Abstract
Alzheimer’s disease (AD) and post-stroke cognitive impairment (PSCI) are the leading causes of progressive dementia related to neurodegenerative and cerebrovascular injuries in elderly populations. Despite decades of research, patients with these conditions still lack minimally invasive, low-cost, and effective diagnostic and treatment methods. MicroRNAs (miRNAs) play a vital role in AD and PSCI pathology. As they are easily obtained from patients, miRNAs are promising candidates for the diagnosis and treatment of these two disorders. In this study, we performed complete sequencing analysis of miRNAs from 24 participants, split evenly into the PSCI, post-stroke non-cognitive impairment (PSNCI), AD, and normal control (NC) groups. To screen for differentially expressed miRNAs (DE-miRNAs) in patients, we predicted their target genes using bioinformatics analysis. Our analyses identified miRNAs that can distinguish between the investigated disorders; several of them were novel and never previously reported. Their target genes play key roles in multiple signaling pathways that have potential to be modified as a clinical treatment. In conclusion, our study demonstrates the potential of miRNAs and their key target genes in disease management. Further in-depth investigations with larger sample sizes will contribute to the development of precise treatments for AD and PSCI.
Similar content being viewed by others
Introduction
Dementia encompasses a range of syndromes characterized by a decline in cognitive and executive abilities, significantly affecting daily life1. Globally, over 50 million people suffer from dementia, and the number of patients is projected to rise to 152 million by 20502. In younger individuals, they are often associated with severe brain trauma and infections. Among the elderly, common causes include Alzheimer's disease (AD), post-stroke cognitive impairment (PSCI), vascular dementia (VaD), and dementia with Lewy bodies (DLB). Dementia prevalence has seen an increase in the older population (≥ 60 years old), with the age-standardized rate ranging between 5 and 7%. Notably, this rate approximately doubles every 5 years3. Unfortunately, the diagnosis and treatment of both AD and PSCI present major challenges. Genetic research is crucial for understanding AD/PSCI etiology and paving the way for novel therapeutic options.
Approximately 60–80% of projected dementia cases are attributed to AD2. Because pathological changes in AD occur decades before the symptoms are detectable, early diagnosis is key to effective treatment. The primary characteristic of AD is impaired clearance of β-amyloid (Aβ), and the formation and accumulation of hyperphosphorylated tau protein tangles. These pathological features arise from abnormal processing of amyloid precursor protein by β-/γ-secretases, leading to the production and aggregation of Aβ40 and Aβ42 monomers4,5. Hyperphosphorylated tau proteins contribute to synaptic disruption, neuronal dysfunction, and development of neurofibrillary tangles6. In addition, cerebrovascular lesions, oxidative stress (OS), neuroinflammation, and abnormal glucose metabolism all combine to play complex roles in AD initiation and progression7. Currently, AD diagnosis relies on neuropsychological and neuroimaging assessments, biomarker-based lumbar puncture, and positron emission tomography (PET). These methods are both costly and invasive, limiting their clinical implementation. Furthermore, available treatments currently can only address symptoms, while they can improve quality of life temporarily, therapies that halt or reverse disease progression have not been developed, despite over 2000 clinical trials testing various targets and treatment strategies8.
Research on non-coding RNAs (ncRNAs) have advanced rapidly in recent years, especially with the development of high-throughput sequencing, whole-gene sequencing, and bioinformatics. Of particular interest are microRNAs (miRNAs), 18–21 nucleotides that regulate the expression of downstream genes through cleaving or post-transcriptionally inhibiting target mRNAs. Through this process, miRNAs modulate numerous intracellular and extracellular signaling pathways in the neocortex, hippocampus, and limbic systems. Specifically, miRNAs are involved in regulating tight connections in the blood–brain barrier (BBB), affecting neuronal growth, proliferation, and differentiation. They also participated in the regulation of synaptic homeostasis and plasticity9,10,11. As the CNS ages, miRNA abundance alters in accordance with neural development and differentiation12,13,14. In-depth analysis of miRNAs is hoped to uncover novel pathogenic pathways or pathogenic factors in known pathways that can benefit AD therapies.
Compared with PET or protein analysis of cerebrospinal fluid, the major advantage of miRNAs is their lack of invasiveness while providing high volumes of data. A recent systematic review cross-validated 250 miRNAs associated with AD, highlighting 10 miRNAs (miR-107, miR-26b, miR-30e, miR-34a, miR-485, miR-200c, miR-210, miR-146a, miR-34c, and miR-125b) in the peripheral blood of patients15. These were hypothesized to be deregulated 20 years before clinical symptom onset15 and are associated with major biological processes, including immune system function, cell cycle regulation, gene expression, neuron growth factor signaling, Wnt signaling, cellular senescence, and Rho GTPase activity. Of these, the involvement of miRNAs in autophagy16 and neuronal mitochondria are particularly relevant to AD etiology17. Notably, miR-455-3p upregulation enhances synaptic activity, suggesting a potential neuroprotective role18. In contrast, miR-34a dysregulation causes dysfunction in resting presynaptic and postsynaptic activity19. Other specific links between miRNAs and AD include miR-29c-3p, miR-193b, miR-132, miR-103, and miR-181c regulating Aβ activity; miR-181c, miR-31, miR-9320, and miR-14621 modulating microglial inflammatory response in the CNS; as well as miR-125b22, miR-26b, miR-483-5p, and miR-502-3p influencing tau protein activity and NFT formation23. Additionally, miR-532-5p targets EPHA4 to ameliorate BBB damage24, while miR-126 targets EFHD2 to promote the formation and modulation of contextual fear memory in patients with AD25. The available data all point to the importance of miRNAs in influencing AD onset and symptoms.
To be classified as PSCI, patient performance on a multidomain neuropsychological assessment or the Montreal Cognitive Assessment must be lower than the fifth percentile of local normative data in at least one cognitive domain26. The condition is a common consequence of ischemic stroke that causes long-term disability and lowers quality of life27,28. Among dementias, PSCI has the second highest incidence after AD29. Pooled data analyses indicated a prevalence of 38%30 PSCI and 18.4%31 post-stroke dementia in the first year after stroke32. Although PSCI is largely attributable to vascular factors, neurodegenerative changes are also common in elderly patients and can interact with PSCI33. A pivotal prerequisite for PSCI is cerebral small vessel disease (CSVD), characterized by arteriolar sclerosis, lacunar infarction, cortical and subcortical microinfarction, as well as pathological changes to white matter (including myelin loss and axonal abnormalities)34. Post-stroke inflammation may also play a role in amyloid deposition during PSCI development35.
Diagnosis of PSCI currently relies on clinical symptoms, neuropsychology, and neuroimaging. Deep learning (DL)-based cognitive evaluation using FDG-PET is an important technique for assessing cognitive dysfunction36. Infarct location and stroke characteristics can predict PSCI independently of demographic and vascular risk factors, particularly the presence of multiple acute infarcts, total infarct volume, and left cerebral hemispheric location27,37. In terms of PSCI biomarkers, numerous potential targets have been proposed, including indicators of genetic polymorphisms, biomarkers in the cerebrospinal fluid and plasma, inflammatory mediators, and peripheral miRNAs38. However, thus far, none can discriminate vulnerable patients robustly.
Research using TaqMan Low-Density Arrays and single TaqMan assays have demonstrated that miR-10b, miR29a-3p, and miR-130b-3p are downregulated in the plasma of patients with PSCI and AD, especially miR-130b-3p in AD39. Experiments rats showed that intravenous (iv) injections of miR-20a-3p improved acute stroke outcomes in male and female rats, while also ameliorating stroke-induced cognitive decline in female rats40. Additionally, intracerebroventricular (ICV) injection of miR-210-5p agonist aggravated synaptic loss and cognitive impairment in VD rats, specifically through regulating synaptosomal-associated protein of 25 KDa41. Serum miRNA profiling of patients and healthy controls identified seven miRNAs (miR-1228-5p, miR-1268a, miR-1268b, miR-4433b-3p, miR-6090, miR-6752-5p, and miR-6803-5p) that could predict the risk of cerebrovascular disorders before stroke onset42. Furthermore, acupuncture appeared to attenuate inflammation-related PSCI through inhibiting the miR-93-mediated TLR4/MyD88/NF-κB signaling pathway20. Similarly, the TNFα–miR-501-3p–ZO-1 axis modulates BBB to affect working memory deficits and white matter lesions43. Overall, therapies based on miRNAs may provide novel insights into inhibiting PSCI progression.
Although numerous preclinical studies involving miRNA therapeutics have been conducted, only a few have moved into clinical development. One of the biggest challenges in developing these treatments is the identification of suitable miRNAs and their targets for each disease type44,45,46. In this study, we aimed to provide more candidates through high-throughput sequencing, screening, and functional analysis of differentially expressed miRNAs (DE-miRNAs). Our findings should expand current understanding of miRNAs related to cognitive decline and provide options for further research on miRNA-based treatment.
Results
Characterization of participants
Medical histories did not differ across the AD, PSCI, post-stroke non-cognitive impairment (PSNCI), and normal control (NC) groups. Sex ratios, exhibited between-group variation (P < 0.05), but pair-to-group comparisons after Bonferroni correction showed no significant differences between pairs (Table S1). Upon assessment, we identified between-group differences in the MMSE, MoCA, and various cognitive domains (P < 0.05) (Table 1).
Bioinformatics analysis of AD and control (NC)
We identified 2263 common miRNAs between the AD and NC groups (Fig. 1A; Table S2). With thresholds of P < 0.05 and |log2FC|≥ 1, we found 14 upregulated and 17 dysregulated DE-miRNAs (Fig. 1B). Of these 31 DE-miRNAs, relevant data for hsa-miR-320a-5p, hsa-miR-519a-2-5p, hsa-miR-520b-5p, and hsa-miR-10400-3p were not retrieved from the TargetScan Human database. The final analysis included 27 DE-miRNAs that targeted 384 genes.
Bioinformatics analysis comparing Alzheimer’s disease (AD) and normal controls (NC). (A) Sequencing cluster map between the AD and NC groups. Different colors indicate relative expression levels (log2-transformed). Blue, below-mean expression; red, above-mean expression. The colored bar at the top of the panel represents participating groups (blue represents AD and red represents NC). The colored bar on the right side of the panel indicates divisions based on K-means. (B) Volcano plot of DE-miRNAs between AD and NC groups. On the x-axis, the dotted line indicates DE-miRNAs that satisfied the |log2FC|≥ 1 cut-off. On the y-axis, the dotted line indicates DE-miRNAs that satisfied P < 0.05. Red, highly expressed; purple, lowly expressed. (C) Gene Ontology (GO) enrichment results between AD and NC. P values are on the X-axis (significance of associations between gene sets and GO terms). The Y-axis displays GO terms. (D) The KEGG47,48,49 pathway analysis between AD and NC. Gene ratio is proportion of genes in a pathway to total genes analyzed (x-axis). Signaling pathways are on the y-axis. Bubble size indicates the number of genes enriched in a given pathway. Bubble color reflects significance level (bluer is smaller). P-values are log-transformed.
After gene set enrichment analyses (GSEA), we restricted our display to the top 10 Gene Ontology (GO) terms per category and the principal 10 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways (Fig. 1C,D; Table S3 and S4). Target genes were significantly enriched in the biological processes of mesenchymal and fibrocyte proliferation, neurogenesis, mRNA processing, stem cell proliferation, and nervous system development. They were located in multiple organelles, including cytoplasmic stress particles, transcription inhibitor complexes, nuclide spots, nuclear specks, integral components of the endoplasmic reticulum membrane, and apical plasma membrane. In terms of molecular function, target genes were involved in protein cysteine S-acyltransferase activity, protein serine muscle enzyme activity regulation, insulin-like growth factor 1, and growth factor-binding hormone. The results of KEGG analysis suggested that target genes mainly participated in the phosphatidylinositol 3-kinase (PI3K)-Akt, MAPK, and FoxO signaling pathways, as well as in small cell lung cancer and prostate cancer (Fig. 1D).
We then constructed a protein–protein interaction (PPI) network of DE-miRNA target genes, comprising 364 gene/protein nodes and 655 non-redundant interacting edges (Fig. 2A). Analysis in MOCDE and CytoHubba yielded nine key genes: ATM, MYC, PTEN, CDKN1B, CCNB1, HIF1A, CDK2, BMI1, and IGF1R. Analysis of interactions between DE-miRNAs and key genes (Fig. 2B) revealed that a single DE-miRNA has multiple targets, while one gene can affect several downstream signaling pathways.
Target genes of DE-miRNAs between AD and NC. (A) STRING visualization of the protein–protein interaction (PPI) network. The darker the color, the larger the font, and the larger the background area, the greater the weight of this gene in the PPI network and its correlation with other genes. (B) DE-miRNAs plus their key target genes.
Bioinformatics analysis of AD and PSCI
We identified 2242 miRNAs that were common across AD and PSCI groups (Table S2; Fig. 3A). Of these, 22 were DE-miRNAs (10 upregulated and 12 dysregulated) (Fig. 3B). Using biogenic analysis, we then conducted GSEA on DE-miRNA target genes (Table S5 and S6); the top 10 GO terms per category and top 10 KEGG pathways are shown (Fig. 3C,D). Target genes participate in the positive regulation of cellular catabolic processes, nuclear-transcribed mRNA poly(A) tail shortening and viral life cycle, negative regulation of cellular macromolecule biosynthetic processes, and cellular amide metabolic processes. Additionally, they are involved in the cellular components of growth cones, asymmetric synapses, neuron-to-neuron synapses, and postsynaptic specialization. Finally, the target genes play a role in regulating enzyme inhibitor activity, protein kinase activity, kinase activity, protein phosphatase inhibitor activity, DNA-binding transcription activator activity, and molecular adaptor activity. The results of KEGG analysis indicated that these genes are mainly involved in cell catabolism, senescence, and endocytosis, as well as the Hippo and p53 signaling pathways. Key target genes of these DE-miRNAs are CDKN1A, CDK6, and CCND2 (Fig. 4A,B).
Bioinformatics analysis comparing Alzheimer’s disease (AD) and post-stroke cognitive impairment (PSCI). (A) Sequencing cluster map between the AD and PSCI groups. Different colors indicate relative expression levels (log2-transformed). Blue, below-mean expression; red, above-mean expression. The colored bar at the top of the panel represents participating groups (blue represents AD and red represents PSCI). The colored bar on the right side of the panel indicates divisions based on K-means. (B) Volcano plot of DE-miRNAs between AD and PSCI groups. On the x-axis, the dotted line indicates DE-miRNAs that satisfied the |log2FC|≥ 1 cut-off. On the y-axis, the dotted line indicates DE-miRNAs that satisfied P < 0.05. Red, highly expressed; purple, lowly expressed. (C) Gene Ontology (GO) enrichment results between AD and PSCI. P values are on the X-axis (significance of associations between gene sets and GO terms). The Y-axis displays GO terms. (D) The KEGG47,48,49 pathway analysis between AD and PSCI. Gene ratio is proportion of genes in a pathway to total genes analyzed (x-axis). Signaling pathways are on the y-axis. Bubble size indicates the number of genes enriched in a given pathway. Bubble color reflects significance level (bluer is smaller). P-values are log-transformed.
Target genes of DE-miRNAs between AD and PSCI. (A) STRING visualization of the protein–protein interaction (PPI) network. The darker the color, the larger the font, and the larger the background area, the greater the weight of this gene in the PPI network and its correlation with other genes. (B) DE-miRNAs plus their key target genes.
Bioinformatics analysis of PSCI and PSNCI
We identified 2238 common miRNAs across the PCI and PSNCI groups (Table S2; Fig. 5A). Of these, 22 DE-miRNAs were found (11 upregulated and 11 dysregulated) (Fig. 5B), but data for hsa-miR-10401-5p was absent from the TargetScan Human database, resulting in 21 DE-miRNAs used for target gene prediction (Table S7 and S8). Only the top 10 GO terms and the top 10 KEGG pathways are displayed (Fig. 5C,D). The results of GO analysis showed that target genes were enriched in protein dephosphorylation-related biological progression. They are localized to the synaptic vesicle membrane, presynaptic active zone cytoplasmic component, and cell cortex region. Additionally, they participate in metabolic progresses related to small GTPase binding, GDP binding, GTPase activity, acidic amino acid transmembrane transporter activity, and hormone binding. Next, the results of KEGG analysis showed that target genes are involved in FoxO, AMPK, and P53 signaling pathways. Construction of a PPI network then identified BMI1, CDK2, CCNB1, KRAS, CCND1, CDK6, MYCN, AR, E2F3, and GSK3B as most important genes interacting with DE-miRNAs (Fig. 6A,B).
Bioinformatics analysis comparing post-stroke cognitive impairment (PSCI) and normal post-stroke non-cognitive impairment (PSNCI). (A) Sequencing cluster map between the PSCI and PSNCI groups. Different colors indicate relative expression levels (log2-transformed). Blue, below-mean expression; red, above-mean expression. The colored bar at the top of the panel represents participating groups (blue represents PSCI and red represents PSNCI). The colored bar on the right side of the panel indicates divisions based on K-means. (B) Volcano plot of DE-miRNAs between PSCI and PSNCI groups. On the x-axis, the dotted line indicates DE-miRNAs that satisfied the |log2FC|≥ 1 cut-off. On the y-axis, the dotted line indicates DE-miRNAs that satisfied P < 0.05. Red, highly expressed; purple, lowly expressed. (C) Gene Ontology (GO) enrichment results between PSCI and PSNCI. P values are on the X-axis (significance of associations between gene sets and GO terms). The Y-axis displays GO terms. (D) The KEGG47,48,49 pathway analysis between PSCI and PSNCI. Gene ratio is proportion of genes in a pathway to total genes analyzed (x-axis). Signaling pathways are on the y-axis. Bubble size indicates the number of genes enriched in a given pathway. Bubble color reflects significance level (bluer is smaller). P-values are log-transformed.
Target genes of DE-miRNAs between PSCI and PSNCI. (A) STRING visualization of the protein–protein interaction (PPI) network. The darker the color, the larger the font, and the larger the background area, the greater the weight of this gene in the PPI network and its correlation with other genes. (B) DE-miRNAs plus their key target genes.
Comparison of DE-miRNAs and target genes across three groups
Three DE-miRNAs (hsa-miR-519a-2-5p, hsa-miR-520b-5p, hsa-miR-5703) were differentially expressed in both AD vs. NC and AD vs. PSCI. Four DE-miRNAs (hsa-miR-5582-3p, hsa-miR-8485, hsa-miR-6131, hsa-miR-449b-5p) were differentially expressed in AD vs. PSCI and PSCI vs. PSNCI. Only one DE-miRNA (hsa-miR-5186) was differentially expressed in AD vs. NC and PSCI vs. PSNCI (Fig. 7A; Table S9). Notably, no DE-miRNAs were common across all three groups, but target gene expression exhibited between-group variation (Fig. 7B; Table S10). Our analysis identified 38 differentially expressed target genes for AD vs. NC and AD vs. PSCI, 39 for AD vs. PSCI and PSCI vs. PSNCI, and 47 for AD vs. NC and PSCI vs. PSNCI. Thirteen target genes were differentially expressed across all three comparative pairs.
Venn diagrams representing DE-miRNAs and target genes common across all three groups with cognitive impairments. (A) DE-miRNAs between patients with different cognitive impairments (AD, PSCI, PSNCI) and healthy controls (NC). Red circle, AD and NC; orange circle, AD and PSCI; green circle, PSCI and PSNCI. (B) Differentially expressed target genes between patients with different cognitive impairments and healthy controls. Blue circle, AD and NC; purple circle, AD and PSCI; yellow circle, PSCI and PSNCI.
Discussion
By collecting data from patients with different dementia subtypes and comparing plasma miRNA expression, our research successfully identified novel miRNAs related to AD and PSCI. Moreover, we linked those miRNAs to their target genes, providing considerable clarity on the molecular mechanisms underlying AD and PSCI.
With the development of genome-wide association studies (GWAS) and single-cell sequencing technologies, miRNAs are increasingly relevant as biomarkers for non-invasive diagnosis and treatment. Different miRNA combinations have been investigated for their suitability in diagnosing cognitive disorders, including AD and PSCI50. Similarly, our target gene prediction and GSEA identified 14 DE-miRNAs. Of these, hsa-miR-548as-5p, hsa-miR-5186, and hsa-miR-449b-5p were differentially expressed across all three patient groups. Functional analysis with KEGG demonstrated that DE-miRNAs participate in AD and PSCI progression through their target genes in the cell cycle, cell senescence processes, as well as FoxO, PI3K, Hippo, and p53 signaling pathways. We also identified several novel DE-miRNAs, including hsa-miR-548as-5p, hsa-miR-519c-3p, hsa-miR-6883-5p, hsa-miR-5186, hsa-miR-548ak, and hsa-miR-147a. The remaining DE-miRNAs have been implicated in infectious diseases51, endocrine diseases52 and neoplastic diseases53,54,55,56, but not in CNS diseases or any cognitive disorders. Notably, changes in cellular developmental pathways do not yield the same outcome in every disease; in cancers, they cause rapid cell growth and proliferation, whereas in CNS disorders, they accelerate neuron aging and death. Understanding the specific characteristics of neoplastic diseases will be useful for producing novel interventions and targets in AD and PSCI57.
We compared DE-miRNAs and target genes across three pairs of groups (AD vs. NC, AD vs. PSCI, and PSCI vs. PSNCI) to better understand which miRNA expression patterns are common or unique across disease states. None of the identified DE-miRNAs were present in all three groups. However, has-miR-5582-3, hsa-miR-8485, hsa-miR-6131, hsa-miR-449b-5p, and hsa-miR-5186 were important for both AD and PSCI development, while hsa-miR-519a-2-5p, hsa-miR-520b-5p, and hsa-miR-5703 were unique to AD, allowing for differential diagnosis from NC and PSCI. Furthermore, we identified 13 target genes (STK35, GRID1, ZNF268, PARP11, RAB5B, PDE7A, RNF40, KCND1, UBN2, TTPAL, CACNG8, ORAI2, and XIAP) that may be vital in AD and PSCI development.
In AD, Aβ deposition and tau neurofibrillary entanglement are classic pathological changes that lead to amyloid precursor deposition, autophagy, apoptosis, and mitochondrial dysfunction. Interestingly, post-stroke degenerative gene expression in hippocampal neurons cause similar pathological changes58, suggesting that gene expression patterns may be the source of mechanistic overlap between AD and PSCI. In particular, p53 regulation of miRNAs is responsive to stressors, thus influencing gene stability and protein function; both Aβ deposition and tau hyperphosphorylation in the CNS can be attributed to p5359,60,61,62. Indeed, the p53-miRNA interaction has been widely researched as a target for treating neurodegenerative diseases. Our study identified hsa-miR-548as-5p, hsa-miR-5186, hsa-miR-548ak, hsa-miR-147a, hsa-miR-6868-3p, and hsa-miR-449b-5p as part of the p53-related signaling pathway.
Research on other diseases and related signaling mechanisms have helped us understand the pivotal roles of several DE-miRNAs in the occurrence of AD and PSCI. First, hsa-miR-187-5p, hsa-miR-3157-5p, and hsa-miR-519c-3p—upregulated in the AD group—are implicated in the PI3K signaling pathway. This is a lipid kinase that participates in the intracellular signaling cascade, as well as cell silencing, survival, and growth under physiological and disease states63,64. Mesenchymal stem cell-derived exosomal miR-223 can protect neurons from apoptosis through protein tyrosine phosphatase (PTEN), which targets the PI3K/Akt pathway65. Additionally, hsa-miR-4655-5p and hsa-miR-3157-5p are involved in regulating mitogen-activated protein kinase (MAPKs) pathways. In mammals, MAPKs include c-Jun NH2-terminal kinase (JNK), p38 MAPK, and extracellular signal-regulated kinase (ERK). The pathway is implicated in cancer and neurodegenerative diseases through its regulation of cell proliferation, differentiation, and intrinsic immunity66. In particular, p38 MAPK affects microglia- and astrocyte-mediated neuroinflammation. This process has been linked to RNase III enzyme Drosha, which accelerates miRNA maturation. Evaluation of postmortem tissues from AD patients and AD rat models suggests that Aβ oligomers induce p38 MAPK-dependent phosphorylation of Drosha, leading to its bidirectional distribution between the neuronal cytoplasm and nucleus. Drosha overexpression can protect neurons from Aβ oligomeric-induced apoptosis67. Furthermore, MAPK can affect glucose metabolism in the CNS, promote catecholamine uptake, regulate neuronal growth, and alter neurotransmitter GABA expression via modulating insulin signal transduction pathways68. Triggering these processes can potentially improve cognitive function in patients with AD and PSCI.
Key genes were screened to further explore the mechanisms underlying AD and PSCI. The tumor suppressor PTEN is regulated by hsa-miR-519c-3p and mediates disease progression through activating the PI3K/serine-threonine kinase (AKT) signaling pathway. Pathological tau protein and intracellular cholesterol enrichment trigger premature PTEN activation, accelerating the microglia toxicity that occurs when synapses or neurons are exposed to phosphoserine apoptotic signals69,70. In vitro and in vivo AD models have confirmed that PTEN inhibition can effectively alleviate synaptic and cognitive damage71. In particular, the isoform PTENα can affect olfactory behavior in AD mice and regulate the endocytosis of olfactory bulb neurons through direct dephosphorylation of endocytosis proteins, a phenomenon also observed in patients with Parkinson's disease and DLB72. In addition to PTENα, PTEN-induced kinase 1 (PINK1) controls the specific clearance of dysfunctional or excess mitochondria through selective autophagy regulation55. PINK1 overexpression can reverse abnormal changes in mitochondrial dynamics, autophagy defects, and abnormal energy metabolism in the hippocampus. Other functions of PINK1 include upregulating antioxidant proteins to lower oxidative stress and inhibiting stress-induced neuronal apoptosis via the Nrf2 pathway73. In an AD mouse model, plasma PTEN-PDZ complex is relatively stable and crosses the BBB to become PTEN, resulting in synaptic protection and improved cognitive function. These two characteristics (plasma stability and BBB permeability) make the PTEN-PDZ complex a promising candidate for parenteral (intravenous and intramuscular) and naso-cerebral administration74.
Transcription factors regulate miRNA production in response to external stimuli and are especially active in the brain75. Under ischemia and hypoxia, transcription factors inactivate neurons within a few minutes, causing serious brain dysfunction. Sensitivity to damage increases with increasing age and greater impairment to the energy transport capacity of cells. In this study, HIF1A (regulated by hsa-miR-519c-3p) encodes the alpha subunit of hypoxia-inducible factor-1 (HIF-1). As a key transcription factor, HIF-1 is an important mediator of cellular and systemic homeostasis. HIF-1α upregulates BACE1 and γ-secretase in ischemic anoxic mouse models and primary neuron culture, inducing Aβ production during brain tissue hypoperfusion or hypoxia76. Likewise, the mTOR-HIF1α complex induces acute inflammation in microglia exposed to Aβ. Metabolic reprogramming from oxidative phosphorylation to glycolysis eventually causes abnormal metabolism, cytokine secretion, and phagocytosis77,78. In addition to its effect on Aβ, HIF-1α also influences tau neurofibrillary tangles under chronic hypoxia. In SD mouse hippocampal neurons after hypoxia treatment, an increase in HIF-1α is accompanied by tau protein hyperphosphorylation, along with a decrease in protein phosphatase 2A and leucin carboxymethyltransferase 1. These molecular changes are linked to cognitive impairment among SD mice and were verified in primary hippocampal neurons and C6/tau cells79. Thus, inhibiting HIF-1α may alleviate hypoxia-related cognitive impairment. Moreover, HIF1α could also cause remission of some PSCI symptoms, specifically irreversible white matter and neuronal damage. The mechanism involves HIF1α activation of the PKA pathway to regulate angiogenesis induced by vascular endothelial growth factor and erythropoietin72,80.
Another important target gene we identified here is the neuronal gene BMI181. Whole-genome sequencing on samples from patients with late-onset AD revealed that BMI expression decreased, in conjunction with changes to Aβ and P-tau82. BMI1 inhibits the transcription of tau-associated tubulin, and knocking out BMI1 in neurons leads to Aβ secretion/deposition, P-Tau aggregation, and neurodegenerative changes83. A clinical cohort study comprising 1565 patients with AD found a significant correlation between BMI1 expression and Aβ1-42 levels in cerebrospinal fluid83. Taken together, available data strongly implicate BMI1 as a potential therapeutic target for AD.
The widespread use of miRNAs as a non-invasive screening method has revealed that a single miRNA or miRNA complex in plasma possesses differential diagnostic efficacy. A key obstacle in identifying relevant target miRNAs is the considerable heterogeneity in miRNA expression. Our study identified several miRNAs that are entirely novel or have only been minimally documented in a few publications associated with non-neurological disorders. Thus, little data are available on whether they can truly serve as biomarkers of PSCI or AD. In AD, bidirectional interactions between neuroinflammatory OS and the proteins Aβ and tau could also influence miRNA expression. Moreover, key miRNA synthesis enzymes such as Drosha, Dicer, and AGO2 are downregulated by stressful conditions such as hypoxia, causing aberrant miRNA expression that may interfere with diagnostic accuracy84,85,86.
One notable limitation of plasma miRNA sequencing is that it only reflects miRNA expression of the disease stage at sampling. While such data can be applied to differential diagnosis and prognosis, it obviously does not reflect miRNA expression patterns throughout the disease course. In this study, miRNA expression represented the post-symptomatic state because only clinically diagnosed participants were included. However, DE-miRNAs have considerable potential for diagnosis before requiring patients to undergo expensive and invasive examinations such as PET scans and lumbar punctures. Beyond an earlier diagnosis, DE-miRNA data also provides more information to clinicians regarding specific disease types, suggesting major benefits in addressing current shortcomings. To overcome the obstacles of heterogeneous miRNA expression, samples at different stages of disease progression are needed to identify common regulatory miRNAs that can then be therapeutically manipulated87.
Therapies based on miRNAs are currently in the form of mimics or inhibitors to influence downstream signaling pathways. However, further research is needed to determine optimal drug delivery in humans. In the future, anti-microRNA (AM) strategies could be implemented for the amelioration and clinical management of AD. This novel therapeutic approach restores multiple miRNA-regulated gene targets via the use of selectively stabilized AM species. Moreover, only a relatively small amount of miRNA families need to be modulated or neutralized to re-establish neuronal homeostasis in the AD-affected brain88.
In conclusion, through high-throughput sequencing and bioinformatics, we identified novel miRNAs, their key target genes, and highlighted potential mechanisms of their involvement in AD and PSCI. Importantly, our findings provided an empirical basis for using miRNAs as peripheral biomarkers and therapeutic targets.
Methods
Participants
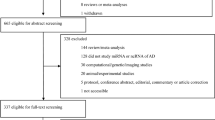
The study selected 18 patients admitted to the Department of Neurology at the First Hospital of Jilin University between 2021 and 2022. Based on etiology and MoCA, patients were divided into three groups (n = 6 per group): AD, PSCI, and post-stroke non-cognitive impairment (PSNCI). Six normal controls (NC) were also included.
Patients were included in the PSCI or PSNCI group if they were 50–80 years of age, met the WHO diagnostic criteria for acute ischemic stroke (AIS), and could complete the relevant scale assessment. They were further subdivided into PSNCI if MoCA ≥ 22 and PSCI if MoCA < 2289. Patients were included in AD group if they met the NINCDS-ADRDA diagnostic criteria and confounding factors were excluded. Finally, spouses of the patients were included as NC, matched by age, sex, education and previous medical history.
Individuals were excluded if they exhibited other CNS conditions that cause cognitive impairment, including metabolic encephalopathy, Parkinson's syndrome, Huntington's disease, subdural hematoma, normal cranial pressure hydrocephalus, brain tumors, and brain trauma. Displaying other systemic conditions linked to cognitive decline also resulted in exclusion; these included hypothyroidism, vitamin (B12, folic acid, niacin) deficiency, severe anemia, liver or renal insufficiency, hyponatremia, hypocalcemia, neurosyphilis, HIV infection, and alcohol/drug abuse. Finally, disorders or any other characteristics that negatively impacted the ability to perform tasks and scale operations led to exclusion. These included mental disorders (e.g., depression and schizophrenia), nuclear magnetic contraindications, refusal of blood collection, serious diseases of vital organs, and sensory impairments (e.g., blindness, aphasia, deafness).
Data collection
Patient age, sex, education, other demographics, and related medical history (e.g., hypertension, diabetes, coronary heart disease) were collected. Professional neurologists conducted all neuropsychological assessments of participants to obtain relevant baseline data. After an 8-h fasting period, peripheral venous blood samples were collected, and plasma was separated for storage at − 80 °C.
Statistical analysis
Statistical analyses were performed in IBM SPSS Statistics version 25 (IBM, Armonk, NY, USA) and R version 4.3.2 (R Foundation for Statistical Computing, Vienna, Austria). Sociodemographic and neuropsychological characteristics were subjected to descriptive analyses. Continuous variables were presented as mean ± SD or median (interquartile range), and categorical variables were presented as frequencies (percentages). Between-group differences were compared with chi-square tests, one-way ANOVA, or Kruskal–Wallis rank sum test. Significance was set at P < 0.05.
Microarray assay of miRNA
Human tissue specimens were collected from the Department of Biobank, Division of Clinical Research, for RNA extraction using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). MiRNA sequencing was performed using an Illumina NextSeq 500 sequencing platform (San Diego, CA, USA). Double-stranded cDNA was synthesized using the dUTP method and high-fidelity PCR polymerase to ensure strand specificity. Constructed libraries were assessed in an Agilent 2100 Bioanalyzer and subjected to quantitative real-time PCR (qRT-PCR).
Based on quantification results, libraries were mixed in equal amounts and used for sequencing. Single-stranded DNA (1.8 pM) generated from denaturing with 0.1 M NaOH were loaded onto the reagent cartridge. Sequencing was performed for 50 cycles on a NextSeq system using the NextSeq 500/550 V2 kit (#FC-404-2005, Illumina). The Solexa pipeline v1.8 (Off-Line Base Caller v1.8) was used for image analysis and base calling. Sequencing quality was examined with FastQC.
Expression profiles of miRNAs were calculated based on mapped read counts. Differentially expressed miRNAs (P < 0.05, |log2FC|≥ 1) were screened in R package edgeR90, then matched to reference sequences in miRDeep291. The expression of miRNAs was represented as counts per million mapped reads (CPM) after transformation.
Bioinformatics analysis
Sequencing cluster maps of the three groups were generated with R (v4.3.2) packages ggplot2 and k-means clustering. To select DE-miRNA, volcano plots were created with ggplot2. Next, DE-miRNA target genes were predicted with reference to three databases: MiRDIP (http://ophid.utoronto.ca/mirDIP/index.jsp)92, miRNet93 (https://www.mirnet.ca/), and TargetScan Human (https://www.targetscan.org/vert_80/)94. Hits in all three databases comprised the final list of target genes.
Functional analyses with GO95 and KEGG96 were visualized using R packages ClusterProfiler and ggplot2. Significant enrichment (P < 0.05 with Benjamin-Hochberg corrections) was determined with Fisher’s exact test. Next, the PPI network of target genes was generated in STRING (https://cn.String-db.org/) and visualized in Cytoscape (v3.9.1)97. Target genes were processed in MCODE (node score cutoff: 0.2, K-Core: 2, Max. Depth: 100) and CytoHubba (MCC algorithm) to identify key genes (those that passed both sets of conditions). Finally, a network of signaling pathways involving DE-miRNAs and their key genes was constructed.
Ethical approval and informed consent
All methods were performed in accordance with the relevant guidelines and regulations (For ex. Declaration of Helsinki). The studies involving human participants were reviewed and approved by the ethics committee of The First Hospital of Jilin University (ethics number: 19K023-003). All subjects provided written informed consent to participate in this study. The sequencing information of all participants has been filed with the China Center for Human Genetic Resources Management.
Data availability
All microarray data used in this study are publicly available through the Gene Expression Omnibus (GEO) database, accession numbers: GSE218605 (https://www.ncbi.nlm.nih.gov/geo/). The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2021) in the National Genomics Data Center (Nucleic Acids Res 2022), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA-Human: HRA003714), which are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human98,99.
References
Gale, S. A., Acar, D. & Daffner, K. R. Dementia. Am. J. Med. 131, 1161–1169. https://doi.org/10.1016/j.amjmed.2018.01.022 (2018).
2022 Alzheimer's disease facts and figures. Alzheimers Dement 18, 700-789, https://doi.org/10.1002/alz.12638 (2022).
Ferencz, B. & Gerritsen, L. Genetics and underlying pathology of dementia. Neuropsychol. Rev. 25, 113–124. https://doi.org/10.1007/s11065-014-9276-3 (2015).
Cortes-Canteli, M. & Iadecola, C. Alzheimer’s disease and vascular aging: JACC focus seminar. J. Am. Coll Cardiol. 75, 942–951. https://doi.org/10.1016/j.jacc.2019.10.062 (2020).
Tiwari, S., Atluri, V., Kaushik, A., Yndart, A. & Nair, M. Alzheimer’s disease: Pathogenesis, diagnostics, and therapeutics. Int. J. Nanomedicine 14, 5541–5554. https://doi.org/10.2147/IJN.S200490 (2019).
Belloy, M. E., Napolioni, V. & Greicius, M. D. A quarter century of APOE and Alzheimer’s disease: Progress to date and the path forward. Neuron 101, 820–838. https://doi.org/10.1016/j.neuron.2019.01.056 (2019).
Alzheimer’s, A. 2016 Alzheimer’s disease facts and figures. Alzheimers Dement 12, 459–509. https://doi.org/10.1016/j.jalz.2016.03.001 (2016).
Grabowska-Pyrzewicz, W., Want, A., Leszek, J. & Wojda, U. Antisense oligonucleotides for Alzheimer’s disease therapy: from the mRNA to miRNA paradigm. EBioMedicine 74, 103691. https://doi.org/10.1016/j.ebiom.2021.103691 (2021).
Toyama, K., Mogi, M. & Tsao, P. S. microRNA-based biomarker for dementia. Aging (Albany NY) 11, 1329–1330. https://doi.org/10.18632/aging.101868 (2019).
Lu, T. X. & Rothenberg, M. E. MicroRNA. J. Allergy Clin. Immunol. 141, 1202–1207. https://doi.org/10.1016/j.jaci.2017.08.034 (2018).
Siedlecki-Wullich, D., Minano-Molina, A. J. & Rodriguez-Alvarez, J. microRNAs as early biomarkers of Alzheimer’s disease: A synaptic perspective. Cells https://doi.org/10.3390/cells10010113 (2021).
Giorgi Silveira, R. et al. MicroRNAs expressed in neuronal differentiation and their associated pathways: Systematic review and bioinformatics analysis. Brain Res. Bull. 157, 140–148. https://doi.org/10.1016/j.brainresbull.2020.01.009 (2020).
Ma, F., Zhang, X. & Yin, K. J. MicroRNAs in central nervous system diseases: A prospective role in regulating blood-brain barrier integrity. Exp. Neurol. 323, 113094. https://doi.org/10.1016/j.expneurol.2019.113094 (2020).
Holohan, K. N., Lahiri, D. K., Schneider, B. P., Foroud, T. & Saykin, A. J. Functional microRNAs in Alzheimer’s disease and cancer: Differential regulation of common mechanisms and pathways. Front. Genet. 3, 323. https://doi.org/10.3389/fgene.2012.00323 (2012).
Swarbrick, S., Wragg, N., Ghosh, S. & Stolzing, A. Systematic review of miRNA as biomarkers in Alzheimer’s disease. Mol. Neurobiol. 56, 6156–6167. https://doi.org/10.1007/s12035-019-1500-y (2019).
Zhang, H., Liang, J. & Chen, N. The potential role of miRNA-regulated autophagy in Alzheimer’s disease. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23147789 (2022).
Gowda, P., Reddy, P. H. & Kumar, S. Deregulated mitochondrial microRNAs in Alzheimer’s disease: Focus on synapse and mitochondria. Ageing Res. Rev. 73, 101529. https://doi.org/10.1016/j.arr.2021.101529 (2022).
Kumar, S., Reddy, A. P., Yin, X. & Reddy, P. H. Novel MicroRNA-455-3p and its protective effects against abnormal APP processing and amyloid beta toxicity in Alzheimer’s disease. Biochim. Biophys. Acta Mol. Basis Dis. 186, 2428–2440. https://doi.org/10.1016/j.bbadis.2019.06.006 (1865).
Sarkar, S. et al. Expression of microRNA-34a in Alzheimer’s disease brain targets genes linked to synaptic plasticity, energy metabolism, and resting state network activity. Brain Res. 1646, 139–151. https://doi.org/10.1016/j.brainres.2016.05.026 (2016).
Wang, L. et al. Acupuncture attenuates inflammation in microglia of vascular dementia rats by inhibiting miR-93-mediated TLR4/MyD88/NF-kappaB signaling pathway. Oxid. Med. Cell. Longev. 2020, 8253904. https://doi.org/10.1155/2020/8253904 (2020).
Wu, H. Z. Y. et al. Differential blood miRNA expression in brain amyloid imaging-defined Alzheimer’s disease and controls. Alzheimers Res. Ther. 12, 59. https://doi.org/10.1186/s13195-020-00627-0 (2020).
Vergallo, A. et al. MiRNA-15b and miRNA-125b are associated with regional Abeta-PET and FDG-PET uptake in cognitively normal individuals with subjective memory complaints. Transl. Psychiatry 11, 78. https://doi.org/10.1038/s41398-020-01184-8 (2021).
Klyucherev, T. O. et al. Advances in the development of new biomarkers for Alzheimer’s disease. Transl. Neurodegener. 11, 25. https://doi.org/10.1186/s40035-022-00296-z (2022).
Liang, X. et al. Exosomal miR-532-5p induced by long-term exercise rescues blood-brain barrier function in 5XFAD mice via downregulation of EPHA4. Aging Cell. 22, e13748. https://doi.org/10.1111/acel.13748 (2023).
Xue, B., Qu, Y., Zhang, X. & Xu, X. F. miRNA-126a-3p participates in hippocampal memory via alzheimer’s disease-related proteins. Cereb. Cortex 32, 4763–4781. https://doi.org/10.1093/cercor/bhab515 (2022).
Weaver, N. A. et al. Strategic infarct locations for post-stroke cognitive impairment: A pooled analysis of individual patient data from 12 acute ischaemic stroke cohorts. Lancet Neurol. 20, 448–459. https://doi.org/10.1016/S1474-4422(21)00060-0 (2021).
Pendlebury, S. T., Rothwell, P. M. & Oxford Vascular Study. Incidence and prevalence of dementia associated with transient ischaemic attack and stroke: analysis of the population-based Oxford Vascular Study. Lancet Neurol 18, 248–258, https://doi.org/10.1016/S1474-4422(18)30442-3 (2019).
Barbay, M. et al. Prevalence of poststroke neurocognitive disorders using National Institute of Neurological Disorders and Stroke-Canadian Stroke Network, VASCOG criteria (vascular behavioral and cognitive disorders), and optimized criteria of cognitive deficit. Stroke 49, 1141–1147. https://doi.org/10.1161/STROKEAHA.117.018889 (2018).
Huang, Y. Y. et al. Post-stroke cognitive impairment: Epidemiology, risk factors, and management. J. Alzheimers Dis. 86, 983–999. https://doi.org/10.3233/JAD-215644 (2022).
Sexton, E. et al. Systematic review and meta-analysis of the prevalence of cognitive impairment no dementia in the first year post-stroke. Eur. Stroke J. 4, 160–171. https://doi.org/10.1177/2396987318825484 (2019).
Craig, L., Hoo, Z. L., Yan, T. Z., Wardlaw, J. & Quinn, T. J. Prevalence of dementia in ischaemic or mixed stroke populations: Systematic review and meta-analysis. J. Neurol. Neurosurg. Psychiatry 93, 180–187. https://doi.org/10.1136/jnnp-2020-325796 (2022).
Balasubramanian, P. et al. Time-restricted feeding (TRF) for prevention of age-related vascular cognitive impairment and dementia. Ageing Res. Rev. 64, 101189. https://doi.org/10.1016/j.arr.2020.101189 (2020).
Horikawa, I. Balancing and differentiating p53 activities toward longevity and no cancer?. Cancer Res. 80, 5164–5165. https://doi.org/10.1158/0008-5472.CAN-20-3080 (2020).
Kalaria, R. N. The pathology and pathophysiology of vascular dementia. Neuropharmacology 134, 226–239. https://doi.org/10.1016/j.neuropharm.2017.12.030 (2018).
Ghoge, H., Sharma, S., Sonawalla, S. & Parikh, R. Cerebrovascular diseases and depression. Curr. Psychiatry Rep. 5, 231–238. https://doi.org/10.1007/s11920-003-0048-7 (2003).
Lee, R., Choi, H., Park, K. Y., Kim, J. M. & Seok, J. W. Prediction of post-stroke cognitive impairment using brain FDG PET: Deep learning-based approach. Eur. J. Nucl. Med. Mol. Imaging 49, 1254–1262. https://doi.org/10.1007/s00259-021-05556-0 (2022).
Pendlebury, S. T. & Rothwell, P. M. Prevalence, incidence, and factors associated with pre-stroke and post-stroke dementia: A systematic review and meta-analysis. Lancet Neurol. 8, 1006–1018. https://doi.org/10.1016/S1474-4422(09)70236-4 (2009).
Mijajlovic, M. D. et al. Post-stroke dementia—A comprehensive review. BMC Med. 15, 11. https://doi.org/10.1186/s12916-017-0779-7 (2017).
Ragusa, M. et al. miRNAs plasma profiles in vascular dementia: biomolecular data and biomedical implications. Front. Cell. Neurosci. 10, 51. https://doi.org/10.3389/fncel.2016.00051 (2016).
Sampath, D. et al. Sex differences in cognitive impairment after focal ischemia in middle-aged rats and the effect of iv miR-20a-3p treatment. Neurobiol. Aging 129, 168–177. https://doi.org/10.1016/j.neurobiolaging.2023.05.001 (2023).
Ren, Z. et al. MicroRNA-210-5p contributes to cognitive impairment in early vascular dementia rat model through targeting Snap25. Front. Mol. Neurosci. 11, 388. https://doi.org/10.3389/fnmol.2018.00388 (2018).
Sonoda, T. et al. Serum microRNA-based risk prediction for stroke. Stroke 50, 1510–1518. https://doi.org/10.1161/STROKEAHA.118.023648 (2019).
Toyama, K. et al. MicroRNA-mediated therapy modulating blood-brain barrier disruption improves vascular cognitive impairment. Arterioscler. Thromb. Vasc. Biol. 38, 1392–1406. https://doi.org/10.1161/ATVBAHA.118.310822 (2018).
Li, Z. & Rana, T. M. Therapeutic targeting of microRNAs: Current status and future challenges. Nat. Rev. Drug Discov. 13, 622–638. https://doi.org/10.1038/nrd4359 (2014).
Cheng, C. J. & Slack, F. J. The duality of oncomiR addiction in the maintenance and treatment of cancer. Cancer J. 18, 232–237. https://doi.org/10.1097/PPO.0b013e318258b75b (2012).
Rupaimoole, R., Han, H. D., Lopez-Berestein, G. & Sood, A. K. MicroRNA therapeutics: principles, expectations, and challenges. Chin. J. Cancer 30, 368–370. https://doi.org/10.5732/cjc.011.10186 (2011).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Asanomi, Y. et al. Dementia subtype prediction models constructed by penalized regression methods for multiclass classification using serum microRNA expression data. Sci. Rep. 11, 20947. https://doi.org/10.1038/s41598-021-00424-1 (2021).
Bagasra, O., Shamabadi, N. S., Pandey, P., Desoky, A. & McLean, E. Differential expression of miRNAs in a human developing neuronal cell line chronically infected with Zika virus. Libyan J. Med. 16, 1909902. https://doi.org/10.1080/19932820.2021.1909902 (2021).
Zhou, X. et al. Therapeutic effects of (5R)-5-hydroxytriptolide on fibroblast-like synoviocytes in rheumatoid arthritis via lncRNA WAKMAR2/miR-4478/E2F1/p53 axis. Front. Immunol. 12, 605616. https://doi.org/10.3389/fimmu.2021.605616 (2021).
Ghanbari, R. et al. Decreased expression of fecal miR-4478 and miR-1295b-3p in early-stage colorectal cancer. Cancer Biomark. 15, 189–195. https://doi.org/10.3233/CBM-140453 (2015).
Meng, Q., Liu, M. & Cheng, R. LINC00461/miR-4478/E2F1 feedback loop promotes non-small cell lung cancer cell proliferation and migration. Biosci. Rep. https://doi.org/10.1042/BSR20191345 (2020).
Quinn, P. M. J., Moreira, P. I., Ambrosio, A. F. & Alves, C. H. PINK1/PARKIN signalling in neurodegeneration and neuroinflammation. Acta Neuropathol. Commun. 8, 189. https://doi.org/10.1186/s40478-020-01062-w (2020).
Yue, N., Ye, M., Zhang, R. & Guo, Y. MiR-449b-5p targets lncRNA PSMG3-AS1 to suppress cancer cell proliferation in lung adenocarcinoma. BMC Pulm. Med. 20, 152. https://doi.org/10.1186/s12890-020-01189-5 (2020).
Varela, L. & Garcia-Rendueles, M. E. R. Oncogenic pathways in neurodegenerative diseases. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23063223 (2022).
Muralidar, S., Ambi, S. V., Sekaran, S., Thirumalai, D. & Palaniappan, B. Role of tau protein in Alzheimer’s disease: The prime pathological player. Int. J. Biol. Macromol. 163, 1599–1617. https://doi.org/10.1016/j.ijbiomac.2020.07.327 (2020).
Jazvinscak Jembrek, M., Slade, N., Hof, P. R. & Simic, G. The interactions of p53 with tau and Ass as potential therapeutic targets for Alzheimer’s disease. Prog. Neurobiol. 168, 104–127. https://doi.org/10.1016/j.pneurobio.2018.05.001 (2018).
Sabapathy, K. & Lane, D. P. Understanding p53 functions through p53 antibodies. J. Mol. Cell Biol. 11, 317–329. https://doi.org/10.1093/jmcb/mjz010 (2019).
Sargolzaei, J., Etemadi, T. & Alyasin, A. The P53/microRNA network: A potential tumor suppressor with a role in anticancer therapy. Pharmacol. Res. 160, 105179. https://doi.org/10.1016/j.phrs.2020.105179 (2020).
Goeman, F., Strano, S. & Blandino, G. MicroRNAs as key effectors in the p53 network. Int. Rev. Cell Mol. Biol. 333, 51–90. https://doi.org/10.1016/bs.ircmb.2017.04.003 (2017).
Liu, R. et al. PI3K/AKT pathway as a key link modulates the multidrug resistance of cancers. Cell Death Dis. 11, 797. https://doi.org/10.1038/s41419-020-02998-6 (2020).
Tewari, D., Patni, P., Bishayee, A., Sah, A. N. & Bishayee, A. Natural products targeting the PI3K-Akt-mTOR signaling pathway in cancer: A novel therapeutic strategy. Semin. Cancer Biol. 80, 1–17. https://doi.org/10.1016/j.semcancer.2019.12.008 (2022).
Wei, H. et al. Mesenchymal stem cell-derived exosomal miR-223 regulates neuronal cell apoptosis. Cell Death Dis. 11, 290. https://doi.org/10.1038/s41419-020-2490-4 (2020).
Kim, E. K. & Choi, E. J. Compromised MAPK signaling in human diseases: an update. Arch Toxicol 89, 867–882. https://doi.org/10.1007/s00204-015-1472-2 (2015).
Xu, H. et al. p38 MAPK-mediated loss of nuclear RNase III enzyme Drosha underlies amyloid beta-induced neuronal stress in Alzheimer’s disease. Aging Cell 20, e13434. https://doi.org/10.1111/acel.13434 (2021).
Sedzikowska, A. & Szablewski, L. Insulin and insulin resistance in Alzheimer’s disease. Int. J. Mol. Sci. https://doi.org/10.3390/ijms22189987 (2021).
Benetatos, J. et al. PTEN activation contributes to neuronal and synaptic engulfment by microglia in tauopathy. Acta Neuropathol. 140, 7–24. https://doi.org/10.1007/s00401-020-02151-9 (2020).
Roca-Agujetas, V., de Dios, C., Abadin, X. & Colell, A. Upregulation of brain cholesterol levels inhibits mitophagy in Alzheimer disease. Autophagy 17, 1555–1557. https://doi.org/10.1080/15548627.2021.1920814 (2021).
Knafo, S. et al. PTEN recruitment controls synaptic and cognitive function in Alzheimer’s models. Nat. Neurosci. 19, 443–453. https://doi.org/10.1038/nn.4225 (2016).
Yuan, Y. et al. PTENalpha regulates endocytosis and modulates olfactory function. FASEB J. 33, 11148–11162. https://doi.org/10.1096/fj.201900588RR (2019).
Wang, X. J. et al. PINK1 overexpression prevents forskolin-induced tau hyperphosphorylation and oxidative stress in a rat model of Alzheimer’s disease. Acta Pharmacol. Sin. 43, 1916–1927. https://doi.org/10.1038/s41401-021-00810-5 (2022).
Lalatsa, A., Sun, Y., Gamboa, J. I. & Knafo, S. Preformulation studies of a stable PTEN-PDZ lipopeptide able to cross an in vitro blood-brain-barrier model as a potential therapy for Alzheimer’s disease. Pharm Res 37, 183. https://doi.org/10.1007/s11095-020-02915-8 (2020).
Sacco, A. et al. Regulatory miRNAs in cardiovascular and Alzheimer’s disease: A focus on copper. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23063327 (2022).
Alexander, C. et al. Hypoxia Inducible Factor-1alpha binds and activates gamma-secretase for Abeta production under hypoxia and cerebral hypoperfusion. Mol. Psychiatry 27, 4264–4273. https://doi.org/10.1038/s41380-022-01676-7 (2022).
Grubman, A. et al. Transcriptional signature in microglia associated with Abeta plaque phagocytosis. Nat. Commun. 12, 3015. https://doi.org/10.1038/s41467-021-23111-1 (2021).
Ashok, B. S., Ajith, T. A. & Sivanesan, S. Hypoxia-inducible factors as neuroprotective agent in Alzheimer’s disease. Clin. Exp. Pharmacol. Physiol. 44, 327–334. https://doi.org/10.1111/1440-1681.12717 (2017).
Lei, L. et al. HIF-1alpha causes LCMT1/PP2A deficiency and mediates tau hyperphosphorylation and cognitive dysfunction during chronic hypoxia. Int. J. Mol. Sci. https://doi.org/10.3390/ijms232416140 (2022).
Yu, W. et al. Connexin43 promotes angiogenesis through activating the HIF-1alpha/VEGF signaling pathway under chronic cerebral hypoperfusion. J. Cereb. Blood Flow Metab. 41, 2656–2675. https://doi.org/10.1177/0271678X211010354 (2021).
El Hajjar, J. et al. Heterochromatic genome instability and neurodegeneration sharing similarities with Alzheimer’s disease in old Bmi1+/− mice. Sci. Rep. 9, 594. https://doi.org/10.1038/s41598-018-37444-3 (2019).
Hogan, R., Flamier, A., Nardini, E. & Bernier, G. The role of BMI1 in late-onset sporadic Alzheimer’s disease. Genes (Basel) https://doi.org/10.3390/genes11070825 (2020).
Flamier, A. et al. Modeling late-onset sporadic Alzheimer’s disease through BMI1 deficiency. Cell Rep. 23, 2653–2666. https://doi.org/10.1016/j.celrep.2018.04.097 (2018).
Rupaimoole, R. et al. Author correction: Hypoxia-mediated downregulation of miRNA biogenesis promotes tumour progression. Nat. Commun. 11, 2867. https://doi.org/10.1038/s41467-020-16661-3 (2020).
Rupaimoole, R. et al. Hypoxia-upregulated microRNA-630 targets Dicer, leading to increased tumor progression. Oncogene 35, 4312–4320. https://doi.org/10.1038/onc.2015.492 (2016).
van den Beucken, T. et al. Hypoxia promotes stem cell phenotypes and poor prognosis through epigenetic regulation of DICER. Nat. Commun. 5, 5203. https://doi.org/10.1038/ncomms6203 (2014).
Rupaimoole, R. & Slack, F. J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 16, 203–222. https://doi.org/10.1038/nrd.2016.246 (2017).
Jaber, V. R., Zhao, Y., Sharfman, N. M., Li, W. & Lukiw, W. J. Addressing Alzheimer’s disease (AD) neuropathology using anti-microRNA (AM) strategies. Mol. Neurobiol. 56, 8101–8108. https://doi.org/10.1007/s12035-019-1632-0 (2019).
Liu, J. H. et al. Elevated blood neutrophil to lymphocyte ratio in older adults with cognitive impairment. Arch. Gerontol. Geriatr. 88, 104041. https://doi.org/10.1016/j.archger.2020.104041 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Friedlander, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 Accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52. https://doi.org/10.1093/nar/gkr688 (2012).
Hauschild, A. C. et al. MirDIP 5.2: Tissue context annotation and novel microRNA curation. Nucleic Acids Res. 51, D217–D225. https://doi.org/10.1093/nar/gkac1070 (2023).
Chang, L., Zhou, G., Soufan, O. & Xia, J. miRNet 2.0: Network-based visual analytics for miRNA functional analysis and systems biology. Nucleic Acids Res. 48, W244–W251. https://doi.org/10.1093/nar/gkaa467 (2020).
McGeary, S. E. et al. The biochemical basis of microRNA targeting efficacy. Science https://doi.org/10.1126/science.aav1741 (2019).
The Gene Ontology, C. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338. https://doi.org/10.1093/nar/gky1055 (2019).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361. https://doi.org/10.1093/nar/gkw1092 (2017).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Chen, T. et al. The genome sequence archive family: Toward explosive data growth and diverse data types. Genomics Proteomics Bioinform. 19, 578–583. https://doi.org/10.1016/j.gpb.2021.08.001 (2021).
Members, C.-N. & Partners. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2023. Nucleic Acids Res 51, D18-D28, https://doi.org/10.1093/nar/gkac1073 (2023).
Acknowledgements
We thank the Department of Biobank, Division of Clinical Research for providing human tissues, RNA Sequencing service were provided by Aksomics, and we would like to thank Editage (www.editage.com) for English language editing.
Funding
The funding was provided by the General Program of the National Natural Science Foundation of China (Grant No. 82071442); the Jilin Provincial Department of Finance (Grant No. JLSWSRCZX2021-004); the Major Chronic Disease Program of the Ministry of Science and Technology of China (Grant No. 2018YFC1312301); and STI2030-Major Projects (No. 2021ZD0201802).
Author information
Authors and Affiliations
Contributions
L.S. designed and supervised the study. W.Z. drafted the manuscript. C.W., G.Z., A.Z. and Y.Q. reviewed the literature. M.Z., C.W., G.Z. and L.S. revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhai, W., Zhao, M., Wei, C. et al. Biomarker profiling to determine clinical impact of microRNAs in cognitive disorders. Sci Rep 14, 8270 (2024). https://doi.org/10.1038/s41598-024-58882-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58882-2
- Springer Nature Limited