Abstract
Acute myeloid leukemia (AML) exhibits pronounced heterogeneity and chemotherapy resistance. Aberrant programmed cell death (PCD) implicated in AML pathogenesis suggests PCD-related signatures could serve as biomarkers to predict clinical outcomes and drug response. We utilized 13 PCD pathways, including apoptosis, pyroptosis, ferroptosis, autophagy, necroptosis, cuproptosis, parthanatos, entotic cell death, netotic cell death, lysosome-dependent cell death, alkaliptosis, oxeiptosis, and disulfidptosis to develop predictive models based on 73 machine learning combinations from 10 algorithms. Bulk RNA-sequencing, single-cell RNA-sequencing transcriptomic data, and matched clinicopathological information were obtained from the TCGA-AML, Tyner, and GSE37642-GPL96 cohorts. These datasets were leveraged to construct and validate the models. Additionally, in vitro experiments were conducted to substantiate the bioinformatics findings. The machine learning approach established a 6-gene pan-programmed cell death-related genes index (PPCDI) signature. Validation in two external cohorts showed high PPCDI associated with worse prognosis in AML patients. Incorporating PPCDI with clinical variables, we constructed several robust prognostic nomograms that accurately predicted prognosis of AML patients. Multi-omics analysis integrating bulk and single-cell transcriptomics revealed correlations between PPCDI and immunological features, delineating the immune microenvironment landscape in AML. Patients with high PPCDI exhibited resistance to conventional chemotherapy like doxorubicin but retained sensitivity to dasatinib and methotrexate (FDA-approved drugs for other leukemias), suggesting the potential of PPCDI to guide personalized therapy selection in AML. In summary, we developed a novel PPCDI model through comprehensive analysis of diverse programmed cell death pathways. This PPCDI signature demonstrates great potential in predicting clinical prognosis and drug sensitivity phenotypes in AML patients.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) arises from the abnormal clonal expansion of myeloid hematopoietic progenitor cells and represents a highly heterogeneous group of hematological malignancies1. Despite its low incidence, AML remains the most common type of acute leukemia in adults and the second most prevalent leukemia overall2. While novel treatment options continue to emerge, clinical outcomes of AML patients over 5 years remain disappointing, with many patients experiencing relapse and developing resistance to therapy, leading to treatment failure and mortality3. Prognosis in AML depends on several factors including disease history4, age5, and cytogenetic and molecular abnormalities such as mutations in NPM1, CEBPA, FLT3, and KIT6. Although incremental advancements have been made in the diagnosis and treatment of AML, favorable responses are often limited to a minority of patients, and the emergence of drug resistance frequently results in therapeutic failure, especially for elderly AML patients, with the majority succumbing to the disease within one year of diagnosis1,3,5,7. Therefore, the development of novel prognostic biomarkers to enable personalized risk stratification and prediction of therapeutic response is critically needed.
Programmed cell death (PCD), which enables the proper formation of organs during embryonic development, plays a fundamental role not only in animal development but also in the maintenance of tissue homeostasis8,9,10,11,12. Dysfunction of PCD is associated with multiple pathological conditions and the development of various diseases, including immune disorders, neurodegeneration, tissue damage, and cancers, such as breast cancer, liver cancer, and AML9,10,13,14,15. Currently, PCD consists of 13 major cell death patterns, including apoptosis, pyroptosis, ferroptosis, autophagy, necroptosis, cuproptosis, parthanatos, entotic cell death, netotic cell death, lysosome-dependent cell death (LDCD), alkaliptosis, oxeiptosis, and disulfidptosis10,16,17. Apoptosis is the most well-studied PCD pathway. Defective apoptosis is a hallmark of AML, contributing to treatment resistance18. Restoring apoptosis sensitivity in leukemia cells represents a promising therapeutic strategy19. Autophagy is a tightly regulated homeostatic process that controls cell survival and cell death. It involves the encapsulation of cytoplasmic contents within autophagosomes, which then fuse with lysosomes for degradation and recycling of the sequestered material20. Defects in autophagy can promote leukemogenesis in AML by enabling the survival of pre-leukemic cells8. Additionally, autophagy has been implicated in resistance to cytarabine11 and targeted therapies like FLT3 in AML21. Disulfidptosis is a recently discovered iron-dependent form of programmed cell death induced by disulfide stress resulting from the accumulation of incorrect disulfide bonds within the endoplasmic reticulum16. While the role of disulfidptosis in AML pathogenesis remains to be fully elucidated, it may be involved in the perturbation of redox homeostasis in AML cells. Ferroptosis is an iron- and ROS-dependent form of PCD characterized by lipid peroxidation and loss of membrane integrity17. Aberrant expression of genes regulating ferroptosis, including downregulation of GPX4 and altered expression of NRF2, has been observed in AML, which may disrupt redox homeostasis12,22.
Other forms of PCD such as necroptosis, pyroptosis, lysosome-dependent cell death, and entotic cell death may also contribute to AML pathogenesis23. However, the association between the multiple PCD pathways as a whole and AML biology requires further study. Elucidating the roles of PCD-related genes could provide insight into AML pathogenesis and reveal potential therapeutic targets. In this study, we identified six pan-programmed cell death-related genes (PPCDS) demonstrating differential expression in AML. We subsequently developed a PPCDS index (PPCDI) to investigate potential associations between PPCDS and AML prognosis. The genetic and mutational landscape of these PPCDS in AML was characterized. Prognostic models incorporating PPCDS and PPCDI data were then constructed to predict overall survival (OS) in both training and validation AML cohorts. Furthermore, we examined interactions between PPCDS/PPCDI levels and immune features in AML. Additionally, the immunotherapy and targeted therapy response related to PPCDS/PPCDI status in AML was evaluated and verified.
Material and methods
Data acquisition and preprocessing
RNA sequencing (RNA-seq) expression data (log2 transformed) were obtained for 129 AML patients from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.com). Additionally, the expression profiling data for three external AML cohorts were acquired from the Tyner’s cohort24, GSE37642-GPL9625, and Target (https://ocg.cancer.gov/programs/target) datasets, respectively. Corresponding clinicopathological characteristics for AML patients from these three cohorts were also collected. Furthermore, single-cell RNA sequencing (scRNA-seq) data from 20 AML patient samples were retrieved from the GSE185993 dataset26.
PPCD gene list
A total of 3,246 PCD-related genes were compiled from the KEGG database, GSEA gene sets, and previous publications by integrating 13 major forms of PCD. These encompassed genes involved in apoptosis (n = 580), pyroptosis (n = 52), ferroptosis (n = 91), autophagy (n = 370), necroptosis (n = 104), cuproptosis (n = 14), parthanatos (n = 9), entotic cell death (n = 15), netotic cell death (n = 8), lysosome-dependent cell death (n = 220), alkaliptosis (n = 7), oxeiptosis (n = 5), and disulfidptosis (n = 1771), as categorized based on prior reports10,17 (Table S1).
Identification of the PPCDS in AML
To identify genes potentially affecting prognosis in AML, differential expression analysis was conducted comparing deceased and surviving AML patients across the three datasets using the limma package. A P-value threshold of < 0.05 was utilized to determine statistical significance. Intersecting the differentially expressed genes (DEGs) identified in each cohort yielded a final DEG list. This DEG list was then cross-referenced with genes associated with the various PCD forms to derive the PCD-related differentially expressed genes (PPCDS) signature.
Construction of the PPCDS signature
To further identify the PPCDS associated with prognosis in AML, 10 machine learning algorithms were implemented, including Supervised Principal Components (SuperPC), CoxBoost, stepwise Cox regression, Least Absolute Shrinkage and Selection Operator (LASSO), Generalized Boosted Regression Modeling (GBM), Survival Support Vector Machine (Survival-SVM), Partial Least Squares regression for Cox models (plsRcox), Ridge regression, Elastic Net (Enet), and Random Survival Forest (RSF). A total of 73 algorithm combinations were evaluated. To develop a robust PPCDS signature with optimal stability, the 73 models were trained in a leave-one-out cross-validation (LOOCV) framework27. Model performance was assessed using the concordance index (c-index), from which the final optimal algorithm combination was selected.
Consensus clustering and principal component analysis
Consensus clustering (CC) analysis for the TCGA and Tyner’s cohort was performed utilizing the ConsensusClusterPlus R package28. The algorithm was configured for agglomerative hierarchical clustering by partitioning around medoids, with a Pearson correlation distance metric. Resampling of 80% of the sample population was conducted for a total of 1000 iterations. Principal component analysis (PCA) was implemented using the R stats package to normalize the expression data. This involved z-score calculation followed by dimensionality reduction through principal component extraction using the prcomp function. This generated an optimized low-dimensional matrix for subsequent analyses.
Variation analysis of the PPCDS
Based on PPCDI level, patients in the TCGA-AML cohort were divided into PPCDI-high and low subgroups, the mutation features of patients in various groups were explored using the maftools package. Additionally, we examined the alteration landscape of each PPCDS in AML based on 165 samples from the TCGA-AML dataset using cBioPortal database (with full expression, mutation, and CAN data)29.
Functional enrichment analysis
Gene set enrichment analysis was conducted on the DEGs using the clusterProfiler and ggplot2 R packages. Enrichment for Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways and Gene Ontology (GO) biological process terms was evaluated. Additionally, the Gene Set Variation Analysis (GSVA) was utilized to assess differences in HALLMARK gene set functions between the high and low PPCDI groups across the three cohorts, referencing the h.all.v2022.1.Hs.symbols.gm database30.
Construction of the nomogram model for PPCDS
Prognostic nomogram models incorporating PPCDI and relevant clinical variables were constructed to predict the clinical outcomes of AML patients across the three cohorts31. Model clinical utility was evaluated through calibration analysis, decision curve analysis (DCA), and time-dependent receiver operating characteristic (ROC) curve analysis implemented in the rms, survival, and timeROC R packages, respectively32,33.
Distribution and expression of PPCDS at the single cell level
Single-cell RNA sequencing data from 20 AML samples were obtained from the GEO database (GSE185993) and analyzed using the Seurat and SingleR packages26. Quality control and preprocessing were performed, involving data normalization, variable feature selection, scaling, dimensionality reduction with PCA, batch effect correction via Harmony, clustering with shared nearest neighbor modularity optimization, UMAP dimensionality reduction, and differential expression analysis to identify cluster markers. This yielded a total of 42,475 high quality cells for downstream analyses. SingleR analysis coupled with known literature-based markers were utilized to assign cell type labels26. Marker gene functional enrichment for each cluster was evaluated with the ClusterGVis34, org.Hs.eg.db and enrichCluster R packages. Copy number variation (CNV) scores were calculated for each proliferating cells using inferCNV algorithm, with T cells set as the reference35. Pseudotime trajectory analysis was conducted using the CytoTRACE36 and Monocle237 packages. Furthermore, the PPCDS scores were calculated for each cell using the AddModuleScore function.
Tumor microenvironment analysis
The expression profiles of PPCDS along with abundances of infiltrating immune cell populations in the tumor microenvironment were analyzed for TCGA-AML and Tyner AML cohorts. Estimation of immune cell type scores was performed by applying a series of computational methods including TIMER, QUANTISEQ, EPIC, MCPCOUNTER, and ESTIMATE algorithms through the IOBR package38.
Immunotherapy and drug sensitivity prediction
To investigate potential associations between the PPCDS/PPCDI signature and immune checkpoint molecule expression in AML, six immune checkpoint-related transcripts were selected for analysis, including CD274, CTLA4, LAG3, PDCD1, TIGIT, and SIGLEC15. The expression profiles of these genes were extracted from both the TCGA-AML and Tyner AML cohorts. The Tumor Immune Dysfunction and Exclusion (TIDE) algorithm39 was utilized to predict differential immunotherapy response between high and low PPCDI subgroups. Additionally, the “oncoPredict” R package40 was leveraged to evaluate differences in drug sensitivity.
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
This investigation was sanctioned by the Ethics Committee of the First Affiliated Hospital of Anhui Medical University. The study was conducted in accordance with the protocol, the principles of the “Declaration of Helsinki” in its current version and the requirements of the Chinese law. The cohort comprised twelve patients with acute myeloid leukemia (AML) newly diagnosed at the institution’s Hematology Department, alongside twelve healthy adults ascertained from the hospital’s Health Examination Center, all of whom provided written informed consent. Peripheral blood specimens were the biological material of choice, with AML diagnoses aligning with the guidelines delineated for the management of adult acute myeloid leukemia in China, acute promyelocytic leukemia excepted.
The transcriptional profiles of genes within the PPCDI cluster were scrutinized by leveraging RT-qPCR analysis to compare the expression between AML and normative healthy samples. The extraction of total RNA was facilitated by the Fastagen RNA extraction kit, subsequent to which cDNA synthesis was executed utilizing the M-MLV reverse transcriptase kit (Takara, Japan), adhering strictly to the protocols prescribed by the manufacturer. Quantitative assessments were conducted employing the ABI PRISM 7500 Sequence Detection System (Applied Biosystems) in conjunction with a dedicated RT-qPCR reagent kit (Promega). Quantitative data were procured via the standard curve method and the expression values were normalized against those of the β-actin (human) gene, serving as an internal reference.
Statistical analysis
Statistical analyses were conducted using R software (v4.2.1). Between-group comparisons of two groups were performed using Student’s t-test or Wilcoxon rank sum test as appropriate, while Kruskal-Wallis one-way analysis of variance was applied for comparisons involving more than two groups. Kaplan-Meier survival curves were generated and differences between curves were assessed. Associations between continuous variables were evaluated using Spearman’s rank correlation analysis. A p-value threshold of 0.05 was set for statistical significance in all analyses.
Ethics statement
This study was reviewed and approved by the First Affiliated Hospital Ethics Committee of Anhui Medical University. Ethics approval number 2023342. Before participating in the study, all patients and healthy volunteers provided written informed consent.
Results
Identification of pan-programmed cell death-related genes and model construction
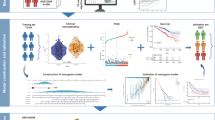
This study utilized 129 patients with complete clinical data from TCGA database as the training cohort. The validation cohorts consisted of 398 patients from the Tyner’s dataset and 417 patients from the GSE37642-GPL96 dataset. Clinical parameters for patients across the three cohorts are presented in Table S2, and the study workflow is outlined in the Fig. 1. To identify robust prognosis-associated DEGs, the intersection of DEGs from the three cohorts was derived, yielding 140 common DEGs (Fig. 2A,B, Tables S3–S5). Moreover, we performed a hypergeometric test to evaluate the overlap of the three datasets’ DEGs. The test yielded a p-value of 7.00e-49, demonstrating that the overlap is highly significant and unlikely to be due to statistical chance (Fig. 2B). GO and KEGG enrichment analysis revealed these 140 DEGs were highly enriched for pathways related to cell adhesion and lipid metabolism (Fig. 2C, Table S6). By overlaying the DEGs with genes associated with specific PCD forms, 30 pan-PCD related DEGs were identified, including 15 disulfidptosis, 1 pyroptosis, 2 ferroptosis, 3 lysosome-dependent cell death, 2 netotic cell death, 9 apoptosis, and 5 autophagy genes (Fig. S1, Table S7).
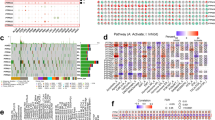
Identification of PPCDS and construction of the PPCDI signature. (A, B) Venn analysis of differentially expressed genes (DEGs) from three AML cohorts identified 140 intersecting DEGs. (C) Gene Ontology and KEGG enrichment analysis of the 140 DEGs. (D) Evaluation of C-indices for 73 prediction models generated via leave-one-out cross-validation in the TCGA-AML, Tyner, and GSE37642-GPL96 cohorts. (E) Relative importance of PPCDS for AML prognosis based on recursive feature elimination. (F) Cross-validation of the PPCDI signature constructed via LASSO regression. (G) Genomic location, expression, and correlation of PPCDS in the TCGA cohort. PPCDS: pan-programmed cell death-related genes; PPCDI: PPCD index.
To identify optimal algorithms for constructing PPCDI, 73 machine learning models utilizing 10 different algorithms were evaluated. As shown in Fig. 2D, StepCox, RSF + StepCox, and LASSO + StepCox exhibited the highest average c-index (Table S8). To mitigate variable selection bias, the RSF and LASSO methods were intersected to derive a robust PPCDS signature, followed by multivariate Cox regression for final model building. The PPCDI score for each patient was calculated by summing the products of gene expression and corresponding coefficients. Patients were stratified into PPCDI-high and low subgroups based on the PPCDI median value. The RSF analysis identified 15 top-ranked genes by importance (Fig. 2E). By intersecting with the LASSO-selected genes, 6 core PPCDS genes were defined: ATP6V0E2, DDIT4, ETS2, DOCK1, CLEC11A, and HGF (Fig. 2F). Chromosomal locations and co-expression patterns were examined for each PPCDS (Fig. 2G). Most PPCDS demonstrated positive correlation in the TCGA-AML cohort, except for CLEC11A and HGF (Fig. 2G). Additionally, univariate Cox regression and Kaplan-Meier analyses were used to evaluate the associations between individual PPCDS and AML prognosis across cohorts. ATP6V0E2, DDIT4, ETS2, and DOCK1 conferred risk, while CLEC11A and HGF had protective effects (Fig. S2). Consistent results were observed in the Tyner’s and GSE37642-GPL96 cohorts (Fig. S3A,B). To ensure the accuracy of the above findings, we incorporated another outside cohort study involving 279 AML patients from Target datasets, analyzing the association between PPCDS and the clinical outcomes of AML patients. The findings indicated that the PPCDS displayed consistent behavior across all cohorts, further validating their potential as biomarkers (Fig. S3C).
Construction of PPCDI and its association with clinical characteristics of AML
Multivariate Cox regression analysis was conducted in the TCGA-AML cohort to determine the coefficients for each PPCDS gene in the PPCDI model: PPCDI = 0.27673 × ATP6V0E2exp + 0.099852 × DDIT4exp + 0.13156 × ETS2exp + 0.090291 × DOCK1exp-0.094115 × CLEC11Aexp-0.14942 × HGFexp. Of notice, elevated PPCDI significantly correlated with worse clinical outcomes and older patient age across all three AML cohorts (Fig. 3A–C). Using the PPCDS expression data, AML patients in the TCGA-AML and Tyner cohorts were clustered into two groups (C1 and C2) via consensus clustering (Fig. 3D,E). Kaplan-Meier analysis revealed patients in cluster C2 had improved OS compared to C1 (Fig. 3F,G, P < 0.001). Furthermore, alluvial diagram analysis demonstrated the majority of patients in cluster C1 had higher PPCDI scores and older age (Fig. 3H,I). The expression patterns and its association with the clinical features of AML patients in TCGA-AML cohort was shown in Fig. 3J.
Association between PPCDI and clinical features of AML. (A-C) Box plots represent the differences in PPCDI distribution across various groups within the three AML cohorts. A p-value of less than 0.05 indicates that the differences observed are statistically significant. (D, E) Unsupervised hierarchical clustering of PPCDS in TCGA-AML and Tyner cohorts, defining two molecular clusters at k = 2. (F, G) Kaplan-Meier analysis of prognosis among molecular clusters. (H, I) Alluvial diagrams depicting interrelationships between PPCDI groups, age, molecular clusters, and survival status. (J) Heatmap of PPCDS and clinical features.
Alteration landscape of PPCDS in AML
Somatic mutation analysis was performed comparing AML patients stratified into high and low PPCDI groups. Common mutated genes identified included DNMT3A, RUNX1, and NPM1, with higher mutation frequencies observed in the PPCDI-high subgroup (Fig. S4A). To further investigate the genomic alteration landscape of individual PPCDS in AML, the cBioPortal database was analyzed. PPCDS exhibited low alteration frequencies overall, with ETS2 harboring the highest mutation rate at 10% (Fig. S4B).
High PPCDI correlates with worse AML survival and is enriched in oncogenic pathways
Patients in the TCGA-AML, Tyner, and GSE37642-GPL96 cohorts were stratified into high and low PPCDI groups based on calculated PPCDI scores. Elevated PPCDI significantly associated with poor clinical outcomes of AML patients (Fig. 4A,B). Principal component analysis clearly separated the AML patients into PPCDI-high and low subgroups in each cohort (Fig. 4C). GSVA was subsequently performed to evaluate differences in HALLMARK pathway activity between PPCDI-defined subgroups in the Tyner cohort, revealing enriched pathways (Fig. 4D,E). Notably, cross-cohort analysis identified consistent enrichment of oncogenic pathways, including KRAS_SIGNALING_UP, IL2_STAT5_SIGNALING, P53_PATHWAY, and TGF_BETA_SIGNALING pathways (Fig. 4E, Table S9).
Performance of the PPCDI prognostic model in training and validation cohorts. (A) PPCDI distribution by survival status and time in TCGA-AML, Tyner, and GSE37642-GPL96 cohorts. (B) Kaplan-Meier analyses of overall survival in PPCDI-high vs PPCDI-low groups across cohorts. (C) Principal component analysis of PPCDI in the three cohorts. (D) GSEA of HALLMARK pathways between PPCDI groups in Tyner’s cohort. (E) Venn diagram of enriched HALLMARK pathways across cohorts.
Validation of the prognostic predictive ability of PPCDI in AML cohorts
Univariate and multivariate Cox regression analyses were performed to determine if PPCDI could independently predict AML prognosis. As shown in Fig. 5A, PPCDI emerged as a significant risk factor in the univariate model (HR = 2.718, 95% CI 1.935–3.819, P < 0.001). The association remained significant after adjusting for potential confounders (HR = 2.335, 95% CI 1.537–3.547, P < 0.001; Fig. 5A). A prognostic nomogram was then developed for 1-, 3-, and 5-year OS prediction in the TCGA cohort using stepwise multivariable Cox regression. The model incorporated PPCDI along with age and FAB stage as risk factors (Fig. 5B). Calibration curves demonstrated excellent concordance between predicted and observed outcomes (Fig. 5C). DCA revealed the nomogram and PPCDI outperformed other examined predictors (Fig. 5D). Significant survival differences were observed between high and low-risk groups stratified by nomogram score (Fig. 5E). The predictive utility of PPCDI was validated in the other two AML cohorts, with nomograms and calibration curves displayed in Fig. S5A–D. Evaluation of time-dependent area under the curve values demonstrated superior accuracy of the nomogram in predicting the clinical outcomes of AML patients in all AML cohorts (Fig. 5F–H). These results highlight the potential clinical utility of the PPCDI signature and nomogram for prognostic risk stratification, facilitating personalized therapy, and improving outcomes in AML patients.
Development and evaluation of the prognostic nomogram. (A) Univariate and multivariate Cox regression of PPCDI and clinical variables in TCGA-AML cohort. (B) Prognostic nomogram predicting AML survival. (C) Calibration plots of nomogram-predicted 1-, 3-, and 5-year overall survival probabilities in TCGA-AML cohort. (D) Decision curve analysis for 3-year survival prediction by the nomogram in TCGA-AML. (E) Kaplan-Meier analysis of nomogram-defined risk groups in TCGA-AML. (F–H) Time-dependent ROC analysis of the nomogram in TCGA-AML, Tyner, and GSE37642-GPL96 cohorts.
Single-cell analysis suggested PPCDS score correlates with the development of AML
To examine PPCDS expression and distribution at single-cell resolution, scRNA-seq data from AML patient samples (GSE185993) were analyzed. After quality control, 42,475 cells were retained for downstream analysis (Fig. S6A). Clustering identified 28 subpopulations, which were annotated into 7 cell types (Fig. 6A, Fig. S6B). Marker gene expression profiles were generated for each cluster (Fig. 6B, Fig. S6C), with enrichment analysis of the top 5 biological processes. The expression and distribution patterns of PPCDS were shown in Fig. S6D. CNV scores were calculated for proliferative cells using the inferCNV algorithm, with T-cells as reference (Fig. 6C,D). This delineated CNV-low (clusters 11, 13) and CNV-high (clusters 1, 8, 15, 16, 19, 26) subsets (Fig. 6D). Pseudotime trajectory analysis via CytoTRACE and Monocle2 was then performed to predict differentiation trajectories of proliferative subpopulations. A clear lineage progression was observed from CNV-low to CNV-high states (Fig. 6E–G). PPCDS scores were calculated for each cell using AddModuleScore (Fig. S6E). Notably, PPCDS score significantly correlated with malignant status of proliferative cells, linking PPCDS expression to AML pathogenesis (Fig. 6H).
Characterization of AML cell states and hierarchy. (A) Louvain clustering identifying 28 clusters in 42475 AML cells. (B) Heatmap of top differentially expressed genes across 7 cell types; top 5 enriched biological processes listed. (C) Hierarchical heatmap of large-scale copy number variations (CNVs) in proliferating cells. (D) CNV scores across proliferating cell clusters. (E, F) Developmental trajectory prediction by CytoTRACE. Boxplots show median and interquartile range. (G) Monocle 2 trajectory analysis combining two proliferating cell subclusters. (H) Comparison of PPCDI scores between CNV-low and high subclusters.
PPCDI can predict immunotherapy and targeted therapy response in AML
Additional analyses were conducted to identify differences in other relevant features between PPCDI-high and -low groups. First, associations were examined between PPCDS/PPCDI levels and the infiltrating of various immune cells calculated using various algorithms. Interestingly, ImmuneScore, ESTIMATEScore, and infiltration levels of CD4 + T cells, CD8 + T cells, NK cells, neutrophils, macrophages, and myeloid dendritic cells demonstrated significant positive correlations with PPCDI scores and most PPCDS genes including ATP6V0E2, DDIT4, ETS2, and DOCK1. Conversely, these immune features negatively correlated with CLEC11A and HGF expression in both TCGA-AML and Tyner cohorts (Fig. 7A, Tables S10, 11). Next, expression of six immune checkpoint molecules was analyzed, revealing predominantly positive associations between checkpoint genes and PPCDS/PPCDI levels (Fig. 7B,C, Tables S10, 11). Strikingly, PPCDS and PPCDI also positively correlated with TIDE and dysfunction scores in AML, implying potential superiority of immunotherapy responses in PPCDI-low patients (Fig. 7B).
Association between PPCDI/PPCDS and immune profiles as well as drug sensitivity in AML. (A) Heatmap of PPCDS/PPCDI correlations with immune cell infiltration estimated by various algorithms. (B) PPCDS/PPCDI associations with immune checkpoint expression and TIDE scores. (C) Expression of six immune checkpoint molecules in PPCDI high vs low groups. (D) Bubble chart relating drugs, PPCDI, and PPCDS. Size and color indicate strength of association. (E) Correlations between PPCDS and classical AML therapeutic targets. Red and blue lines indicate positive and negative correlations, respectively; solid and dashed lines indicate significant (P < 0.05) and non-significant (P > 0.05) correlations. (F) Boxplots of drug IC50 values in PPCDI high vs low groups. (G) Correlations between drug IC50 values and PPCDI scores of individual AML patients.
To elucidate the relationship between the model and drug sensitivity, we calculated the half maximal inhibitory concentration (IC50) values for each drug in TCGA-AML samples, aiming to identify drugs with significant differential activity. The landscape of correlations and significance between drug sensitivities and PPCDI is presented in Fig. 7D and Table S12. We observed that the IC50 of doxorubicin was higher, while the IC50 values of dasatinib and methotrexate were lower in the PPCDI-high group (Fig. 7D–G). These findings imply that AML patients with elevated PPCDI levels tend to be resistant to standard chemotherapy regimens, yet may remain sensitive to two other FDA-approved drugs originally developed for different leukemias. Therefore, dasatinib and methotrexate represent promising options for salvage therapy in chemotherapy-resistant AML. The associations between model genes and established AML therapeutic targets are presented in Fig. 7E.
In vitro experiments validation of the clinical significance of PPCDS in AML
RT-qPCR analysis was employed to ascertain the disparities in the expression of PPCDS mRNA between cohorts of normal healthy individuals and those with newly diagnosed AML. The sequences of the PCR primers targeting PPCDS are delineated in Table S13. As illustrated in Fig. 8A–D, a marked upregulation in the expression levels of ATP6V0E2, ETS2, DOCK1, and DDIT4 mRNA was observed in the AML cellular samples relative to the normal counterparts, corroborating the insights gleaned from bioinformatics analyses. Although the expression levels of CLEC11A and HGF did not show significant changes in the tumor cohort (ns, P > 0.05), as depicted in Fig. 8E,F, this observation is inconsistent with predictions from bioinformatics analyses, indicating that further elucidation of their expression at the protein level is warranted.
In-vitro experiments validation of the significance of PPCDS in AML. (A-F) RT-qPCR analysis of PPCDS mRNA expression in normal healthy individuals and those with newly diagnosed AML. Data in (A-I) are presented as mean ± SD (n = 6 per group) and p-values by two-tailed unpaired Student’s t-test are indicated. **P < 0.01, ns, P > 0.05.
Discussion
To the best of our knowledge, this is the first study to comprehensively investigate 13 diverse programmed cell death pathways, identify six PPCDS in AML, and construct a PPCDI signature using the TCGA-AML cohort. Validation of the PPCDI signature in two external AML cohorts (Tyner’s cohort, GSE37642-GPL96) confirms its performance. To evaluate the clinical utility of PPCDS in AML, we developed a nomogram incorporating PPCDI and relevant clinical factors, and verified its predictive applicability. Importantly, our findings reveal significant associations between PPCDI and the tumor microenvironment and drug sensitivity in AML. These results underline the potential value of PPCDI for stratifying AML patients and guiding personalized therapy.
PCD comprises biological and pathological processes that regulate tissue homeostasis, stress response, and cell proliferation, which is intricately associated with anti-tumor immunity in various cancers9,10,13,17,20. While prior studies have described roles for discrete PCD pathways in some cancers10,14, the genetic landscape, multimolecular mechanisms, and immune functions of pan-PCD genes in AML remain poorly characterized. Herein, we constructed a PPCDI signature based on six key PPCDS-ATP6V0E2, DDIT4, ETS2, DOCK1, CLEC11A, and HGF. We demonstrate this PPCDI signature can predict prognosis and therapy response in individual AML patients.
In a range of cancer types, the genes ATP6V0E2, DDIT4, ETS2, DOCK1, CLEC11A, and HGF have been subjected to extensive research. However, their functional roles and the mechanisms by which they are regulated within the context of AML remain insufficiently characterized. To corroborate the insights gleaned from our bioinformatics analysis, RT-qPCR was employed to assess the expression levels of PPCDS in samples from both healthy individuals and those diagnosed with AML. The findings largely corroborated the anticipated patterns of expression. Nonetheless, the levels of HGF and CLEC11A did not align with the bioinformatic predictions, signaling the necessity for further validation at the protein level.
ATP6V0E2 encodes a component of vacuolar ATPase (V-ATPase) that acidifies intracellular compartments. As mentioned previously, anlotinib can upregulate ATP6V0E2 expression and enhance lysosomal activity in colon cancer cells, resulting in attenuated apoptosis41. Of interest, AML cells also depend on lysosomal function to sustain redox homeostasis and survival. As such, increased ATP6V0E2 expression and lysosomal activation in AML could similarly protect leukemia cells from drug-induced apoptosis. DDIT4 is a stress-responsive protein that participates in mTOR pathway regulation, which is induced by various cellular stresses including DNA damage, oxidative stress, and nutrient deprivation42. Multiple studies have found high expression of DDIT4 associated with poor prognosis in AML patients43. DDIT4 may promote AML cell survival under chemotherapy by modulating autophagy and mTOR activity. ETS2 encodes an ETS family transcription factor that regulates gene expression and participates in the RAS/MAPK cascade44. Stromal ETS2 in the mouse pancreas helps recruit immunosuppressive cells like myeloid-derived suppressor cells and T regulatory cells, as well as chemokines45. Intriguingly, ETS2 may play a role in thymocyte proliferation and survival through a Myc-dependent pathway46. DOCK1 is a guanine nucleotide exchange factor that modulates Rho family GTPase signaling, which is critical for multiple cellular process, such as cell adhesion and migration, and actin cytoskeleton47. Specifically, the Dock1/Rac pathway is specifically targetable in the Npm1cA/+; Smc3Δ/+ AML with concurrent NPM1cA and cohesin mutations. Targeting this pathway could benefit this subtype, demonstrating the potential of genetics-guided precision oncology48. Conversely, upregulation of CLEC11A and HGF correlates with improved AML prognosis through various mechanisms49,50, consistent with our findings.
In this study, we did not observe significant associations between mutations in FLT3, IDH1, RAS, and NPMc and overall survival in TCGA-AML patients. The relatively low frequency of alterations in these individual genes within the cohort may partially account for the lack of detectable survival correlations. Moreover, AML is recognized as a high-risk disease driven by co-occurring combinations of mutations across many genes. Individual mutations alone may not exert sufficient prognostic influence48. In this study, we found upregulation of ATP6V0E2, DDIT4, ETS2, DOCK1 and downregulation of CLEC11A and HGF associated with poorer AML survival. A PPCDI signature calculated via multivariate Cox regression demonstrated robust prognostic performance, suggesting the PPCDI could be a potential prognostic biomarker in AML patients.
Increasing evidence indicates that tumor-microenvironment interactions influence T cell infiltration and immune cell function, relevant to immunotherapy27,39. We observed strong positive PPCDS/PPCDI correlations with immune infiltrates, indicating genetic-immunological crosstalk. High PPCDI patients exhibited elevated immune checkpoint molecules, implying heightened evasion and immunosuppression. Consistently, high PPCDI associated with increased TIDE and dysfunction scores, suggesting immunotherapy resistance. Moreover, elevated PPCDI associated with increased doxorubicin IC50 values, which may account for the poor prognosis resulting from chemotherapy resistance. In contrast, the IC50 values of dasatinib and methotrexate was lower in the PPCDI-high group, suggesting these agents as potential therapeutic alternatives for refractory AML.
Despite the potential of our model, some limitations warrant acknowledgement. First, the retrospective patient recruitment may have introduced partial bias. Second, the clinical utility of this model lacks validation in phase 3 randomized controlled trials, which are essential to inform treatment decisions. Third, due to the lack of IC50 data for other commonly used AML chemotherapeutic agents such as daunorubicin, cytarabine, mitoxantrone, and etoposide in the GDSC database, we were unable to investigate the association between PPCDI and the sensitivity of AML cells to these conventional chemotherapeutic drugs. Consequently, further investigations are warranted to assess the responsiveness of PPCDI to these standard AML chemotherapeutic agents. Although our study provides some preliminary evidence through in vitro PCR experiments, these results present limitations. To further validate the role and function of PPCDS in AML, more extensive protein level and functional experiments, such as Western blotting, co-immunoprecipitation, and cellular functional assays are required to comprehensively elucidate the specific mechanisms and roles of PPCDS.
In summary, we developed a six-gene pan-programmed cell death signature that provides novel insights into the interrelationships between cell death pathways and the clinical, molecular, and immunological landscapes of AML. Additionally, we constructed a robust PPCDS-based model capable of effectively predicting prognosis and responses to immunotherapy and targeted therapy in AML patients. Our findings indicate the potential clinical utility of PPCDS and PPCDI for prognostication and guiding personalized therapeutic decisions in AML patients undergoing immunotherapy and targeted therapies.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Sweet, K. et al. Phase I clinical trial of selinexor in combination with daunorubicin and cytarabine in previously untreated poor-risk acute myeloid leukemia. Clin. Cancer Res. 26, 54–60. https://doi.org/10.1158/1078-0432.CCR-19-2169 (2020).
Short, N. J. et al. Advances in the treatment of acute myeloid leukemia: New drugs and new challenges. Cancer Discov. 10, 506–525. https://doi.org/10.1158/2159-8290.CD-19-1011 (2020).
Kayser, S. & Levis, M. J. Updates on targeted therapies for acute myeloid leukaemia. Br. J. Haematol. 196, 316–328. https://doi.org/10.1111/bjh.17746 (2022).
Garzon, R. et al. A phase 1 clinical trial of single-agent selinexor in acute myeloid leukemia. Blood 129, 3165–3174. https://doi.org/10.1182/blood-2016-11-750158 (2017).
Appelbaum, F. R. et al. Age and acute myeloid leukemia. Blood 107, 3481–3485. https://doi.org/10.1182/blood-2005-09-3724 (2006).
Jiang, X. et al. Disruption of Wnt/β-catenin exerts antileukemia activity and synergizes with FLT3 inhibition in FLT3-mutant acute myeloid leukemia. Clin. Cancer Res. 24, 2417–2429. https://doi.org/10.1158/1078-0432.CCR-17-1556 (2018).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221. https://doi.org/10.1056/NEJMoa1516192 (2016).
Lalaoui, N., Johnstone, R. & Ekert, P. G. Autophagy and AML—food for thought. Cell Death Differ. 23, 5–6. https://doi.org/10.1038/cdd.2015.136 (2016).
Hu, D. et al. Genetic landscape and clinical significance of cuproptosis-related genes in liver hepatocellular carcinoma. Genes Dis. 11, 516–519. https://doi.org/10.1016/j.gendis.2023.03.010 (2024).
Zou, Y. et al. Leveraging diverse cell-death patterns to predict the prognosis and drug sensitivity of triple-negative breast cancer patients after surgery. Int. J. Surg. (London, England) 107, 106936. https://doi.org/10.1016/j.ijsu.2022.106936 (2022).
Piya, S., Andreeff, M. & Borthakur, G. Targeting autophagy to overcome chemoresistance in acute myleogenous leukemia. Autophagy 13, 214–215. https://doi.org/10.1080/15548627.2016.1245263 (2017).
Lu, B. et al. The Role of ferroptosis in cancer development and treatment response. Front. Pharmacol. https://doi.org/10.3389/fphar.2017.00992 (2018).
Zhu, Y., He, J., Li, Z. & Yang, W. Cuproptosis-related lncRNA signature for prognostic prediction in patients with acute myeloid leukemia. BMC Bioinform. 24, 37. https://doi.org/10.1186/s12859-023-05148-9 (2023).
Hu, D. et al. Multimolecular characteristics of cell-death related hub genes in human cancers: A comprehensive pan-cancer analysis. Cell Cycle (Georgetown, Tex.) https://doi.org/10.1080/15384101.2022.2101337 (2022).
Linkermann, A., Stockwell, B. R., Krautwald, S. & Anders, H. J. Regulated cell death and inflammation: An auto-amplification loop causes organ failure. Nat. Rev. Immunol. 14, 759–767. https://doi.org/10.1038/nri3743 (2014).
Liu, X. et al. Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nat. Cell Biol. 25, 404–414. https://doi.org/10.1038/s41556-023-01091-2 (2023).
Tang, D., Kang, R., Berghe, T. V., Vandenabeele, P. & Kroemer, G. The molecular machinery of regulated cell death. Cell Res. 29, 347–364. https://doi.org/10.1038/s41422-019-0164-5 (2019).
Mohr, S. et al. Hoxa9 and Meis1 cooperatively induce addiction to Syk signaling by suppressing miR-146a in acute myeloid leukemia. Cancer Cell 31, 549-562.e511. https://doi.org/10.1016/j.ccell.2017.03.001 (2017).
Sarraf Yazdy, M., Mato, A. R. & Cheson, B. D. Combinations or sequences of targeted agents in CLL: Is the whole greater than the sum of its parts (Aristotle, 360 BC)?. Blood 133, 121–129. https://doi.org/10.1182/blood-2018-08-869503 (2019).
Levine, B. & Kroemer, G. Biological functions of autophagy genes: A disease perspective. Cell 176, 11–42. https://doi.org/10.1016/j.cell.2018.09.048 (2019).
Koschade, S. E. et al. Translatome proteomics identifies autophagy as a resistance mechanism to on-target FLT3 inhibitors in acute myeloid leukemia. Leukemia 36, 2396–2407. https://doi.org/10.1038/s41375-022-01678-y (2022).
Cheng, C. et al. Inhibition of Nrf2-mediated glucose metabolism by brusatol synergistically sensitizes acute myeloid leukemia to Ara-C. Biomed. Pharmacother. Biomed. Pharmacother. 142, 111652. https://doi.org/10.1016/j.biopha.2021.111652 (2021).
Sun, Y., Wang, R., Xie, S., Wang, Y. & Liu, H. A novel identified necroptosis-related risk signature for prognosis prediction and immune infiltration indication in acute myeloid leukemia patients. Genes https://doi.org/10.3390/genes13101837 (2022).
Tyner, J. W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526–531. https://doi.org/10.1038/s41586-018-0623-z (2018).
Herold, T. et al. Isolated trisomy 13 defines a homogeneous AML subgroup with high frequency of mutations in spliceosome genes and poor prognosis. Blood 124, 1304–1311. https://doi.org/10.1182/blood-2013-12-540716 (2014).
Naldini, M. M. et al. Longitudinal single-cell profiling of chemotherapy response in acute myeloid leukemia. Nat. Commun. 14, 1285. https://doi.org/10.1038/s41467-023-36969-0 (2023).
Liu, Z. et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat. Commun. 13, 816. https://doi.org/10.1038/s41467-022-28421-6 (2022).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: A class discovery tool with confidence assessments and item tracking. Bioinformatics (Oxford, England) 26, 1572–1573. https://doi.org/10.1093/bioinformatics/btq170 (2010).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. https://doi.org/10.1126/scisignal.2004088 (2013).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
Wu, J. et al. A nomogram for predicting overall survival in patients with low-grade endometrial stromal sarcoma: A population-based analysis. Cancer Commun. (London, England) 40, 301–312. https://doi.org/10.1002/cac2.12067 (2020).
Blanche, P., Dartigues, J. F. & Jacqmin-Gadda, H. Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Stat. Med. 32, 5381–5397. https://doi.org/10.1002/sim.5958 (2013).
Alba, A. C. et al. Discrimination and calibration of clinical prediction models: Users’ guides to the medical literature. JAMA 318, 1377–1384. https://doi.org/10.1001/jama.2017.12126 (2017).
Zhang, J. ClusterGVis: One-step to Cluster and Visualize Gene Expression Matrix. (2022)
Tirosh, I., Izar, B. & Prakadan, S. M. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352(6282), 189–196. https://doi.org/10.1126/science.aad0501 (2016).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411. https://doi.org/10.1126/science.aax0249 (2020).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386. https://doi.org/10.1038/nbt.2859 (2014).
Zeng, D. et al. IOBR: Multi-omics immuno-oncology biological research to decode tumor microenvironment and signatures. Front. Immunol. 12, 687975. https://doi.org/10.3389/fimmu.2021.687975 (2021).
Fu, J. et al. Large-scale public data reuse to model immunotherapy response and resistance. Genome Med. 12, 21. https://doi.org/10.1186/s13073-020-0721-z (2020).
Maeser, D., Gruener, R. F. & Huang, R. S. oncoPredict: An R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief. Bioinform. https://doi.org/10.1093/bib/bbab260 (2021).
Sun, X. et al. Transcriptome profiling analysis reveals that ATP6V0E2 is involved in the lysosomal activation by anlotinib. Cell Death Dis. 11, 702. https://doi.org/10.1038/s41419-020-02904-0 (2020).
Sofer, A., Lei, K., Johannessen, C. M. & Ellisen, L. W. Regulation of mTOR and cell growth in response to energy stress by REDD1. Mol. Cell. Biol. 25, 5834–5845. https://doi.org/10.1128/mcb.25.14.5834-5845.2005 (2005).
Cheng, Z. et al. Up-regulation of DDIT4 predicts poor prognosis in acute myeloid leukaemia. J. Cell. Mol. Med. 24, 1067–1075. https://doi.org/10.1111/jcmm.14831 (2020).
Plotnik, J. P. & Hollenhorst, P. C. Interaction with ZMYND11 mediates opposing roles of Ras-responsive transcription factors ETS1 and ETS2. Nucleic Acids Res. 45, 4452–4462. https://doi.org/10.1093/nar/gkx039 (2017).
Pitarresi, J. R. et al. Stromal ETS2 regulates chemokine production and immune cell recruitment during acinar-to-ductal metaplasia. Neoplasia (New York, N.Y.) 18, 541–552. https://doi.org/10.1016/j.neo.2016.07.006 (2016).
Zaldumbide, A., Carlotti, F., Pognonec, P. & Boulukos, K. E. The role of the Ets2 transcription factor in the proliferation, maturation, and survival of mouse thymocytes. J. Immunol. 169, 4873–4881. https://doi.org/10.4049/jimmunol.169.9.4873 (2002).
Gadea, G. & Blangy, A. Dock-family exchange factors in cell migration and disease. Eur. J. Cell Biol. 93, 466–477. https://doi.org/10.1016/j.ejcb.2014.06.003 (2014).
Meyer, A. E. et al. Combinatorial genetics reveals the Dock1-Rac2 axis as a potential target for the treatment of NPM1; Cohesin mutated AML. Leukemia 36, 2032–2041. https://doi.org/10.1038/s41375-022-01632-y (2022).
Kentsis, A. et al. Aberrant expression of hepatocyte growth factor induces autocrine MET activation providing a novel therapeutic target in acute myeloid leukemia. Blood 116, 1042. https://doi.org/10.1182/blood.V116.21.1042.1042 (2010).
Yin, C. et al. High expression of CLEC11A predicts favorable prognosis in acute myeloid leukemia. Front. Oncol. 11, 608932. https://doi.org/10.3389/fonc.2021.608932 (2021).
Acknowledgements
We gratefully acknowledge all patients who participated in our study as well as the support from their families, and we also thank the investigators and staff in this clinical study for their valuable contributions.
Funding
This study was supported by the Key Research and Development Program of Anhui Province (Grant No. 202104j0702).
Author information
Authors and Affiliations
Contributions
DTH designed research, XXP performed research, YQ analyzed data, and was a major contributor in writing the manuscript, MZY performed Writing—Review & Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qin, Y., Pu, X., Hu, D. et al. Machine learning-based biomarker screening for acute myeloid leukemia prognosis and therapy from diverse cell-death patterns. Sci Rep 14, 17874 (2024). https://doi.org/10.1038/s41598-024-68755-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-68755-3
- Springer Nature Limited