carolina at Charlotte (2016).
Farynyk, D. ABC Tracker Support. https://abctracker.atlassian.net/wiki/spaces/ABCTS/pages/458795/FAQ (2020).
Patman, J., Michael, S. C. J., Lutnesky, M. M. F. & Palaniappan, K. BioSense: real-time object tracking for animal movement and behavior research. IEEE Applied Imagery Pattern Recognition Workshop (AIPR) 1–8 (2018).
Patman, J., Michael, S. C. J., Lutnesky, M. M. F. & Palaniappan, K. BioSense. https://ieeexplore.ieee.org/document/8707411 (2018).
Werkhoven, Z., Rohrsen, C., Qin, C., Brembs, B. & de Bivort, B. MARGO (Massively Automated Real-time GUI for Object-tracking), a platform for high-throughput ethology. PLoS ONE 14, e0224243 (2019).
Werkhoven, Z., Rohrsen, C., Qin, C., Brembs, B. & de Bivort, B. MARGO. https://github.com/de-Bivort-Lab/margo (2020).
Swierczek, N. A., Giles, A. C., Rankin, C. H. & Kerr, R. A. High-throughput behavioral analysis in C. elegans. Nat. Methods 8, 592–598 (2011).
Swierczek, N. A., Giles, A. C., Rankin, C. H. & Kerr, R. A. Multi-Worm Tracker. https://sourceforge.net/projects/mwt/ (2019).
Correll, N. et al. SwisTrack: a tracking tool for multi-unit robotic and biological systems. IEEE/RSJ International Conference on Intelligent Robots and Systems 2185–2191 (2006).
Mario, E. Di et al. SwisTrack. https://sourceforge.net/projects/swistrack (2019).
Zhang, Z. A flexible new technique for camera calibration. IEEE Trans. Pattern Anal. Mach. Intell. 22, 1330–1334 (2000).
Rao, S. R. et al. Small animal video tracking for activity and path analysis using a novel open-source multi-platform application (AnimApp). Sci. Rep. 9, 12343 (2019).
Pennekamp, F., Schtickzelle, N. & Petchey, O. L. BEMOVI, software for extracting behavior and morphology from videos, illustrated with analyses of microbes. Ecol. Evol. 5, 2584–2595 (2015).
Pennekamp, F., Schtickzelle, N. & Petchey, O. L. BEMOVI. http://bemovi.info/ (2015).
Harmer, A. M. T. & Thomas, D. B. Pathtrackr: an r package for video tracking and analysing animal movement. Methods Ecol. Evol. 10, 1196–1202 (2019).
Harmer, A. M. T. & Thomas, D. B. Pathtrackr. https://github.com/aharmer/pathtrackr (2019).
Madan, C. R. & Spetch, M. L. Visualizing and quantifying movement from pre-recorded videos: the spectral time-lapse (STL) algorithm. F1000Res. 3, 19 (2014).
Mönck, H. J. et al. BioTracker: an open-source computer vision framework for visual animal tracking. Preprint at https://arxiv.org/abs/1803.07985 (2019).
Yamanaka, O. & Takeuchi, R. UMATracker: an intuitive image-based tracking platform. J. Exp. Biol. 221, jeb182469 (2018).
Yamanaka, O. & Takeuchi, R. UMATracker. http://ymnk13.github.io/UMATracker/ (2019).
Geuther, B. Q. et al. Robust mouse tracking in complex environments using neural networks. Commun. Biol. 2, 124 (2019).
Geuther, B. Q. et al. MouseTracking. https://github.com/KumarLabJax/MouseTracking (2019).
Zhiping, X. U. & Cheng, X. E. Zebrafish tracking using convolutional neural networks. Sci. Rep. 7, 42815 (2017).
Itskovits, E., Levine, A., Cohen, E. & Zaslaver, A. A multi-animal tracker for studying complex behaviors. BMC Biol. 15, 29 (2017).
Itskovits, E., Levine, A., Cohen, E. & Zaslaver, A. Multi-Animal Tracker. https://github.com/itskov/MultiAnimalTrackerSuite (2019).
Cuevas, E. V., Zaldivar, D. & Rojas, R. Kalman filter for vision tracking. Freie Universität Berlin, Fachbereich Mathematik und Informatik; Serie B, Informatik (2005).
Kuhn, H. W. The Hungarian method for the assignment problem. Nav. Res. Logist. Q. 2, 83–97 (1955).
Del Moral, P. Nonlinear filtering: interacting particle resolution. C R Acad. Sci. I 325, 653–658 (1997).
Ohayon, S., Avni, O., Taylor, A. L., Perona, P. & Egnor, S. E. R. Automated multi-day tracking of marked mice for the analysis of social behaviour. J. Neurosci. Methods 219, 10–19 (2013).
Ohayon, S., Avni, O., Taylor, A. L., Perona, P. & Egnor, S. E. R. motr: a MOuse TRacker. http://motr.janelia.org/ (2019).
Rodriguez, A. et al. Optical fish trajectory measurement in fishways through computer vision and artificial neural networks. J. Comput. Civ. Eng. 25, 291–301 (2011).
Rodriguez, A., Bermúdez, M., Rabuñal, J. & Puertas, J. Fish tracking in vertical slot fishways using computer vision techniques. J. Hydroinformatics 17, 275–292 (2014).
Sridhar, V. H., Roche, D. G. & Gingins, S. Tracktor. https://github.com/vivekhsridhar/tracktor (2019).
Rao, G. M. & Satyanarayana, C. Visual object target tracking using particle filter: a survey. Int. J. Image Graph Signal Process. 6, 57–71 (2013).
Datta, S. R. Q&A: understanding the composition of behavior. BMC Biol. 17, 44 (2019).
Kabra, M., Robie, A. A., Rivera-Alba, M., Branson, S. & Branson, K. JAABA: interactive machine learning for automatic annotation of animal behavior. Nat. Methods 10, 64–67 (2013).
Linck, V. M. et al. Effects of inhaled Linalool in anxiety, social interaction and aggressive behavior in mice. Phytomedicine 17, 679–683 (2010).
Steele, W. B., Mole, R. A. & Brooks, B. W. Experimental protocol for examining behavioral response profiles in larval fish: application to the neuro-stimulant caffeine. J. Vis. Exp. 137, e57938 (2018).
Dankert, H. QTrack (Cadabra). http://www.vision.caltech.edu/cadabra/ (2009).
Krynitsky, J. et al. Rodent Arena Tracker (RAT): a machine vision rodent tracking camera and closed loop control system. eNeuro 7, ENEURO.0485-19.2020 (2020).
Krynitsky, J. et al. Rodent arena tracker (RAT). https://hackaday.io/project/162481-rodent-arena-tracker-rat (2020).
Feldman, A., Hybinette, M. & Balch, T. The multi-iterative closest point tracker: an online algorithm for tracking multiple interacting targets. J. Field Robot. 29, 258–276 (2012).
Hrolenok, B., Quitmeyer, A., Motter, S., Stolarsky, D. & Migliozzi, B. L. R. Bio-Tracking. http://www.bio-tracking.org/ (2012).
Risse, B., Berh, D., Otto, N., Klämbt, C. & Jiang, X. FIMTrack: an open source tracking and locomotion analysis software for small animals. PLoS Comput. Biol. 13, e1005530 (2017).
Risse, B., Berh, D., Otto, N., Klämbt, C. & Jiang, X. FIMTrack. https://www.uni-muenster.de/Informatik.AGRisse/media/fim-media.html (2017).
Author information
Authors and Affiliations
Contributions
A.R. co-wrote the manuscript and performed most of the analysis of the tracking software. V.P. co-wrote the manuscript and assisted with the analysis of the tracking software. J.H., D.W. and M.A. revised and edited the manuscript and assisted with the analysis of the tracking software.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Peer review information Lab Animal thanks Waseem Abbas, Alfonso Perez-Escudero and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Panadeiro, V., Rodriguez, A., Henry, J. et al. A review of 28 free animal-tracking software applications: current features and limitations. Lab Anim 50, 246–254 (2021). https://doi.org/10.1038/s41684-021-00811-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41684-021-00811-1
- Springer Nature America, Inc.
This article is cited by
-
Metabolic plasticity drives mismatches in physiological traits between prey and predator
Communications Biology (2024)
-
Exploring the adaptive behaviour and environmental acclimation of artificially-bred Chinese sturgeon (Acipenser sinensis) in semi-open marine environment: insights for endangered species conservation
Reviews in Fish Biology and Fisheries (2024)
-
DVT: a high-throughput analysis pipeline for locomotion and social behavior in adult Drosophila melanogaster
Cell & Bioscience (2023)
-
Three-dimensional unsupervised probabilistic pose reconstruction (3D-UPPER) for freely moving animals
Scientific Reports (2023)
-
Beyond the three-chamber test: toward a multimodal and objective assessment of social behavior in rodents
Molecular Autism (2022)