Abstract
Genotyping of gDNA rs12041331 (PEAR1), rs6065 (GP1BA), and rs730012 (LTC4S) can provide systematic guidance on the use of aspirin. However, an accurate, reliable and economical approach to simultaneous detection of the above single nucleotide polymorphisms (SNPs) is not reported. Herein, we designed and substantiated an allele-specific (AS) forward primer-superposed amplification analysis for measurement of the SNPs in PEAR1, GP1BA and LTC4S genes, in which the values of ∆Cq (differences in threshold cycles between the wild-type forward primer-based assay and the mutated-type forward primer-based assay) were employed to decide genotype. Mismatch AS forward primers were screened with the singleplex amplification analysis. Moreover, Cq extension optimized by AS forward primer superposition was observed in the selected forward primer-based triplex analysis. Further, robustness assessment of the triplex analysis showed the amplification efficiency ranging from 0.9 to 1.1. Precision test demonstrated the coefficient of variation of less than 2%. And the detective results of 189 DNA samples was completely concordant with that of commercial Sanger sequencing. In summary, we developed a simple, accurate and economical approach to genotyping of rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S) to provide a valuable pharmacogenomics tool for guidance of aspirin delivery.
Similar content being viewed by others
Introduction
Cardiovascular diseases, consisting of hypertension, hyperlipidemia, coronary heart disease, atherosclerosis, etc., have become the top killers of human health1,2,3. The prevention and treatment of cardiovascular diseases are increasingly becoming a focus of attention in medical field. Because of multifaceted causes and manifestations, the control of cardiovascular disease demands the use of compound formulation, commonly called polypills, which can ominously decrease the incidence of cardiovascular diseases4, especially cerebral apoplexy and myocardial infarct, by 35–50%5,6,7,8. The components of the polypill comprise statins, aspirin and a kind of antihypertensive drug9. The polypill concept is increasingly being recognized and accepted in clinical practice.
As a platelet aggregation antagonist, aspirin is widely used to decline the risk of suspected patients with acute myocardial infarction, and prevent cardiovascular diseases such as recurrence of myocardial infarction, stroke and atherosclerosis10,11,12. In vivo, aspirin can inactivate cyclooxygenase (COX) by irreversible acetylating the hydroxyl group of serine residues in the active part of COX13,14,15. This impedes platelet aggregation through constraining both the metabolism of arachidonic acid (AA) and the production of thromboxane A2 (TXA2)13,16,17,18. Aspirin resistance (AR) and adverse reactions were found in some patients who took routine dose of aspirin19. AR is defined as a lower-than-expected inhibition effect of aspirin on platelet aggregation, raising the risk of recurrent cerebral infarction and other cerebrovascular events20,21,22,23. Clinic studies exhibited a 5–60% incidence of AR and a 1.5% frequency of aspirin hypersensitivity causing allergy and hemorrhage in patients with cardiovascular disease24,25,26.
Aspirin pharmacogenomics evidenced that genetic polymorphism of PEAR1 (rs12041331), GP1BA (rs6065) and LTC4S (rs730012) is associated with AR and adverse reactions27,28,29,30,31. In 2020, a scoring table for genotyping of gDNA rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S) was proposed to provide systematic guidance for aspirin administration32. However, an accurate and reliable approach to simultaneous examination of the above single nucleotide polymorphisms (SNPs) is not reported. Herein, we designed and substantiated an allele-specific (AS) forward primer-superposed amplification analysis for discrimination of the SNPs in PEAR1 (rs12041331), GP1BA (rs6065) and LTC4S (rs730012) genes. The results insinuated that this analysis is a valuable tool to facilitate personalized antiplatelet therapy.
Materials and methods
Design strategy
First, mismatch AS F-primers were screened with singleplex amplification analysis. Next, the selected F-primers-based triplicate analysis was optimized by F-primer superposition to avoid undetermined results. Then, the optimized analysis was validated by robustness assessment and precision evaluation, as well as agreement analysis compared with Sanger sequencing. The values of ∆Cq (differences in threshold cycles between the wild-type F-primer-based amplification assay and the mutated-type F-primer-based amplification assay) were calculated to decide the outcomes.
DNA extraction from buccal swab
The human buccal swab samples used in this study involved 189 Chinese volunteers, which was not a train set classified by aspirin resistance. This study was approved by the Biomedical Research Ethic Committee of Shandong Provincial Hospital (No.2023-417) and has been conducted in accordance with ethical standards and guidelines of the Biomedical Research Ethic Committee of Shandong Provincial Hospital. Authors of this work extend a statement assuring that this work was conducted in accordance with the Declaration of Helsinki and obtained informed consent from all participants. Genomic DNA was extracted by using the QIAamp DNA Mini kit (Cat No. 51304, QIAGEN, Dusseldorf, Germany), and the procedure was carried out according to the instructions. DNA concentration was examined with a NanoPhotometer P360 (Implen GmbH, Munich, Germany). The quality was determined by using OD 260 / 280 ratio. Sanger sequencing was conducted by Personal Biotechnology Co., Ltd (Qingdao, China).
Primers and probes
AS F-primers, reverse primers and hydrolysis probes were designed using Primer Express 3.0, based on the information of the whole gene sequence. The second or fifth mismatch base was introduced at the 3′ end of F-primers, which were screened in a subsequent process. Probes for rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S) were labelled at the 5′ end with the fluorescent dye FAM, VIC, and NED, respectively, and at the 3′ end with the quencher BHQ1, BHQ1, and BHQ3, respectively. The oligonucleotide was synthesized by Sangon Biotech (Shanghai, China).
Real- time amplification assay
Triplex amplification analysis (TaqMan qPCR) was executed in a total of 20 μL reaction mixture, which contained 10 μL AceQ® Universal U+ Probe Master Mix V2 (Vazyme, Nanjing, China), 0.2 μM of each wild/mutated-type F-primer, 0.2 μM of each reverse primer, 0.1 μM of hydrolysis probe and 10 ng DNA template. The singleplex amplification analysis was conducted according to the same protocol. A robotic liquid handling workstation (epMotion 5075 vt, Germany) was utilized to dispense the mix. The reaction protocols started with a contamination digestion step for 2 min at 37℃ and a pre-denaturation step for 5 min at 95 ℃, followed by 45 cycles of 95 ℃ for 10 s, and 60 ℃ for 35 s. Fluorescence data were collected at 60 ℃. These amplifications were performed on the ABI7500 Real-Time PCR Instrument (ThermoFisher Scientific Inc., MA, USA).
Data analysis
GraphPad Prism software version 9.5 (GraphPad Software, Inc., San Diego, CA) was used to conduct data analysis and graphing.
Results and discussion
Screening of mismatch AS F-primers by singleplex amplification analysis
AS F-primers with a second or fifth mismatch base at 3' terminus were screened by detection of homozygote/heterozygote using singleplex real-time amplification analysis (10 ng DNA/test). And ΔCq (differences in threshold cycles between the wild-type F-primer-based amplification assay and the mutated-type F-primer-based amplification assay) was utilized to determine genotype. The principle for selection of the F-primer is the following: (a) no undetermined result was observed. (b) the Cq value was approximately 35 when wild homozygotes were detected in mutated-type F-primer-based amplification assay or when mutated homozygotes were measured in wild-type F-primer-based amplification assay. The original Cq values obtained from singleplex amplification analysis were shown in Table 1. And the selected sequences were shown in Table 2.
PCR-based analysis is a convenient tool to discriminate SNPs. Genetic polymorphism specific-binding molecules in PCR-based analysis comprise dsDNA-binding dye, AS probe and primer33,34. The dsDNA-binding dye-based high-resolution dissolution curve (HRM) assay needs specific equipment module. Besides diseconomy, it is time-consuming and laborious to discover appropriate Minor Groove Binder (MGB) probe31,35. For enhancement of AS primer specificity, base mismatch is more economic than locked nucleic acid (LNA) decoration32. In present study, the mismatch AS primers as polymorphism specific-binding molecules were screened and utilized to discriminate homozygotes/heterozygotes for rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S).
Development of triplex amplification analysis optimized by F-primer superposition
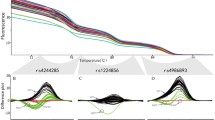
Based on the selected mismatch AS F-primers, we developed a triplex amplification analysis optimized by F-primer superposition. Undetermined results were observed when homozygotes were measured by un-optimized triplex amplification analysis (Fig. 1A). The extension of Cq was forced by the mismatch AS F-primer superposition, which was implemented with the addition of 0.01 μM mutated/wild-type F-primer into 0.2 μM wild/mutated-type F-primer-based amplification assay (Fig. 1B). The results showed that all outcomes of homozygotes were positive, suggesting that the mismatch AS F-primer superposition can improve detective convenience via omitting positive controls in the triplex amplification analysis.
Amplification plots of the triplex amplification analysis. The wild-type F-primer-based amplification assay and the mutated-type F-primer-based amplification assay were used to detect wild-type, mutated-type and heterozygous gDNA. (A) Amplification plots of un-optimized triplex amplification analyses. (B) Amplification plots of optimized triplex amplification analyses. Represented amplification plots are shown. The reaction was run in duplicate, in which 10 ng genomic DNA was inputted. Wt gDNA, wild-type genomic DNA; Mut gDNA, mutated-type genomic DNA; Het gDNA, heterozygote genomic DNA; WF primer, wild-type forward primer; MF primer, mutated-type forward primer.
Robustness assessment of triplex amplification analysis
To assess robustness of the triplex amplification analysis, the heterozygote was gradually reduced to generate gDNA samples at levels of 40 ng, 20 ng, 10 ng, 5 ng, 2.5 ng and 1.25 ng. Reactions were run in duplicate with three independent experiments. We used the following formula to calculate the amplification efficiency: 10−1/slope − 1, when the logarithm of the template concentration was plotted on the x-axis and Cq was plotted on the y-axis. The results demonstrated that the amplification efficiency calculated from standard curve ranged from 0.9 to 1.1 (Fig. 2), and limit of detection (LOD) was at least 1.25 ng/test.
Robustness assessment of the triplex amplification analysis. The robustness assessment was executed by employing mismatch allele-specific forward primers targeting single nucleotide polymorphism to simultaneously detect heterozygotes for PEAR1 (rs12041331), GP1BA (rs6065) and LTC4S (rs730012). Serial dilutions of heterozygote (1.25–40 ng) were measured by the triplex amplification analysis. Reactions were run in duplicate with three independent experiments. (A) Standard curve of the triplex amplification analysis. Amplification efficiency (Eff) % and R2 are shown. Data are expressed as mean ± SE. (B) Amplification plots of the robustness assessment. Represented amplification plots are shown. WF primer, wild-type forward primer; MF primer, mutated-type forward primer.
Precision evaluation of triplex amplification analysis
The precision of the triplex amplification analysis was evaluated by detection of genomic DNA at 10 ng/test and 2.5 ng/test levels. Each specimen was tested in eight plicates by two operators with two reagent lots every day over 5 days (n = 80/specimen) at one site. A total of eighty Cq values were collected to calculate the coefficient of variance (CV). The results revealed that CV value was < 2% for all days, specimens, replicates, operators and reagent lots combined. Figure 3 shows the intra-day CV for PEAR1(rs12041331), GP1BA (rs6065) and LTC4S (rs730012).
Intra-day coefficient of variant (CV) for PEAR1 (rs12041331), GP1BA (rs6065) and LTC4S (rs730012). Genomic DNA at 10 ng (A) or 2.5 ng (B) level was measured in eight plicates by two operators with two reagent lots every day over 5 days (n = 80/specimen) at one site. Data were expressed as CV median in wild (a) or mutated-type (b) forward primer-based amplification assay.
Agreement analysis between triplex amplification analysis and Sanger sequencing
We conducted the triplex amplification analysis on each of 189 samples, in which 89, 165, and 136 specimens were defined as homozygotes for rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S), respectively, while 100, 24, and 53 samples were defined as heterozygotes for rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S), respectively. The cut-off values for genotyping were described in Table 3.
Pharmacogenomics appears that some SNPs are more likely to initiate AR and adverse reactions36,37. Behaving as a kind of platelet transmembrane protein, the platelet endothelial aggregation receptor 1 (PEAR1) plays an important role in platelet aggregation. And genetic polymorphism of the rs12041331 in PEAR1 gene can obviously affect the inhibitive effect of aspirin on platelet aggregation38. Glycoprotein Ib-alpha (GP1BA) gene encodes platelet surface membrane glycoprotein (GPIb) that is a heterodimer consisting of bisulfide-linked α and β subunits, and acts as a receptor for von Willebrand factor(VWF)39. Genetic polymorphism of the rs6065 in GP1BA gene was evidenced to correlate aspirin resistance40,41. It was documented that patient carrying C-type allele for rs730012 in leukotriene C4 synthase (LTC4S) gene are prone to aspirin-induced urticaria42,43. Consulting to a scoring table proposed by Guangdong Pharmaceutical Association (Guangzhou, China)32 (Table 4), a triplex amplification analysis to detect genetic polymorphism of gDNA rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S) was designed and substantiated in this study. The results of the agreement analysis indicated the genotyping outlined by the triplex amplification analysis is consistent with the results obtained from Sanger sequencing.
In summary, we established a simple, efficient and accurate approach to the determination of genetic polymorphism of gDNA rs12041331 (PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S), which can be used to guide aspirin delivery to reduce AR and adverse reaction.
Data availability
Primer and probe sequences related to this study are available within this manuscript. The primers and probes were designed in accordance with the gene sequences of rs12041331(PEAR1), rs6065 (GP1BA) and rs730012 (LTC4S), which are available in NCBI database. Raw data are available from the corresponding authors upon reasonable request.
References
Flora, G. D. & Nayak, M. K. A brief review of cardiovascular diseases, associated risk factors and current treatment regimes. Curr. Pharm. Des. 25, 4063–4084. https://doi.org/10.2174/1381612825666190925163827 (2019).
Roth, G. A. et al. Global, regional, and national age-sex-specific mortality for 282 causes of death in 195 countries apnd territories, 1980–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet 392, 1736–1788. https://doi.org/10.1016/s0140-6736(18)32203-7 (2018).
Kyu, H. H. et al. Global, regional, and national disability-adjusted life-years (DALYs) for 359 diseases and injuries and healthy life expectancy (HALE) for 195 countries and territories, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet 392, 1859–1922. https://doi.org/10.1016/s0140-6736(18)32335-3 (2018).
Yusuf, S. & Pinto, F. J. The polypill: From concept and evidence to implementation. Lancet 400, 1661–1663. https://doi.org/10.1016/s0140-6736(22)01847-5 (2022).
Yusuf, S. et al. Blood-pressure and cholesterol lowering in persons without cardiovascular disease. N. Engl. J. Med. 374, 2032–2043. https://doi.org/10.1056/NEJMoa1600177 (2016).
Roshandel, G. et al. Effectiveness of polypill for primary and secondary prevention of cardiovascular diseases (PolyIran): A pragmatic, cluster-randomised trial. Lancet 394, 672–683. https://doi.org/10.1016/s0140-6736(19)31791-x (2019).
Joseph, P. et al. Fixed-dose combination therapies with and without aspirin for primary prevention of cardiovascular disease: An individual participant data meta-analysis. Lancet 398, 1133–1146. https://doi.org/10.1016/s0140-6736(21)01827-4 (2021).
Yusuf, S. et al. Polypill with or without aspirin in persons without cardiovascular disease. N. Engl. J. Med. 384, 216–228. https://doi.org/10.1056/NEJMoa2028220 (2021).
Webster, R., Castellano, J. M. & Onuma, O. K. Putting polypills into practice: Challenges and lessons learned. Lancet 389, 1066–1074. https://doi.org/10.1016/s0140-6736(17)30558-5 (2017).
Meher, S., Duley, L., Hunter, K. & Askie, L. Antiplatelet therapy before or after 16 weeks’ gestation for preventing preeclampsia: An individual participant data meta-analysis. Am. J. Obstet. Gynecol. 216, 121-128.e122. https://doi.org/10.1016/j.ajog.2016.10.016 (2017).
Huang, J. et al. Aspirin and heparin for the prevention of pre-eclampsia: Protocol for a systematic review and network meta-analysis. BMJ Open 9, e026920 (2019).
Kim, I. S. et al. Reduced metabolic activity of gut microbiota by antibiotics can potentiate the antithrombotic effect of aspirin. Biochem. Pharmacol. 122, 72–79 (2016).
Karsten, S. Aspirin and platelets: The antiplatelet action of aspirin and its role in thrombosis treatment and prophylaxis. Sem. Thromb Hemost. 23, 349–356 (1997).
Zhao, Y., Yang, S. & Wu, M. Mechanism of improving aspirin resistance: Blood-activating herbs combined with aspirin in treating atherosclerotic cardiovascular diseases. Front. Pharmacol. 12, 794417 (2021).
Finneran, M. M., Gonzalez-Brown, V. M., Smith, D. D., Landon, M. B. & Rood, K. M. Obesity and laboratory aspirin resistance in high-risk pregnant women treated with low-dose aspirin. Am. J. Obstet. Gynecol. 220, 385-e1 (2019).
Floyd, C. N. & Ferro, A. Antiplatelet drug resistance: Molecular insights and clinical implications. Prostaglandins Other Lipid Mediat. 120, 21–27. https://doi.org/10.1016/j.prostaglandins.2015.03.011 (2015).
Fuster, V. & Sweeny, J. M. Aspirin: A historical and contemporary therapeutic overview. Circulation 123, 768–778. https://doi.org/10.1161/circulationaha.110.963843 (2011).
Franchi, F. & Angiolillo, D. J. Novel antiplatelet agents in acute coronary syndrome. Nat. Rev. Cardiol. 12, 30–47. https://doi.org/10.1038/nrcardio.2014.156 (2015).
Gurbel, P. A. & Tantry, U. S. Antiplatelet drug resistance and variability in response: The role of antiplatelet therapy monitoring. Curr. Pharm. Des. 19, 3795 (2013).
Eikelboom, J. W. et al. Aspirin-resistant thromboxane biosynthesis and the risk of myocardial infarction, stroke, or cardiovascular death in patients at high risk for cardiovascular events. Circulation 105, 1650–1655. https://doi.org/10.1161/01.cir.0000013777.21160.07 (2002).
Gum, P. A., Kottke-Marchant, K., Welsh, P. A., White, J. & Topol, E. J. A prospective, blinded determination of the natural history of aspirin resistance among stable patients with cardiovascular disease. J. Am. Coll. Cardiol. 41, 961–965. https://doi.org/10.1016/s0735-1097(02)03014-0 (2003).
Patrono, C. Aspirin resistance: definition, mechanisms and clinical read-outs. J. Thromb. Haemost. JTH 1, 1710–1713. https://doi.org/10.1046/j.1538-7836.2003.00284.x (2003).
Yi, X., Zhou, Q., Lin, J. & Chi, L. Aspirin resistance in Chinese stroke patients increased the rate of recurrent stroke and other vascular events. Int. J. Stroke Off. J. Int. Stroke Soc. 8, 535–539. https://doi.org/10.1111/j.1747-4949.2012.00929.x (2013).
Du, G., Lin, Q. & Wang, J. H. A brief review on the mechanisms of aspirin resistance. Int. J. Cardiol. 220, 21–26. https://doi.org/10.1016/j.ijcard.2016.06.104 (2016).
Woessner, K. M. Aspirin desensitization for cardiovascular disease. Curr. Opin. Allergy Clin. Immunol. 15, 314–322. https://doi.org/10.1097/aci.0000000000000189 (2015).
Iwamoto, J., Saito, Y., Honda, A. & Matsuzaki, Y. Clinical features of gastroduodenal injury associated with long-term low-dose aspirin therapy. World J. Gastroenterol. 19, 1673–1682. https://doi.org/10.3748/wjg.v19.i11.1673 (2013).
Fujiwara, T. et al. Exploratory aspirin resistance trial in healthy Japanese volunteers (J-ART) using platelet aggregation as a measure of thrombogenicity. Pharmacogenomics J. 7, 395–403. https://doi.org/10.1038/sj.tpj.6500435 (2007).
Würtz, M., Nissen, P. H., Grove, E. L., Kristensen, S. D. & Hvas, A. M. Genetic determinants of on-aspirin platelet reactivity: focus on the influence of PEAR1. PloS ONE 9, e111816. https://doi.org/10.1371/journal.pone.0111816 (2014).
Lewis, J. P. et al. Genetic variation in PEAR1 is associated with platelet aggregation and cardiovascular outcomes. Circ.-Cardiovasc. Genet. 6, 184. https://doi.org/10.1161/circgenetics.111.964627 (2013).
Kauskot, A. et al. A novel mechanism of sustained platelet αIIbβ3 activation via PEAR1. Blood 119, 4056–4065. https://doi.org/10.1182/blood-2011-11-392787 (2012).
Ansari, N., Najafi, S., Shahrabi, S. & Saki, N. PEAR1 polymorphisms as a prognostic factor in hemostasis and cardiovascular diseases. J. Thromb. Thrombolysis 51, 89–95. https://doi.org/10.1007/s11239-020-02149-w (2021).
Association, G. P. Guidelines on individualized pharmaceutical care for antiplatelet drugs based on pharmacogenomics. Pharmacy Today, 030 (2020). Chinese with English abstract.
Nikodem, D., Cłapa, T. & Narożna, D. HRM-PCR in medical diagnostic. Postepy Biochem. 67, 59–63. https://doi.org/10.18388/pb.2021_373 (2021).
Matsuda, K. PCR-based detection methods for single-nucleotide polymorphism or mutation: Real-time PCR and its substantial contribution toward technological refinement. Adv. Clin. Chem. 80, 45–72. https://doi.org/10.1016/bs.acc.2016.11.002 (2017).
Cheng, H., Liu, Y., Fu, M., Liu, J. & Chen, G. Taqman-MGB FQ-PCR for human GPIIIa, GP1BA and PEAR1 SNPs. Clin. Lab. https://doi.org/10.7754/Clin.Lab.2022.220132 (2022).
Wang, J. et al. Association among PlA1/A2 gene polymorphism, laboratory aspirin resistance and clinical outcomes in patients with coronary artery disease: An updated meta-analysis. Sci. Rep. 9, 13177. https://doi.org/10.1038/s41598-019-49123-y (2019).
Wang, H. et al. Association of GPIa and COX-2 gene polymorphism with aspirin resistance. J. Clin. Lab. Anal. 32, e22331. https://doi.org/10.1002/jcla.22331 (2018).
Lewis, J. P. et al. Genetic variation in PEAR1, cardiovascular outcomes and effects of aspirin in a healthy elderly population. Clin. Pharmacol. Ther. 108, 1289–1298. https://doi.org/10.1002/cpt.1959 (2020).
Douglas, H. et al. Platelet membrane glycoprotein Ibalpha gene-5T/C Kozak sequence polymorphism as an independent risk factor for the occurrence of coronary thrombosis. Heart (British Cardiac Society) 87, 70–74. https://doi.org/10.1136/heart.87.1.70 (2002).
Matsubara, Y., Murata, M., Watanabe, G. & Ikeda, Y. Enhancing effect of the (145)Met-allele of GPIb alpha on platelet sensitivity to aspirin under high-shear conditions. Thromb. Res. 123, 331–335. https://doi.org/10.1016/j.thromres.2008.02.008 (2008).
Hong, X. et al. Simultaneous genotyping of human platelet alloantigen-1 to 28bw systems by multiplex polymerase chain reaction sequence-based typing. Vox Sang. 112, 360–366. https://doi.org/10.1111/vox.12507 (2017).
Sánchez-Borges, M. et al. The A-444C polymorphism in the leukotriene C4 synthase gene is associated with aspirin-induced urticaria. J. Investig. Allergol. Clin. Immunol. 19, 375–382 (2009).
Shi-E, L. et al. Association of LTC4S gene polymorphisms with childhood asthma and responsiveness to montelukast. Chin. J. Child Health Care. 26, 835–838, 842 (2018).
Acknowledgements
The work was supported by grants from the key project of international cooperation of Qilu University of Technology (No. QLUTGJHZ2018008).
Author information
Authors and Affiliations
Contributions
L.W.J. and H.Y. secured funding and designed the experiment, who are also guarantors of this work and, as such, had full access to all of the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis; L.C.Y., Y.P., Z.J., L.X.Q., L.Y.Q., Z.J. performed experiments and analyzed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, CY., Yang, P., Zheng, J. et al. Establishment of a forward primers-superposed amplification analysis for accurate aspirin pharmacogenomic measurement. Sci Rep 14, 880 (2024). https://doi.org/10.1038/s41598-024-51458-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-51458-0
- Springer Nature Limited