Abstract
Hippophae tibetana, one of the highest-altitude woody plants endemic to the Qinghai-Tibet Plateau, primarily thrives on riverbanks formed by glacial meltwater. As a dioecious species, it demonstrates significant ecological and economic value in extreme alpine environments. However, the lack of sex identification techniques outside of the flowering period severely limits research on sex ratio, differentiation, and breeding. There is an urgent need to develop effective sex-linked molecular markers that are independent of developmental stages, but current research in this area remains limited. This study developed a set of accurate sex-linked molecular markers for the rapid identification of male and female individuals of H. tibetana. Through whole-genome resequencing of 32 sexually differentiated H. tibetana samples, this study offers strong evidence supporting chromosome 2 as the sex chromosome and successfully identified key loci related to sex determination on this chromosome. Utilizing these loci, we, for the first time, developed three reliable pairs of sex-specific molecular markers, which exhibited high accuracy during validation across various geographic populations, offering an effective tool for the sex identification of H. tibetana. Additionally, this study lays the groundwork for further research into the mechanisms of sex determination and the evolution of sex chromosomes in H. tibetana.
Similar content being viewed by others
Introduction
Hippophae tibetana Schltdl, belonging to the family Elaeagnaceae and the genus Hippophae, is endemic to the Qinghai-Tibet Plateau1. This species naturally occurs in high-altitude regions of Gansu, Qinghai, Tibet, and Sichuan provinces in China2. Its altitude range spans from 2800 to 5200 m, making it the highest-altitude species within the genus Hippophae1,3. H. tibetana is dioecious, reproducing primarily through seeds and clonal propagation3. This species is noted for its exceptional clonal propagation ability and nitrogen-fixing capacity, playing a significant role as a pioneer plant in ecological restoration of riverbanks, grasslands, and bare lands following glacier retreat4. Moreover, H. tibetana is a dual-purpose plant, valuable for its medicinal, edible, and oil extraction properties5,6,7.
As the ecological and economic value of H. tibetana becomes increasingly recognized, this plant is attracting more attention. However, existing research primarily focuses on its chemical composition8, pharmacological effects5,9, phylogeography1,10, and genomics3,11, with relatively few studies on its sex differentiation. Studying the adaptive evolution of male and female plants of H. tibetana, a dioecious and clonal species distributed in the high-altitude regions of the Qinghai-Tibet Plateau, is of great significance. Nevertheless, before flowering, it is challenging to distinguish male and female individuals of H. tibetana based on phenotypic characteristics. Additionally, due to the small inflorescences and short flowering period, it is not easy to quickly identify the sex even during the flowering phase12. This poses a challenge to studying the adaptive evolution of male and female plants, highlighting the urgent need for a method that can accurately identify the sex of H. tibetana without being constrained by time. In recent years, molecular marker technology has been widely used for sex identification in various dioecious plants, such as Actinidia arguta (Siebold & Zucc.) Planch. ex Miq13, Spinacia oleracea L14, and Carica papaya L15. Although some sex-specific molecular markers have been developed for different species of Hippophae16,17,18,19,20, their applicability is often limited to specific geographic regions or varieties, lacking universal applicability and reliability21. This not only poses challenges for sex identification but also hinders the progress of in-depth research on the mechanisms of sex determination in Hippophae.
Although H. tibetana is diploid22, its sex determination system has not been accurately defined. Wang et al.3 aligned the RAD-seq data of male and female H. tibetana to the high-quality reference genome of the species. They identified that the 100 kb interval at positions 52,850,000–52,950,000 on chromosome 2 might be related to sex regulation or differentiation. However, because of the limitations of sequencing data, they could not identify reliable sex determination loci. With the advancement of high-throughput sequencing technology, whole-genome data analysis has become a commonly employed means for examining sex chromosomes and determining sex in nonmodel organisms13,23. This technology is particularly important for woody plants that require a long time to reach sexual maturity and flower for sex differentiation. It allows for the identification of sex before flowering, thereby optimizing the male-to-female ratio in breeding programs and reducing cultivation costs. Obtaining individual genome sequence information through high-throughput sequencing and aligning it with the reference genome can reveal single-nucleotide polymorphisms (SNPs) and insertions/deletions (InDels) in the whole-genome24, potentially allowing for the precise identification of genetic differences between male and female individuals.
In this study, utilizing the whole-genome data of H. tibetana published by Wang et al.3, we conducted whole-genome resequencing of 32 sexually differentiated H. tibetana individuals. We confirmed the sex chromosomes of H. tibetana and identified sex-linked molecular markers that can aid in differentiating between male and female plants.
Materials and methods
Sample collection and sequencing
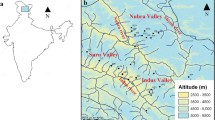
In June 2023, we collected samples from male and female H. tibetana plants in the field from six populations. The sex of the plants was determined on the basis of the morphological characteristics of their flowers. From each population, four female and four male plants were randomly selected for fresh leaf sampling (refer to Fig. 1 for sampling locations). To represent a broader genetic diversity and avoid collecting samples from plants of the same maternal line, we ensured a minimum distance of 100 m between sampled individuals of the same sex. The voucher specimens of male and female H. tibetana plants are preserved in the Key Laboratory of Biodiversity and Environment on the Qinghai-Tibetan Plateau, Ministry of Education, School of Ecology and Environment, Tibet University, Lhasa 850,000, China, under the voucher number LQ202306013. We extracted total DNA from the samples by using the modified CTAB method25 and assessed DNA quality and concentration through 1.0% agarose gel electrophoresis and the NanoDrop 2000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). DNA samples that met the quality criteria were sent to Novogene Co., Ltd. (Beijing, China) for DNA library construction and then sequenced on the Illumina NovaSeq X Plus with PE150 at a sequencing depth of 30 ×. Each sample generated approximately 47 GB of raw data (Supplementary Table S1).
Sampling locations of the six H. tibetana populations in Tibet. DR: Dingri County, Tibet; DX: Dangxiong County, Tibet; MZGK: Mozhugongka County, Tibet; BS: Basu County, Tibet. Generated using ggplot2 v. 3.3.5 and sf v. 1.0-3 packages in R v. 4.1.2. URL: https://ggplot2.tidyverse.org/.
Whole-genome population variation detection
The raw sequencing data were first subjected to quality control and cleaning using Fastp v.0.2126 with default parameters, and then assessed for quality with FastQC v.0.11.9 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). The cleaned data were aligned to the H. tibetana reference genome by using BWA v.2.2.127 and converted to BAM format with samtools v.1.1928. InDel identification and filtering were conducted using GATK4 v. 4.229. This process included removing PCR duplicates, detecting single-sample and population variants, and marking and filtering InDels and low-quality sites using the following criteria: “QD < 2.0||FS > 200.0||SOR > 10.0||MQRankSum < − 12.5||ReadPosRankSum < − 8.0.” Further filtering was performed using Plink v.1.930, excluding sites with a minor allele frequency of < 0.01 and missing genotype data exceeding 50%. Finally, 6,554,748 sites were selected from the initial 7,706,351 variant sites for further analysis.
Sex-determining region screening
We used the EMMAX (v intel64-20,120,205) software package31 to conduct a genome-wide association study (GWAS) on the filtered dataset using a mixed linear model (MLM). In this study, we treated male and female sexes as two independent traits to identify gene regions associated with sex determination. To ensure the accuracy of our analysis, we first calculated the kinship matrix using emmax-kin-intel64 and removed samples with a relatedness threshold greater than 0.125 (DX08, RD02, YBJ04, and YBJ14). The GWAS then focused particularly on regions with densely clustered significant variant sites and corresponding site information. Given that the reference genome was derived from a female plant, we focused on the alignment of sequencing reads from both female and male plants to the same site on the reference genome. A Python script (https://github.com/zengzhefei/MaleSpecificIndelFilter.git) was used to identify variant sites with InDels > 20 bp in the male samples. Finally, the selected sites were manually inspected using IGV v.2.1632, and sequences with long InDels in the male samples were identified as candidate regions for subsequent primer design.
Male-specific primer design
We used seqkit v.2.6.133 to extract sequence information corresponding to the identified variant sites from the reference genome for primer design. Initially, we focused on variant regions primarily 30–60 bp in length. To facilitate distinguishing male-specific bands, we aimed to design PCR products between 200 and 300 bp in length. The suitability of a variant region for primer development was determined by examining the distribution of nucleotide bases within approximately 200 bp flanking the variant site, ensuring an even distribution, as regions with excessive repeats hinder effective primer design. We then used primer3 v.2.3.734 to batch design the primers. During the designing of primers, we carefully avoided mismatches, dimers, hairpin structures, and primer dimers. The designed primers were subsequently aligned with the reference genome by using blastn v.2.5.035 to ensure their specificity. After this screening step, primers that demonstrated high specificity were selected and synthesized by Sangon Biotech (Shanghai) Co. Ltd.
Primer universality validation
To validate the specificity of the primers, we conducted conventional PCR amplification on a small set of H. tibetana DNA samples, including two females and two males, by using batch-synthesized primers. The PCR reaction comprised a total volume of 30 μL, including 15 μL of Taq Mix, 1 μL of DNA template, 1 μL each of forward and reverse primer, and 12 μL of deionized water. The PCR program started with an initial denaturation at 95 °C for 3 min, followed by 35 cycles of 95 °C for 15 s, 58 °C for 15 s, 72 °C for 30 s, and concluded with a final extension at 72 °C for 5 min before storage at 4 °C. The PCR products were analyzed using 2% agarose gel electrophoresis. Primers that effectively differentiated between the female and male samples based on product length were selected. These selected primers were then used to validate the accuracy on 48 samples (24 females and 24 males) collected from six H. tibetana populations in Dingri, Basu, Dangxiong, and Mozhugongka, Tibet.
Permission for sample collection
H. tibetana is not listed on the IUCN Red List of Threatened Species, and the collection area is not within a protected zone. Consequently, no specific permissions or licenses were required for sample collection from authorities.
Results
Sex-linked regions of H. tibetana
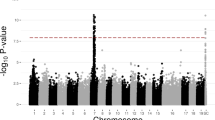
A GWAS was conducted using an MLM, treating male and female as contrasting traits to explore the correlation between sex traits and InDel sites. The analysis revealed that sex-related variations were primarily concentrated on chromosome 2, although a few loci differentially present in males and females were also found on other chromosomes (Fig. 2). Further screening identified 30 regions related to sex determination, with one specific region each on chromosomes 1 and 8, whereas the remaining specific regions were located on chromosome 2 (Supplementary Table S2).
Selection of sex-specific markers
After initial screening for long InDel sites within the sequencing reads of male H. tibetana, we identified 106 variant loci. Based on these findings, we designed 24 pairs of male-specific primers in bulk. After preliminary PCR amplification screening, three primer pairs were successfully obtained, enabling effective differentiation between male and female samples of H. tibetana (Table 1). The coordinates of the variant sites for these three primer pairs were visualized using IGV v.2.1632, which allowed us to observe the alignment of sequencing data with the reference genome. For instance, using the Hiti_05 primer pair, we determined that nearly half of the reads mapping to this site in male individuals included a 53 bp deletion. Both forward and reverse primers were situated in conserved sequences, enhancing the accuracy of amplification (Fig. 3).
Mapping of second-generation sequencing reads from one male and one female individual of H. tibetana on chromosome 2. Within the region outlined by the black dashed box, approximately 50% of reads from the male individual show a 53 bp deletion in the Hiti_05 primer-amplified region. The Sequences section displays the base sequence of the reference genome in the amplified region, with red, green, blue, and orange colors denoting the T, A, C, and G bases, respectively.
Specificity marker accuracy test
To further verify the accuracy of three primer pairs in distinguishing between male and female plants of H. tibetana, we employed the same PCR reaction system used for primer specificity validation. We tested 48 H. tibetana samples (24 female and 24 male) from six populations in Dingri, Basu, Muzhugongka, and Dangxiong, Tibet. The results confirmed that all three primer pairs could accurately differentiate between the tested male and female samples: two bands were produced in all male samples, whereas only one band appeared in female samples (Fig. 4). However, weak bands were observed in some female samples from the Dingri, Basu, Mozhugongka, and Dangxiong-3 populations when using the Hiti_05 primer (Fig. 4). To determine if this phenomenon was related to the annealing temperature, a gradient PCR experiment was conducted with a temperature range of 55–65 °C. The results indicated that male and female individuals could be clearly differentiated at temperatures between 58.8 and 61.1 °C. Outside this range, some female samples also produced faint bands, which could interfere with the differentiation between males and females (Supplementary Fig. S4).
PCR amplification results of 48 H. tibetana samples from different regions using the three pairs of primers (Hiti-05, Hiti-08, Hiti-17). The symbols ♀ and ♂ represent the female and male samples of H. tibetana, respectively. The original gel images generated by the three pairs of primers Hiti-05, Hiti-08, and Hiti-17 can be found in Supplementary Figs. S1, S2, and S3.
Discussion
H. tibetana plays a pivotal role in both ecological and economic sectors, especially in the ecologically sensitive region of the Tibetan Plateau. The plant is useful for ecological restoration and soil conservation. The fruits of H. tibetana are substantially enriched in nutritional components and bioactive compounds, making them valuable to the food, pharmaceutical, and cosmetic industries4,21. However, as a dioecious woody plant, identifying the sex of H. tibetana through phenotypes before flowering is challenging. It takes 3–4 years for the seeds to flower after germination, and the flowering period is brief, which complicates the development of related industries12,21. The sex determination system of H. tibetana remains unclear, limiting our understanding of its sex determination mechanism and hindering the advancement of sex identification techniques. Therefore, developing an accurate and efficient sex identification method is essential for advancing the H. tibetana industry and facilitating in-depth studies on its sex determination mechanism.
Puterova et al.36 studied the transposable elements and satellites in H. rhamnoides genome, concluding that its sex determination system is of the XY type, with the Y chromosome being shorter than the X chromosome. This raises the question of whether H. tibetana also follows a similar XY sex determination system. Based on the resequencing data analysis that clearly distinguishes male and female H. tibetana, this study found a significant number of variant sites in the sequencing reads of male samples aligned to the reference genome, further confirming that the sex determination mechanism of H. tibetana is of the XY type. To date, no efficient molecular marker exists for distinguishing between male and female plants of H. tibetana. Earlier studies, including the one by Korekar et al.17, identified specific RAPD primers that could amplify female samples of H. rhamnoides, but not male samples, leading to the development of SCAR markers, HrX1 and HrX2. However, subsequent studies, including the study by Chawla et al.18, have indicated that HrX1 could help differentiate between the male and female samples of H. tibetana, but our preliminary tests found that this marker did not produce specific amplification in the H. tibetana samples used in this study (Supplementary Fig. S5). Furthermore, we evaluated the sex-specific molecular markers developed by Das et al.19 for H. rhamnoides ssp. turkestanica to determine their applicability to H. tibetana. The results indicated that no bands were amplified, thus precluding sex differentiation (Supplementary Fig. S6). This finding indicates the complexity of sex chromosomes in H. tibetana and suggests that the effectiveness of designed markers may be limited to specific populations. Similar issues have been observed in other species, such as kiwifruit and persimmon37,38, where DNA markers sometimes did not co-segregate with sex-determining genes during meiosis, potentially leading to false positives or negatives.
In recent years, the rapid advancement of sequencing technologies and the assembly of high-quality reference genomes for numerous nonmodel species have opened new avenues for developing molecular markers for sex differentiation in dioecious plants39,40. Several studies have successfully designed molecular markers that can help distinguish between the sex of animals and plants using resequencing data combined with various analysis methods. For instance, She et al.14 analyzed the resequencing data of 10 individuals of S. oleracea (5 females and 5 males), identified potential male-specific regions, and designed primers accordingly. Guo et al.13 conducted pooled sequencing on 15 male and 15 female A. arguta, assembled male-specific kmers, and designed molecular markers. Luo et al.41 analyzed resequencing data from 15 male and 15 female Leiocassis longirostris Günther, identified regions associated with sex based on sex-specific SNPs, and developed highly accurate molecular markers.
This study involved sampling male and female individuals from six populations of H. tibetana to develop molecular markers for accurately identifying the male and female plants of this species. Our sampling strategy was influenced by the chloroplast trnT-trnF phylogeographic study conducted by Wang et al.3 and covered three genetic lineages across the eastern, central, and western regions. This approach was chosen to comprehensively reflect the genetic structure of H. tibetana and ensure the universality of the molecular markers developed. In terms of primer design, we adopted and enhanced the method proposed by Luo et al.41, utilizing next-generation sequencing, to focus on homologous fragments between male and female individuals that contained significant long fragment InDel sites in the male samples. This strategy facilitated the development of sex-specific markers. We successfully designed and validated three pairs of male-specific markers in H. tibetana, which demonstrated effective differentiation in 48 samples (24 females and 24 males) from various origins, providing a useful tool for studying the adaptive evolution of male and female H. tibetana.
Wang et al.3 suggested that chromosome 2 of H. tibetana might be its sex chromosome. Our study showed that loci associated with sex are primarily concentrated on chromosome 2, which aligns with the hypothesis that this chromosome serves as the sex chromosome for H. tibetana. Previous research has indicated that the evolution of dioecy in Hippophae species (seabuckthorn) is relatively recent. For example, H. rhamnoides subsp. turkestanica may exhibit characteristics of subdioecy in its transition to complete dioecy, reflecting a high similarity between the male and female genomes of seabuckthorn, along with functional dynamic partitioning42. Our analysis also revealed multiple regions related to sex determination on chromosome 2 of H. tibetana, and a few sex-differentiating loci on other chromosomes, suggesting a complex evolutionary history of its sex chromosomes. These characteristics position seabuckthorn, particularly H. tibetana, as an ideal subject for studying plant sex differentiation and evolution. Researching these mechanisms in the extreme environments where H. tibetana thrives is particularly significant for understanding the adaptive evolution of male and female plants.
This study introduces a simpler and more efficient method for designing molecular markers by rapidly screening long fragment InDel variations in male individuals. Verification across the male and female samples of H. tibetana from various sources suggests that all three pairs of primers possess significant sex-differentiating capabilities, highlighting their specificity and potential for application. This method is theoretically applicable to other species with XY/ZW sex determination systems, especially those that already have high-quality reference genomes. The methodology presented in this study not only facilitates the development of sex-specific molecular markers but also enhances our understanding of sex evolution mechanisms. We aim to apply this method for developing universal primers applicable across the entire genus of seabuckthorn, further exploring the mechanisms of sex evolution in seabuckthorn. Such research will offer valuable insights into the mechanisms of sex differentiation and adaptive evolution of plants in extreme environments and provide scientific support for the large-scale breeding and precise management of economically useful plants such as seabuckthorn.
Conclusion
In this study, we performed whole-genome resequencing on 32 sexually differentiated individuals of H. tibetana. The results revealed that sex-related loci are predominantly concentrated on chromosome 2, providing robust evidence for its role as the sex chromosome in H. tibetana. Based on this insight, we developed a more efficient method to derive three pairs of accurate sex-linked molecular markers, effectively addressing the challenge of sex identification in this species. These markers were proven highly accurate and reliable for rapid sex identification in H. tibetana, and thus can serve as promising tools for this purpose. This study not only delivers an efficient approach for sex identification in H. tibetana, which can aid in optimizing commercial cultivation and minimizing resource waste, but also lays the groundwork for in-depth investigations into the sex determination mechanism and adaptive evolution of male and female plants in H. tibetana.
Data availability
The raw resequencing data and population genetic variation data generated in this study have been deposited in the China National Center for Bioinformation, with accession numbers CRA017060 (https://ngdc.cncb.ac.cn/gsa) and GVM000732 (http://bigd.big.ac.cn/gvm), respectively.
References
Wang, H. et al. Phylogeographic structure of Hippophae tibetana (Elaeagnaceae) highlights the highest microrefugia and the rapid uplift of the Qinghai-Tibetan Plateau. Mol. Ecol. 19, 2964–2979 (2010).
He, X., Si, J., Zhao, C., Wang, C. & Zhou, D. Potential distribution of Hippophae thibetana and its predicted responses to climate change. J. Desert Res. 43, 101 (2021).
Wang, R. et al. How to survive in the world’s third poplar: Insights from the genome of the highest altitude woody plant, Hippophae tibetana (Elaeagnaceae). Front. Plant Sci. 13, 1051587 (2022).
Ruan, C.-J., Rumpunen, K. & Nybom, H. Advances in improvement of quality and resistance in a multipurpose crop: Sea buckthorn. Crit. Rev. Biotechnol. 33, 126–144 (2013).
Ding, X. et al. Wild plants used by tibetans in Burang Town, characterized by alpine desert meadow, in Southwestern Tibet, China. Agronomy 12, 704 (2022).
Nybom, H., Ruan, C. & Rumpunen, K. The systematics, reproductive biology, biochemistry, and breeding of sea Buckthorn—A review. Genes 14, 2120 (2023).
Boesi, A. Traditional knowledge of wild food plants in a few Tibetan communities. J. Ethnobiol. Ethnomed. 10, 1–19 (2014).
Wei, J. et al. Phenolic compositions and antioxidant activities of Hippophae tibetana and H. rhamnoides ssp. sinensis berries produced in Qinghai-Tibet Plateau. Food Chem. 15, 100397 (2022).
Liu, Y. et al. Identification of the traditional Tibetan medicine “Shaji” and their different extracts through tri-step infrared spectroscopy. J. Mol. Struct. 1124, 180–187 (2016).
Jia, D. R., Liu, T. L., Wang, L. Y., Zhou, D. W. & Liu, J. Q. Evolutionary history of an alpine shrub Hippophae tibetana (Elaeagnaceae): Allopatric divergence and regional expansion. Biol. J. Linn. Soc. 102, 37–50 (2011).
Zhang, G. et al. Chromosome-level genome assembly of Hippophae tibetana provides insights into high-altitude adaptation and flavonoid biosynthesis. BMC Biol. 22, 82 (2024).
Qiong, L., Ngudrub, N. & Duan, S. Q. A preliminary study on flowering date, flower morphology and flower number of Hippophae tibetana. J. Tibet Univ. Nat. (Sci. Ed.) 24, 21–23 (2009).
Guo, D., Wang, R., Fang, J., Zhong, Y. & Qi, X. Development of sex-linked markers for gender identification of Actinidia arguta. Sci. Rep. 13, 12780 (2023).
She, H. et al. Identification of a male-specific region (MSR) in Spinacia oleracea. Hortic. Plant J. 7, 341–346 (2021).
Bui, T. L., Truong, N. T. & Do, T. K. The application of molecular marker in papaya sex determination: From the laboratory to the field. Sci. Hortic. 327, 112872 (2024).
Sharma, A., Zinta, G., Rana, S. & Shirko, P. Molecular identification of sex in Hippophae rhamnoides L. using isozyme and RAPD markers. For. Stud. China 12, 62–66 (2010).
Korekar, G. et al. Identification and validation of sex-linked SCAR markers in dioecious Hippophae rhamnoides L. (Elaeagnaceae). Biotechnol. Lett. 34, 973–978 (2012).
Chawla, A., Kant, A., Stobdan, T., Srivastava, R. B. & Chauhan, R. S. Cross-species application of sex linked markers in H. salicifolia and H. tibetana. Sci. Hortic. 170, 281–283 (2014).
Das, K. et al. ISSR markers for gender identification and genetic diagnosis of Hippophae rhamnoides ssp. turkestanica growing at high altitudes in Ladakh region (Jammu and Kashmir). Protoplasma 254, 1063–1077 (2017).
Zhou, W. et al. Molecular sex identification in dioecious Hippophae rhamnoides L. via RAPD and SCAR markers. Molecules 23, 1048 (2018).
Jain, A., Kumar, A. & Sharma, P. C. The Seabuckthorn Genome (ed Prakash C. S.) 187–212 (Springer, 2022).
Sun, K. et al. Molecular phylogenetics of Hippophae L. (Elaeagnaceae) based on the internal transcribed spacer (ITS) sequences of nrDNA. Plant Syst. Evol. 235, 121–134 (2002).
Yu, L. A., Ma, X., Deng, B., Yue, J. & Ming, R. Construction of high-density genetic maps defined sex determination region of the Y chromosome in spinach. Mol. Genet. Genomics 296, 41–53 (2021).
Varshney, R. K., Terauchi, R. & McCouch, S. R. Harvesting the promising fruits of genomics: Applying genome sequencing technologies to crop breeding. PLoS Biol. 12, e1001883 (2014).
Aboul-Maaty, N. A. F. & Oraby, H. A. S. Extraction of high-quality genomic DNA from different plant orders applying a modified CTAB-based method. Bull. Natl. Res. Cent. 43, 1–10 (2019).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Jung, Y. & Han, D. BWA-MEME: BWA-MEM emulated with a machine learning approach. Bioinformatics 38, 2404–2413 (2022).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): High-performance genomics data visualization and exploration. Briefings Bioinf. 14, 178–192 (2013).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: A cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS One 11, e0163962 (2016).
Untergasser, A. et al. Primer3—New capabilities and interfaces. Nucleic Acids Res. 40, e115–e115 (2012).
Chen, Y., Ye, W., Zhang, Y. & Xu, Y. High speed BLASTN: An accelerated MegaBLAST search tool. Nucleic Acids Res. 43, 7762–7768 (2015).
Puterova, J. et al. Satellite DNA and transposable elements in seabuckthorn (Hippophae rhamnoides), a dioecious plant with small Y and large X chromosomes. Genome Biol. Evol. 9, 197–212 (2017).
Guo, D. et al. Validation of kiwifruit sex molecular markers in Actinidia arguta. J. Fruit Sci. 36, 549–556 (2019).
Zhang, P. X. et al. Validation of a male-linked gene locus (OGI) for sex identification in persimmon (Diospyros kaki Thunb.) and its application in F1 progeny. Plant Breed. 135, 721–727 (2016).
Badenes, M. L., Fernández i Martí, A., Ríos, G. & Rubio-Cabetas, M. J. Application of genomic technologies to the breeding of trees. Front. Genet. 7, 222377 (2016).
da Fonseca, R. R. et al. Next-generation biology: Sequencing and data analysis approaches for non-model organisms. Mar. Genomics 30, 3–13 (2016).
Luo, H. et al. Identification of male sex-specific markers using genome re-sequencing in the Chinese longsnout catfish Leiocassis longirostris. Aquaculture 558, 738392 (2022).
Mangla, Y. et al. Occurrence of subdioecy and scarcity of gender-specific markers reveal an ongoing transition to dioecy in Himalayan seabuckthorn (Hippophae rhamnoides ssp. turkestanica). Heredity 122, 120–132 (2019).
Acknowledgements
The authors wish to extend their sincere gratitude to the National Natural Science Foundation of China for funding this research (Grant No. 31760127). We are also thankful for the financial support from the First-class Discipline Construction Project of Ecology (Zangcaijiaozhi [2023]01), the Tibet Autonomous Region Natural Science Foundation Project (ZRKX2024000019), the Key R&D Project of Science and Technology Program of Tibet Autonomous Region (No. XZ202301ZY0006G), and the Graduate High-level Talent Training Program of Tibet University (2022-GSP-B003). Special acknowledgment is extended to Zhiping Song, Yuguo Wang, and Wenju Zhang for their support by providing the data analysis platform, which was essential for the successful conduct of our experiments.
Funding
This work was supported by the National Natural Science Foundation of China (No. 31760127), the First-class Discipline Construction Project of Ecology (Zangcaijiaozhi [2023] 01), the Tibet Autonomous Region Natural Science Foundation Project (ZRKX2024000019), the Key R&D Project of Science and Technology Program of Tibet Autonomous Region (NO. XZ202301ZY0006G), and the Graduate High-level Talent Training Program of Tibet University (2022-GSP-B003).
Author information
Authors and Affiliations
Contributions
L.Q. and W.Z. contributed to the conception and design of the study. Z.Z., R.W., Y.C., and J.W. conducted the experiments. Z.Z. and R.W. were responsible for data analysis. Z.Z. prepared the initial draft of the manuscript and managed the imagery, incorporating feedback from W.Z., L.Q., R.W., Y.W., and Z.S. All authors participated in revising the draft and approved the final manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeng, Z., Wang, R., Wang, J. et al. Development and validation of sex-linked molecular markers for rapid and accurate identification of male and female Hippophae tibetana plants. Sci Rep 14, 19243 (2024). https://doi.org/10.1038/s41598-024-69918-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69918-y
- Springer Nature Limited