Abstract
Parkinson's disease (PD) is a progressive neurodegenerative disease whose etiology is attributed to development of Lewy bodies and degeneration of dopaminergic neurons in the substantia nigra (SN). Currently, there are no definitive diagnostic indicators for PD. In this study, we aimed to identify potential diagnostic biomarkers for PD and analyzed the impact of immune cell infiltrations on disease pathogenesis. The PD expression profile data for human SN tissue, GSE7621, GSE20141, GSE20159, GSE20163 and GSE20164 were downloaded from the Gene Expression Omnibus (GEO) database for use in the training model. After normalization and merging, we identified differentially expressed genes (DEGs) using the Robust rank aggregation (RRA) analysis. Simultaneously, DEGs after batch correction were identified. Gene interactions were determined through venn Diagram analysis. Functional analyses and protein–protein interaction (PPI) networks were used to the identify hub genes, which were visualized through Cytoscape. A Lasso Cox regression model was employed to identify the potential diagnostic genes. The GSE20292 dataset was used for validation. The proportion of infiltrating immune cells in the samples were determined via the CIBERSORT method. Sixty-two DEGs were screened in this study. They were found to be enriched in nerve conduction, dopamine (DA) metabolism, and DA biosynthesis Gene Ontology (GO) terms. The PPI network and Lasso Cox regression analysis revealed seven potential diagnostic genes, namely SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1, were subsequently validated in peripheral blood samples obtained from healthy control (HC) and PD patients, as well as in the GSE20292 dataset. The results revealed the exceptional sensitivity and specificity of these genes in PD diagnosis and monitoring. Moreover, PD patients exhibited a higher number of plasma cells, compared to HC individuals. The SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1 are potential diagnostic biomarkers for PD. Our findings also reveal the essential roles of immune cell infiltration in both disease onset and trajectory.
Similar content being viewed by others
Introduction
Over the last 25 years, there has been a global increase in the number of individuals affected by, dying from, or suffering from long-term neurological disorder-associated disabilities. This is despite the development of multiple novel diagnostic and treatment methods1. Parkinson’s disease (PD) is a significant contributor to neurological disability, and is the second most prevalent neurodegenerative disorder worldwide. With a global prevalence exceeding 6 million individuals, the condition has exhibited a remarkable 2.5-fold rise in occurrence over the past generation2. In industrialized countries, the estimated prevalence of this condition is 0.3%, which is rarely observed in patients under the age of 40, but its incidence is increasing with advancing age3. The disease is associated with typical motor symptoms, such as parkinsonism, Lewy bodies and dopaminergic neuron degeneration in the substantia nigra (SN)4,5,6. The underlying etiologic factors for this complex disease are attributed to a combination of genetic and environmental factors that affect essential cellular processes7,8. Clinical management of this disease is challenged by limitations of treatment and definitive diagnosis, particularly at the earliest stages of the disease.
Through transcriptomic analysis, studies have identified various differentially expressed genes (DEGs) and dysregulated pathways. Beta-glucocerebrosidase is involved in both the endo-lysosomal pathway and immune responses, which are two critical processes in PD development9. Kurvits et al. used the Robust rank aggregation (RRA) strategy and found that IL18R1, an interleukin receptor associated with proinflammatory responses, was induced by GM-CSF administration and was associated with neuroprotective mechanisms in PD 10,11. Therefore, PD is a potential multisystem disorder. Songyun Zhao et al12. Performed Lasso Cox regression analysis and constructed a model that yielded the most substantial net gain, underscoring the critical role of the advanced risk model in guiding personalized anti-cancer therapy and driving informed decision-making, which is related to PD. Protein–protein interaction (PPI) network analysis is a powerful approach for achieving a comprehensive understanding of biological processes at molecular and systemic levels13. For downstream applications, the STRING database integrates data from various primary databases14. Recently, Kim et al. reported on the functional significance of thiol-oxidoreductase TXNIP in development of LRRK2-associated PD within a three dimensional (3D) environment, highlighting the potential of 3D organoid-based models in advancing therapeutic discovery for sporadic PD15. The significance of immune cell infiltrations in PD onset and progression has been well-established16,17. CIBERSORT, a computational tool, allows for assessment of immune cell proportions based on gene expression data18. Both innate and adaptive immune systems play a role in neuronal death and PD pathogenesis19. Despite remarkable advances in recent years, the etiopathogenesis of PD, encompassing the contribution of biomarkers and underlying biological processes leading to formation of Lewy bodies and dopaminergic neuron loss in the SN, has not been fully elucidated. In this study, we aimed at identifying the potential biomarkers and molecular mechanisms that can promote healthy brain functions and avert PD onset.
We downloaded five datasets from the Gene Expression Omnibus (GEO), combined them as the training set and used another dataset as the validation set. Then, RRA and batch correction were used in the training set to establish the consensus DEGs of PD. Next, Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed to reveal the biological functions of the DEGs. The PPI networks were used to establish interactions among DEGs-related proteins. Lasso Cox regression models were used to identify the possible biomarkers for PD. Finally, 7 genes, namely SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1, were identified to be potential diagnostic genes for PD. To validate the clinical accuracy of the genes, a validation set and reverse transcription-quantitative real-time polymerase chain reaction (RT-qPCR) were performed. CIBERSORT analysis was performed to investigate immune cell infiltrations in PD samples. The goal of this study was to identify the potential diagnostic genes for PD and to characterize immune cell infiltrations in peripheral blood samples from PD patients.
Materials and methods
Data source and pre-processing.
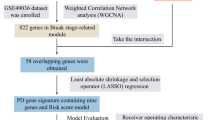
Five PD datasets (GSE7621, GSE20141, GSE20159, GSE20163 and GSE20164) were downloaded from the GEO database (https://www.ncbi.nlm.nih.gov/geo/), merged and used as the training set, which contains 48 healthy control (HC) and 56 PD patients. The GSE20292 dataset, which consists of transcriptional analysis data for the whole SN in PD, was used for validation (Table 1). The ‘sva’ package (https://bioconductor.org/packages/release/bioc/html/sva.html) was used to remove batch effects from the training sets. The principal component analysis (PCA) cluster plot was used to visualize the effects of removing inter-batch differences. The workflow of this study is shown in Fig. 1.
RRA analysis.
The RRA analysis was employed to systematically integrate gene expression information across diverse datasets20. We integrated the R packages ‘limma’ (https://bioconductor.org/packages/release/bioc/html/limma.html) and ‘RobustRankAggreg’ (https://cran.r-project.org/web/packages/RobustRankAggreg/index.html) to identify genes that exhibited statistically significant changes (adjusted P value < 0.05) and those that were statistically significance in adjusted P value < 0.05 and |log fold change (FC) |> 1.0.
Functional enrichment analysis.
The GO and KEGG enrichment analyses of DEGs were conducted using the ‘clusterProfiler’ (https://bioconductor.org/packages/release/bioc/html/clusterProfiler.html) package of R software. Findings of GO annotation analysis were categorized as follows: Biological Process (BP), Molecular Function (MF), and Cellular Components (CC)21,22. Significant enrichment was defined as adjusted P values < 0.05.
Establishment of the PPI network and identification of hub genes.
To establish the interplay of DEGs, PPI network analysis was performed using the STRING database (https://cn.string-db.org/). A stringent interaction score threshold of 0.4 was employed to identify the most reliable and relevant interactions among the DEGs. Genes were denoted by nodes while connections between them were represented by edges. Subsequently, the main regulatory network was constructed and visualized using Cytoscape (https://cytoscape.org/, version 3.9.1). The cytoHubba plugin of Cytoscape was used to identify the hub genes in the PPI network. A systematic evaluation of central genes was performed using a comprehensive repertoire of ten distinct methods: MCC, DMNC, Degree, BottleNeck, EcCentricity, Closeness, MNC, Radiality, Stress, andBetweenness. By synthesizing the results obtained from these diverse approaches, the top 39 hub genes were identified, representing a collective selection based on their high-ranking positions across the ten methods.
Identification of PD biomarkers via Lasso Cox regression analysis.
The Lasso Cox regression analysis, a penalized regression technique, is effective for predicting outcomes and has low correlations, making it suitable for selecting the most relevant features in datasets with various variables. The ‘glmnet’ package(https://cran.r-project.org/web/packages/glmnet/) was used to extract and fit the consensus DEGs expression profiles into the Lasso Cox regression model. To identify the DEGs between HC and PD patients after Lasso Cox regression analysis, a heatmap and volcano map were drawn in the training set.
Evaluation of the diagnostic model in verification set.
A heatmap and a violin map were drawn to identify the differentially expressed hub genes between HC and PD patients in the verification set. Then, the efficacy of distinguishing between HC and PD patients was tested by receiver operating characteristic (ROC) curve analysis.
Immune cell infiltration analysis.
The R package ‘CIBERSORT’ (https://github.com/Moonerss/CIBERSORT) was used to calculate the fractions of immune infiltrating cells in the training set, which were visualized using the boxplot. The Violin Plot was used to identify statistically significant differences in immune cells between HC and PD patients (P value < 0.05).
RT-qPCR
Independent peripheral blood samples from PD patients were used to assess the practical efficacy of our diagnostic model. Clinical whole blood samples were collected from both HC and PD patients, and total DNA were extracted using the FastPure Blood DNA Isolation Mini Kit V2. The β-Actin primer pair was used as the internal control. Relative gene expressions were calculated and normalized via the Ct technique and ΔΔCt method. Clinical information for PD patients used in the RT-qPCR is detailed in Supplementary Table S1. Primer sequences are detailed in Table 2.
Statistical analysis
Data preparation, functional analysis, and modeling were performed using R software (https://cran.r-project.org/, v4.1.3). All P values were two-sided and differences with P < 0.05 and log |FC|< 0.585 were considered significant. Cytoscape was utilized to visualize the PPI network. ROC curves analysis was derived using ‘pROC’ packages (https://cran.r-project.org/web/packages/pROC/). Intergroup comparisons were performed via the Wilcox test.
Ethical approval
In this study, we confirm that all experiments and methods were conducted strictly in accordance with relevant guidelines and regulations. The experimental protocols have been approved by the ethics committee to ensure the legality and ethical compliance of the research. Hereby declare that informed consent was obtained from all human participants, including the use of tissue samples, involved in our research study. The human tissue sample collection for this study have obtained approval from the following institution: The first affiliated hospital of guangxi medical university. The number of the approving: 2023-E258-01.
Results
Identification of DEGs.
Five datasets (GSE7621, GSE20141, GSE20159, GSE20163 and GSE20164) with a total of 104 samples were included as the training set. Batch correction based on the ‘sva’ R package was performed to diminish the potential for bias to be introduced by batch effects on subsequent analyses. Boxplot and PCA revealed the differences before and after batch correction, indicating the successful removal of batch effects (Fig. 2A–D). The R package ‘RobustRankAggreg’ was used to perform the RRA analysis (Fig. 3A). Batch correction was performed on the training set (Fig. 3B). A total of 80 DEGs (nine upregulated and 71 downregulated) between HC and PD patients were identified via RRA and batch correction. Based on the R package of ‘venn’ (https://cran.r-project.org/web/packages/venn/), There are 62 DEGs were detected between RRA analysis and batch correction. (Fig. 3C). In conclusion, by integrating the datasets and treating them as the training set, we were able to filter out 62 DEGs using RRA and batch correction, thereby laying the foundation for subsequent analyses.
GO and KEGG pathway functional enrichment analysis of DEGs.
The GO and KEGG enrichment analyses were performed to determine the functions of the 62 significant DEGs (Fig. 4A,B). The enriched GO terms were nerve conduction, dopamine (DA) metabolism, and DA biosynthesis. The enriched KEGG terms included PD and the cycle of nerve conduction. Association of proteins and the top five GO and KEGG terms were visualized using a cnetplot (Fig. 4C,D). Based on our analysis, it is possible that the identified DEGs are closely associated with the nervous system and may potentially impact PD pathogenesis.
The GO analysis and KEGG pathway analysis of DEGs. (A) Dot plot showing GO analysis of DEGs. (B) Dot plot showing KEGG pathway analysis of DEGs. (C) Circle graph showing the DEGs that were enriched in the top 5 GO categories of BPs. (D) Circle graph showing the DEGs that were enriched in KEGG pathway analysis.
PPI network analysis and hub genes selection
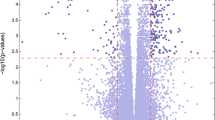
The PPI networks were analyzed in the STRING online database to elucidate on protein interactions among the DEGs (Fig. 5A). Subsequently, the main regulatory network was constructed and visualized using Cytoscape Consequently, a total of 39 genes were identified (Fig. 5B). Ranked and networks of the genes for each approach were listed in Supplementary Table S1 and Supplementary Figure S1. Expression profiles of the hub genes were extracted and integrated into the Lasso Cox regression model to identify the possible biomarkers for PD. To build a model that facilitates quantification of each patient with accuracy, seven of the 39 DEGs, including SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1 were retained by application of the Lasso Cox regression model with a minimum of λ (Fig. 5C). The distribution of these genes is presented in the volcano plot (Fig. 5D). Expression levels of the above genes significantly differed between the two sets as shown in the heatmap (Fig. 5E). These findings imply that the hub genes may have pivotal functions in biological mechanisms underlying PD and are promising therapeutic targets.
PPI network and hub gene selection. (A) The top 62 hub genes in the PPI network of DEGs based on node degree. (B) Hub genes were identified by taking the interplay of the first 39 genes in the ten classification methods of cytoHubba. (C) Hub gene selection in the Lasso Cox regression model. Vertical lines were drawn at optimal values by the minimum criteria and the 1–SE criteria. (D) The hub gene landscape in the volcano map.
Immune cell infiltration analysis.
Progression of neurodegenerative processes in PD may be sustained by changes in immune cell markers that induce or worsen neuroinflammation6,16. Therefore, to establish the constitution of immune cells in PD samples for the purpose of obtaining the difference between HC and PD patients in relative abundance of immune cells, the immune status was evaluated. Constituent ratios for 22 infiltrating immune cell types were as shown in Fig. 6A. The PD patients exhibited higher counts of plasma cells compared with HC (Fig. 6B). The intensity of plasma cell expressions, as evidenced by the image, was closely associated with PD.
Performance of the diagnostic model in SN datasets and samples.
To assess the clinical efficacy of our diagnostic model, we assessed it using SN datasets (GSE20292). Then, we developed a heatmap and a violin plot between HC and PD patients for the seven hub genes (Fig. 7A,B). Hub gene expression levels in the dataset were calculated using the ROC curves to identify the corresponding area under the curve (AUC). In Fig. 7C, AUCs for SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1 in HC and PD patients were 0.864, 0.652, 0.863, 0.934, 0.65, 0.722, and 0.773, respectively. When the seven hub genes were combined into a single variable, their predictive value improved; the AUC for the model was 0.965 (Fig. 7D). Thus, the hub genes have a high predictive accuracy for diagnosis.
Validation of expression level of the seven hub genes in peripheral blood samples.
The RT-qPCR assay was performed to validate the efficacy of the prediction model. Relative expressions of TAC1, PDE6H, KIAA0319, AGTR1, SLC18A2, and PCDH8 in HC were significantly higher than those in PD patients, while those of AXIN1 were significantly lower in HC, relative to PD patients (Fig. 8). These results from peripheral blood samples are consistent with the GSE20292 dataset performed on SN, indicating that these genes hold promise as prospective therapeutic targets for PD patients.
Discussion
The gold standard for diagnosing PD relies on the presence of SN pars compacta degeneration and Lewy pathology, as confirmed by post-mortem pathological examination23,24. Pathologically, PD is a slowly progressing neurological disease that begins years before a diagnosis25. Diagnostic examinations that allow for clear diagnosis during the initial phases of the disease, in particular, do not exist. Therefore, there is a need to develop suitable diagnostic approaches for early diagnosis. Through bioinformatics analyses, we aimed at identifying diagnostic genes that are related to the disease.
In previous studies, we acknowledge that some datasets were utilized in prior research by Zheng et al.26 and Kelly et al27. Zheng ’s study conducted a genome-wide meta-analysis of gene sets in PD, identifying 10 gene sets associated with the disease. Key findings include defects in mitochondrial electron transport, glucose utilization, and glucose sensing occurring early in disease pathogenesis. In Kelly’s study, DEGs were found to be associated with perturbed pathways, including mitochondrial dysfunction and oxidative stress. In our study, based on GSE7621, GSE20141, GSE20159, GSE20163, and GSE20164, RRA and batch correction were performed to identify the consensus DEGs, which were 80 in number. Among them, 71 genes were downregulated while nine genes were upregulated. Functional enrichment analysis revealed that the enriched GO terms were mainly in nerve conduction, DA metabolism, and DA biosynthesis. Pathologically, PD is associated with both neuronal dysfunction and inflammation of the central nervous system, consistent with nerve conduction disorders28. Dysregulated DA is more likely to play a crucial role in early onset of PD, thus, early identification of dysregulated DA should be a priority29. The enriched KEGG terms were also mainly in nerve conduction. Analysis of the PPI network revealed 39 hub genes, which were visualized by Cytoscape. The Lasso Cox regression model was used to assess the diagnostic genes with a high accuracy, which were SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1. CIBERSORT was used to assess immune cell infiltrations in PD. Plasma cells were found to be differentially expressed between HC and PD patients. Compared with previous studies, our study integrates multiple datasets and employs advanced methods such as RRA analysis and immune cell infiltration analysis, we believe that our work contributes to a more comprehensive understanding of the molecular mechanisms underlying PD.
SLC18A2, the vesicular monoamine transporter 2, is important in neurotransmitter transportation. It packages histamine into vesicles in preparation for neurotransmitter release from the presynaptic neuron30. The gene, which is important in the monoaminergic signaling pathway, has been extensively researched on. In the absence of this monoaminergic transporter, histamine immunoreactivity is significantly suppressed in neuronal cell bodies of the brain31. Reduced histamine metabolism in the central nervous system is a preventative measure against PD onset32. Further elucidation of the involved mechanisms will contribute to a better understanding of the disease. Substance P (SP) dysregulation is associated with the etiology of PD. Mice lacking endogenous SP (TAC1-/-) exhibited greater resistance to nigral dopaminergic neurodegeneration than wild-type controls. The neuroinflammatory and dopaminergic neurodegenerative effects of SP are mediated by microglial NOX233, thus, they may shed new light on PD pathophysiology. Protocadherins, which contain PCDH8, are calcium-dependent adhesion molecules that have drawn interest for their potential functions in development of neural circuits and their potential effects on neurological illnesses. Physiologically, PCDH8 is involved in development and maintenance of intrahippocampal circuits34. However, the association between PCDH8 and PD has yet to be established. KIAA0319 is associated with extracellular signaling pathways35,36. Extracellular signaling pathways and endocytosis are both necessary to control neurogenesis37. As a pivotal upstream gatekeeper, KIAA0319 plays crucial roles in neurogenesis by arresting cellular progression at the neural progenitor cell stage. Cell cycle progression is deregulated in PD, and key regulators of the G1/S transition checkpoint are significantly altered38. Moreover, the cell cycle is enriched in GO terms. Even though there is no conclusive proof that KIAA0319 is directly associated with PD, our findings shed light on the subject. PDE6H is associated with changes in circadian rhythms that are involved in aging39, however, the association between PDE6H and PD has not been determined. AXIN1 was overexpressed in hippocampus tissues and cells from MPTP-lesioned mice models of PD. AXIN1 suppression in PD suppressed hippocampus neuron apoptosis. AXIN1 downregulation suppresses DA neuron death via miR-12840. These outcomes suggest a therapeutic potential for AXIN1 in treatment of PD. AGTR1 encodes the angiotensin II type 1 receptor (AT1R), whose expressions decreases in dopaminergic neurons of the SN as PD advances41. In contrast, AT1R upregulation induces the release of pro-inflammatory cytokines, leading to inflammation that culminates in dopaminergic cell death and dysfunction. Blocking AT1R with its antagonist can attenuate neurotoxin-induced degeneration of dopaminergic neurons in the SN42,43. These findings underscore the complex nature of AGTR1’s role in PD and highlight the need for further research.
The GO function analysis revealed that the DEGs were primarily enriched in nerve conduction, DA metabolism, and DA biosynthesis. Nerve conduction is divided into sensory nerve conduction and motor nerve conduction44. Regarding nerve conduction, Toth reported that individuals with PD exhibited slower motor conduction velocities, when compared to HC. Toth hypothesized that peripheral neuropathy in PD could be attributed to levodopa exposure and elevated levels of methylmalonic acid45. Thus, motor nerve transmission abnormalities maybe present in PD patients46. Pathologically, PD is characterized by degeneration of the nigrostriatal dopaminergic system. In 1988, Gotham et al. proposed the ‘dopamine overdose’ hypothesis47. The hypothesis, which proposes an impact on cognitive functions in PD, suggests that the medication doses required to restore DA functions in the most severely affected regions may be too high for less affected areas. According to this theory, the ventral striatum, which remains relatively intact, can become excessively stimulated when DA replacement therapy is administered. Subsequently, this overstimulation affects the limbic system, leading to impaired executive functions mediated by the limbic and orbitofrontal systems, such as learning and risk-taking. Empirical evidence from literature and clinical observations, particularly in relation to DA agonists, provides support for this theory and the emergence of impulse control disorders48. The enriched KEGG terms also included the PD and cycle of nerve conduction. The diagnostic gene, SLC18A2, is also associated with neurotransmitter transport, synaptic vesicle cycle, and dopaminergic synapses. In terms of neurotransmitter transport, mutations in SLC18A2 will impact the transmission of monoamine neurotransmitters, leading to a phenotype that shares characteristics with all monoamine-related disorders49. In terms of the relationship between SLC18A2 and DA, SLC18A2 is a vesicular monoamine transporter that is essential in DA regulation. Suppressed SLC18A2 activities might reduce DA release50.
Studies have reported on significantly changed B cell subpopulation structures in PD51. Given that plasma cells are derived from B cells, the abundance of plasma cells is associated with PD development. Plasma cells may affect PD pathogenesis by influencing the immune microenvironment. However, the relationship between plasma cells and PD has not been fully defined. Systematic studies should be performed to elucidate on possible mechanisms of plasma cells in PD.
Recognizing the practical challenges of obtaining brain tissue for routine diagnosis, we considered the more accessible and commonly used sample type, namely peripheral blood. Our intention was to validate the identified hub genes, derived from PD patients’ brain tissues, in peripheral blood. This approach is motivated by the fact that obtaining brain tissue for diagnosing suspected PD patients is impractical in routine clinical settings. Peripheral blood, being a commonly used and less invasive sample, is more feasible for routine diagnostic checks. We aimed to assess whether the gene expression patterns in blood align with those in brain tissues, with the goal of reducing diagnostic complexity and improving efficiency. However, our study has some limitations. First, even though we combined the datasets due to the lower number of samples in PD, studies with bigger sample sizes should be performed to confirm our findings. Second, we utilized RT-qPCR to validate the conclusions in blood samples of PD patients. Other experimental verification methods should be used to verify our results. Nowadays, larger-scale single cell RNA sequencing analysis and multi-omics with more clinical samples is needed to further elucidate the exact mechanism of the disease. We can make it realize in PD in the further research.
In conclusion, the developed diagnostic model provides new insights for early stage PD diagnosis. Plasma cells were found to be differentially expressed between HC and PD patients. Elucidation of the genetic and immunological mechanisms that underlie the initial signs of PD will unlock new therapeutic avenues. These insights will empower clinicians to effectively intervene with innovative or repurposed anti-inflammatory and immunomodulatory treatments, with the aim of slowing down the progression of this disease.
Conclusion
The SLC18A2, TAC1, PCDH8, KIAA0319, PDE6H, AXIN1, and AGTR1 are associated with PD pathology, and are potential diagnostic markers for PD. Besides, immune cell infiltrations might play an important role in PD. These findings hold promising implications for PD diagnosis and treatment.
Data availability
All sequencing data generated in this study are deposited in the GEO database. (GSE7672, GSE20141, GSE20159, GSE20163, GSE20164, and GSE20292).
Abbreviations
- 3D:
-
3 Dimension
- AT1R:
-
Angiotensin II type 1 receptor
- AUC:
-
Area under the curve
- BP:
-
Biological process
- CC:
-
Cellular component
- DA:
-
Dopamine
- DEGs:
-
Differentially expressed genes
- FC:
-
Fold change
- GEO:
-
Gene expression omnibus
- GO:
-
Gene ontology
- HC:
-
Healthy control
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- MF:
-
Molecular function
- PCA:
-
Principal component analysis
- PD:
-
Parkinson's disease
- PPI:
-
Protein–protein interaction
- ROC:
-
Receiver operating characteristic
- RRA:
-
Robust rank aggregation
- RT-qPCR:
-
Reverse transcription-quantitative real-time polymerase chain reaction
- SN:
-
Substantia nigra
- SP:
-
Substance P
References
Feigin, V. L. et al. Global, regional, and national burden of neurological disorders, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18, 459–480 (2019).
Dorsey, E. R., Sherer, T., Okun, M. S. & Bloem, B. R. The emerging evidence of the Parkinson pandemic. J. Park. Dis. 8, S3–S8 (2018).
Hayes, M. T. Parkinson’s disease and parkinsonism. Am. J. Med. 132, 802–807 (2019).
Tysnes, O.-B. & Storstein, A. Epidemiology of Parkinson’s disease. J. Neural Transm. Vienna Austria 1996(124), 901–905 (2017).
Armstrong, M. J. & Okun, M. S. Diagnosis and treatment of Parkinson disease: A review. JAMA 323, 548 (2020).
Kalia, L. V. & Lang, A. E. Parkinson’s disease. The Lancet 386, 896–912 (2015).
Ascherio, A. & Schwarzschild, M. A. The epidemiology of Parkinson’s disease: risk factors and prevention. Lancet Neurol. 15, 1257–1272 (2016).
Kline, E. M. et al. Genetic and environmental factors in p arkinson’s disease converge on immune function and inflammation. Mov. Disord. 36, 25–36 (2021).
Riboldi, G. M. et al. Transcriptome deregulation of peripheral monocytes and whole blood in GBA-related Parkinson’s disease. Mol. Neurodegener. 17, 52 (2022).
Zeis, P. et al. In situ maturation and tissue adaptation of type 2 innate lymphoid cell progenitors. Immunity 53, 775-792.e9 (2020).
Kurvits, L. et al. Transcriptomic profiles in Parkinson’s disease. Exp. Biol. Med. 246, 584–595 (2021).
Zhao, S. et al. Identification and validation of neurotrophic factor-related gene signatures in glioblastoma and Parkinson’s disease. Front. Immunol. 14, 1090040 (2023).
Tomkins, J. E. & Manzoni, C. Advances in protein-protein interaction network analysis for Parkinson’s disease. Neurobiol. Dis. 155, 105395 (2021).
Szklarczyk, D. et al. The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49, D605–D612 (2021).
Kim, H. et al. Modeling G2019S-LRRK2 sporadic Parkinson’s disease in 3D midbrain organoids. Stem Cell Rep. 12, 518–531 (2019).
Tansey, M. G. et al. Inflammation and immune dysfunction in Parkinson disease. Nat. Rev. Immunol. 22, 657–673 (2022).
Heavener, K. S. & Bradshaw, E. M. The aging immune system in Alzheimer’s and Parkinson’s diseases. Semin. Immunopathol. 44, 649–657 (2022).
Chen, B., Khodadoust, M.S., Liu, C.L., Newman, A.M. & Alizadeh, A.A. Profiling tumor infiltrating immune cells with CIBERSORT. in Cancer Systems Biology (ed. Von Stechow, L.) vol. 1711 243–259 (Springer New York, 2018).
Earls, R. H. et al. Intrastriatal injection of preformed alpha-synuclein fibrils alters central and peripheral immune cell profiles in non-transgenic mice. J. Neuroinflammation 16, 250 (2019).
Kolde, R., Laur, S., Adler, P. & Vilo, J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics 28, 573–580 (2012).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucl. Acids Res. 51, D587–D592 (2023).
Zhang, P.-L., Chen, Y., Zhang, C.-H., Wang, Y.-X. & Fernandez-Funez, P. Genetics of Parkinson’s disease and related disorders. J. Med. Genet. 55, 73–80 (2018).
Levin, J., Kurz, A., Arzberger, T., Giese, A. & Höglinger, G. U. The differential diagnosis and treatment of atypical Parkinsonism. Dtsch. Ärztebl. Int. https://doi.org/10.3238/arztebl.2016.0061 (2016).
Aarsland, D. et al. Parkinson disease-associated cognitive impairment. Nat. Rev. Dis. Primer 7, 47 (2021).
Zheng, B. et al. PGC-1 α, a potential therapeutic target for early intervention in Parkinson’s disease. Sci. Trans. Med. 2(52), 5273 (2011).
Kelly, J., Moyeed, R., Carroll, C., Albani, D. & Li, X. Gene expression meta-analysis of Parkinson’s disease and its relationship with Alzheimer’s disease. Mol. Brain. 12(1), 1 (2019).
Hirsch, E. C. & Hunot, S. Neuroinflammation in Parkinson’s disease: a target for neuroprotection?. Lancet Neurol. 8, 382–397 (2009).
Warren, N., O’Gorman, C., Lehn, A. & Siskind, D. Dopamine dysregulation syndrome in Parkinson’s disease: a systematic review of published cases. J. Neurol. Neurosurg. Psychiatry 88, 1060–1064 (2017).
Cliburn, R. A. et al. Immunochemical localization of vesicular monoamine transporter 2 (VMAT2) in mouse brain. J. Chem. Neuroanat. 83–84, 82–90 (2017).
Baronio, D. et al. Vesicular monoamine transporter 2 (SLC18A2) regulates monoamine turnover and brain development in zebrafish. Acta Physiol. 234(1), e13725 (2022).
Jiménez-Jiménez, F. J., Alonso-Navarro, H., García-Martín, E. & Agúndez, J. A. G. Thr105Ile (rs11558538) polymorphism in the histamine N-methyltransferase (HNMT) gene and risk for Parkinson disease: A PRISMA-compliant systematic review and meta-analysis. Med. Baltimore 95, e4147 (2016).
Wang, Q. et al. Substance P exacerbates dopaminergic neurodegeneration through neurokinin-1 receptor-independent activation of microglial NADPH oxidase. J. Neurosci. 34, 12490–12503 (2014).
Kim, S. Y. et al. The expression of non-clustered protocadherins in adult rat hippocampal formation and the connecting brain regions. Neuroscience 170, 189–199 (2010).
Franquinho, F. et al. The dyslexia-susceptibility protein KIAA0319 inhibits axon growth through Smad2 Signaling. Cereb. Cortex 27, 1732–1747 (2017).
Wu, G.-D., Li, Z.-H., Li, X., Zheng, T. & Zhang, D.-K. microRNA-592 blockade inhibits oxidative stress injury in Alzheimer’s disease astrocytes via the KIAA0319-mediated Keap1/Nrf2/ARE signaling pathway. Exp. Neurol. 324, 113128 (2020).
Cope, E. C. & Gould, E. Adult neurogenesis, glia, and the extracellular matrix. Cell Stem Cell 24, 690–705 (2019).
Paniagua, S. et al. Dyslexia associated gene KIAA0319 regulates cell cycle during human neuroepithelial cell development. Front. Cell Dev. Biol. 10, 967147 (2022).
Baburski, A. Z., Sokanovic, S. J., Andric, S. A. & Kostic, T. S. Aging has the opposite effect on cAMP and cGMP circadian variations in rat Leydig cells. J. Comp. Physiol. B 187, 613–623 (2017).
Zhang, G. et al. HIF-1α/microRNA-128-3p axis protects hippocampal neurons from apoptosis via the Axin1 -mediated Wnt/β-catenin signaling pathway in Parkinson’s disease models. Aging 12, 4067–4081 (2020).
Zawada, W. M. et al. Loss of angiotensin II receptor expression in dopamine neurons in Parkinson’s disease correlates with pathological progression and is accompanied by increases in Nox4- and 8-OH guanosine-related nucleic acid oxidation and caspase-3 activation. Acta Neuropathol. Commun. 3, 9 (2015).
Grammatopoulos, T. N. et al. Angiotensin type 1 receptor antagonist losartan, reduces MPTP-induced degeneration of dopaminergic neurons in substantia nigra. Mol. Neurodegener. 2, 1 (2007).
Sathiya, S. et al. Telmisartan attenuates MPTP induced dopaminergic degeneration and motor dysfunction through regulation of α-synuclein and neurotrophic factors (BDNF and GDNF) expression in C57BL/6J mice. Neuropharmacology 73, 98–110 (2013).
Prolonged compound muscle action potential duration in critical illness myopathy—Goodman—2009—Muscle & Nerve—Wiley Online Library. https://onlinelibrary.wiley.com/doi/https://doi.org/10.1002/mus.21445.
Toth, C. et al. Levodopa, methylmalonic acid, and neuropathy in idiopathic Parkinson disease. Ann. Neurol. 68, 28–36 (2010).
Abbruzzese, G., Vische, M., Ratto, S., Abbruzzese, M. & Favale, E. Assessment of motor neuron excitability in parkinsonian rigidity by the F wave. J. Neurol. 232, 246–249 (1985).
Miah, I. P., Olde Dubbelink, K. T., Stoffers, D., Deijen, J. B. & Berendse, H. W. Early-stage cognitive impairment in Parkinson’s disease and the influence of dopamine replacement therapy. Eur. J. Neurol. 19, 510–516 (2012).
Weintraub, D. et al. Impulse control disorders in Parkinson disease: A cross-sectional study of 3090 patients. Arch. Neurol. 67(5), 589–595 (2010).
Rilstone, J. J., Alkhater, R. A. & Minassian, B. A. Brain dopamine-serotonin vesicular transport disease and its treatment. N. Engl. J. Med. 368, 543–550 (2013).
Li, Q. et al. Genetic variations in the 3′-untranslated region of SLC18A2 are associated with serum FSH concentration in polycystic ovary syndrome patients and regulate gene expression in vitro. Hum. Reprod. 31, 2150–2157 (2016).
Stevens, C. H. et al. Reduced T helper and B lymphocytes in Parkinson’s disease. J. Neuroimmunol. 252, 95–99 (2012).
Funding
This study was supported by the National Natural Science Foundation of China (Grant no: 81860142).
Author information
Authors and Affiliations
Contributions
Conceptualization: J.H., B.L., H.W. and S.C.; Data curation: J.H., B.L. and S.C.; Analysis: J.H., B.L. and S.C.; Writing: J.H., B.L. and S.C.; Preparation: J.H., B.L., C.L. and S.C.; Writing—Review & Editing: J.H., B.L. and S.C.; Funding acquisition: J.H., B.L., S.C. and H.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, J., Li, B., Wei, H. et al. Integrative analysis of gene expression profiles of substantia nigra identifies potential diagnosis biomarkers in Parkinson's disease. Sci Rep 14, 2167 (2024). https://doi.org/10.1038/s41598-024-52276-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52276-0
- Springer Nature Limited