Abstract
The classical hypothesis proposes that the lack of recombination on sex chromosomes arises due to selection for linkage between a sex-determining locus and sexually antagonistic loci, primarily facilitated by inversions. However, cessation of recombination on sex chromosomes could be attributed also to neutral processes, connected with other chromosome rearrangements or can reflect sex-specific recombination patterns existing already before sex chromosome differentiation. Three Coleonyx gecko species share a complex X1X1X2X2/X1X2Y system of sex chromosomes evolved via a fusion of the Y chromosome with an autosome. We analyzed synaptonemal complexes and sequenced flow-sorted sex chromosomes to investigate the effect of chromosomal rearrangement on recombination and differentiation of these sex chromosomes. The gecko sex chromosomes evolved from syntenic regions that were also co-opted also for sex chromosomes in other reptiles. We showed that in male geckos, recombination is less prevalent in the proximal regions of chromosomes and is even further drastically reduced around the centromere of the neo-Y chromosome. We highlight that pre-existing recombination patterns and Robertsonian fusions can be responsible for the cessation of recombination on sex chromosomes and that such processes can be largely neutral.
Similar content being viewed by others
Introduction
Sex chromosomes have been in a multitude of shapes before they assumed a consistent form1,2. They can evolve from a pair of autosomes that acquired a sex-determining locus3. Although many sex chromosomes stay undifferentiated for a quite long evolutionary time4, in many lineages recombination between nascent sex chromosomes becomes suppressed around the sex-determining locus, and the non-recombining zone eventually spreads to almost the entire length of the chromosome pair5. Because of recombination suppression, many Y or W chromosomes diverge from their X or Z counterparts, accumulating deleterious mutations due to Muller’s ratchet6. In extreme cases, Y and W chromosomes may become fully heterochromatic or even lost7,8. Several hypotheses try to explain why there is a suppression of recombination in sex chromosomes. According to the classical model, the cessation of recombination is favored by natural selection preferring linkage between the sex-determining locus and sexually antagonistic alleles, that is, alleles that are beneficial for one sex and detrimental to the other9. This model is supported for example by studies on the guppy fish (Poecilia reticulata), a classical model species for exploring sex chromosome evolution10,11. However, alternative models suggest that stepwise suppression of recombination around the SDL (sex-determining locus) might occur due to neutral processes such as due to emergence of a sex-determining locus in an already ancestrally poorly recombining regions in a given sex, or mutation-induced cessation of recombination12,13. In this case, the presence of sexually antagonistic alleles or at least alleles beneficial only to one sex in the non-recombining region of sex chromosomes would be a consequence of recombination suppression rather than its cause. However, these models do not seem mutually exclusive and sex chromosomes in different lineages might utilize different mechanisms of recombination suppression in the heterogametic sex14.
Geckos (infraorder Gekkota) represent a good model group for studies onmechanisms of recombination suppression in sex chromosomes. Some gecko lineages have a putatively ancestral environmental sex determination system (ESD), while others evolved sex chromosomes of both male (XY) and female (ZW) heterogametic types from different ancestral autosomes15,16,17,18,19. The New World eublepharid genus Coleonyx comprises two lineages: the Central American clade containing C. mitratus, C. elegans and C. nemoralis, and the northern clade containing all the remaining species20,21,22. C. elegans has a complex male heterogametic sex chromosome system with X1X1X2X2 sex chromosomes in females (2n = 32) and X1X2Y sex chromosomes in males (2n = 31). Previously, we have shown that the whole chromosome probe derived from the Y chromosome of C. elegans hybridizes in this species with X1, X2 and Y chromosomes, suggesting that the multiple sex chromosomes evolved via afusion of the ancestral acrocentric Y with an acrocentric autosome now playing the role of a neo-X chromosome22,23. Alternatively, the polymorphic fused chromosomes could have been initially autosomal, and could have acquired the SDL later. The Y chromosome is the only non-acrocentric chromosome in the karyotype of C. elegans. It was suggested that the sex chromosomes of this species are poorly differentiated23,24; however, comparative genome coverage analysis using Illumina reads reveal some Y chromosome areas that are apparently degenerate25. These areas are homologous to parts of chicken (Gallus gallus) chromosomes 1, 6 and 11 (GGA1, GGA6, and GGA11 respectively). qPCR analysis reveal that the same areas are degenerate in C. mitratus as well, but its chromosomes were not cytogenetically characterized. The system of multiple sex chromosomes is missing in other eublepharid geckos and it was demonstrated that congeneric C. brevis possess independently evolved XX/XY sex chromosomes21,23,25.
In this study, we used cross-species chromosome painting with flow-sorted sex chromosome libraries of C. elegans to cytologically confirm their identity with the sex chromosomes of C. mitratus and sequenced these libraries to reveal their genomic content and elucidate the origin of the sex chromosomes of these geckos. Further, we performed immunolocalization of SYCP3, the main protein of the lateral elements of synaptonemal complexes (SC), and MLH1, a mismatch repair protein marking mature recombination nodules, to assess recombination suppression between the Y chromosome and the two X chromosomes. This method is widely used to analyze meiotic pairing and crossing over between chromosomes in vertebrates, including geckos26.
Material and methods
Specimens, DNA sequencing, assembly, annotation and COI barcoding
A group of C. mitratus was acquired from a commercial seller and kept according to the recommendations of Seufer et al.27. The species was identified by morphology as described by Klauber28, and species identification was further supported by similarity analysis of mitochondrial DNA sequences (DNA barcoding). All methods were performed in accordance with the relevant guidelines and regulations. All samples used in this study are listed in Table 1. DNA was extracted from muscle tissue of a male individual using the standard phenol–chloroform technique29. A library for low-coverage genomic sequencing was prepared using TruSeq Nano DNA Low Throughput Library Prep (Illumina), following the manufacturer's protocol. Paired-end sequencing was performed on Illumina MiSeq using ReagentKit v2 with 600-cycles (Illumina). The NGS data were deposited in the NCBI SRA database under accession number PRJNA945407. Raw Illumina reads were trimmed using Cutadapt 4.230. The complete mitochondrial genome for performing DNA barcoding was assembled de novo using GetOrganelle 1.7.7.0 pipeline31. Mitochondrial genome was annotated using MITOS232 and deposited in GenBank under accession number OQ644632. The coding sequence of the cytochrome oxidase subunit I (COI) gene was extracted from the assembly and homology search was performed using the online NCBI BLAST tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and the BOLD database33.
Synaptonemal complex preparation and immunostaining
SC spreads were prepared via drying-down technique described by Peters et al.34. Immunofluorescence staining was performed according to the protocol by Anderson et al.35. Prior to immunostaining, slides were incubated in a solution of 10% PBT (PBS with 3% bovine serum albumin and 0.05% Tween 20) and 90% PBS for 45 min to reduce non-specific antibody binding. Primary antibodies included rabbit polyclonal anti-SYCP3 antibodies (1:500; Abcam, ab15093), human anticentromere antibodies (1:100; Antibodies Inc., 15-234) and mouse monoclonal anti-MLH1 antibodies (1:30, Abcam, ab14206). The slides were incubated with antibodies overnight in a humid box at 37 ℃, and then washed three times in PBS with 0.1% Tween 20 for 15 min each time. Secondary antibodies included Cy3-conjugated goat anti-rabbit (1:500; Jackson ImmunoResearch, 111-165-144), FITC-conjugated donkey anti-human (1:100; Jackson ImmunoResearch, 709-095-149) and FITC-conjugated goat anti-mouse (1:30; Jackson ImmunoResearch, 115-095-003). The slides were incubated with them for 1 h under the same conditions. After washing, the slides were mounted in Vectashield medium with DAPI (Vector Laboratories, cat No. H-1000-10) under the coverslips. Microscopic analysis and image processing were performed as described previously36.
Cell cultures and mitotic chromosome preparation
Cell cultures were prepared from tissues of four C. mitratus embryos dissected from eggs that had been incubated for one month at 28 ℃, and from the thorax tissues of one subadult male. The cultures were established in the Laboratory of Animal Cytogenetics, Institute of Molecular and Cellular Biology, Russia, using enzymatic treatment of tissues as described previously37,38. The cell culture lines were deposited in the Core Facilities Center “Cryobank of cell cultures” IMCB SB RAS. Metaphase chromosome spreads were prepared from chromosome suspensions obtained from early passages of primary fibroblast cultures as described previously39,40,41.
Flow-sorted chromosome libraries and FISH
Flow sorting of C. elegans chromosomes has been previously described23,42. Painting probes were prepared by DOP-PCR amplification of flow sorted chromosomes and labeled with biotin-dUTP and digoxigenin-dUTP (Sigma) by secondary DOP-PCR amplification as described previously43,44. The ribosomal DNA probe was obtained from plasmid DNA (pHr13), containing human partial 28S, full 5.8S, partial 18S ribosomal genes and two internal spacers45. The telomeric DNA probe was generated by PCR with oligonucleotides (TTAGGG)5 and (CCCTAA)546. Labeling was performed using the «FTP-Display» DNA fragmentation kit (DNA-Display, Russia) by incorporation of biotin-dUTP and digoxigenin (dUTP). Dual-color ZooFISH was performed according to a previously published protocol47. Briefly, freshly made chromosome preparations were aged for 1 h at 65 °C and treated with pepsin. Chromosome denaturation was done in 70% formamide with 2 × SSC at 70 °C for 1 min. Hybridization mixture contained a hybridization buffer (50% formamide, 10% dextran sulfate, 2 × SSC), 0.2% Tween 20, 1.5 µg sonicated genomic DNA of C. mitratus and 0.1 µg of each labeled painting probe. Probes were denatured at 95 °C for 5 min and preannealed at 45 °C for 1 h. Hybridization was carried out at 40 °C for 48 h. The slides were analyzed with fluorescence microscope Olympus BX53 using Video-Test-FISH (VideoTestT, Saint-Petersburg, Russia) digital imaging systems.
ChromSeq analysis
The DNA pools of the C. elegans chromosomes Y, X1, and X2 were sequenced as described above. The NGS data were deposited in the NCBI SRA database under accession number PRJNA945407. The resulting reads were aligned to the genome of Eublepharis macularius (Emac_v1.0.1)48 and the genome of Anolis carolinensis (AnoCar2.0)49, improved by the DNA Zoo Consortium50,51, using the DOPseq pipeline52. Synteny between reference genomes was determined using D-GENIES53.
Ethical statement
All manipulations with animals were approved by the Institute of Molecular and Cellular Biology Ethics Committee (statement №01/21 from 26/01/2021).
Results
Species identification
The species identity of animals was confirmed by morphological traits and by DNA barcoding. The COI haplotype of the sequenced specimen showed 99.55% similarity with the available C. mitratus partial COI gene sequence (GenBank record ON873271), and maximum 98.92% similarity with C. mitratus COI sequences in the BOLD database. The assembled C. mitratus mitochondrial genome was deposited in GenBank under accession number OQ644632.
Karyotypes and ZooFISH
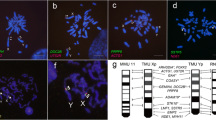
The mitotic karyotypes of the male and of three embryos contained 2n = 31 chromosomes, with one unpaired large metacentric chromosome. One embryo had 2n = 32 chromosomes, all acrocentric. ZooFISH with probes of C. elegans Y and X1 chromosomes painted three chromosomes in the 2n = 31 embryos (two acrocentrics and the metacentric) and four acrocentric chromosomes in the 2n = 32 embryo (Fig. 1a, b). Thus, the metacentric chromosome was identified as the Y chromosome. The terminal parts of X1 and Yq contained the DAPI-negative nucleolus organizer, revealed by FISH with rDNA probe. The telomeric probe hybridized to terminal parts of all chromosomes with no interstitial signals (Fig. 1c). Inverted DAPI images of the metaphases in Fig. 1a–c are presented in Fig. 1d–f, respectively.
Fluorescence in situ hybridization with flow-sorted chromosome-specific probes of male C. elegans, 45S rDNA and telomeric probes on metaphase of C. mitratus (a–c) and inverted DAPI images of the same spreads (d–f). (a) Y chromosome specific probe (red) and X1 chromosome specific probe (green) on C. mitratus male metaphase. (b) Y chromosome-specific probe (red) and X1 chromosome specific probe (green) on C. mitratus female metaphase. (c) 45S rDNA probe (red) and telomeric probe (green) on C. mitratus male metaphase. Bar: 10 µm.
Synapsis and recombination of the sex chromosomes and autosomes
The SC karyotypes of the three adult males had 15 elements: 14 bivalents and the sex trivalent (Fig. 2). Recombination was studied in detail in males 1 and 2. The total SC lengths of these specimens were 186.7 ± 25.5 μm and 194.4 ± 48.9 μm (mean ± SD, 100 spreads per individual analyzed, t-test = 0.14, p = 0.89, no significant difference between individuals). In the sex trivalents, synapsis was initiated at the terminal parts. Telomeric initiation of synapsis is typical for chromosomes of vertebrates and was observed in lizards before26,54. The median part where the centromeric areas of the two X chromosomes fused showed delayed synapsis, indicated by incomplete synapsis when the autosomes were already fully paired, and often non-homologous pairing between the tips of the X chromosomes was observed (Fig. 3). The SC spreads of the males had 18.5 ± 1.1 and 18.7 ± 0.9 MLH1 foci (mean ± SD, 100 spreads per individual analyzed, t-test = 0.13, p = 0.9, no significant difference between individuals). Only spreads with at least one focus at each SC were considered. The sex trivalent had either two MLH1 foci, with one focus per arm (84% and 98% in two males), three foci with two foci in one arm and one in another (6% and 1% in each male, respectively), or rarely only one focus (10% and 1% in each male, respectively). The distal parts of all chromosome arms showed pronounced recombination peaks, and most acrocentric autosomal SCs also displayed minor recombination peaks close to centromeres. The proximal crossovers occurred both as single crossovers and as second crossovers in the chromosomes with MLH1 foci in both peaks. The median parts of all SCs had lower recombination than the ends. No MLH1 foci were detected in the median part of the sex trivalent (Fig. 4).
A pachytene spermatocyte of Coleonyx mitratus, immunolabeled with antibodies to SYCP3 (red), MLH1 (green) and centromeric proteins (green). Centromeres are indicated by larger and diffuse signals, whereas the MLH1 foci are small and round. In the upper tip of the rightmost autosomal bivalent, non-specific binding of anti-centromere antibodies in a heterochromatic area is present. Insert shows a scheme of the sex trivalent. Arrowheads show centromeres at the sex trivalent. Bar: 10 µm.
(a–d) Putative consecutive stages of sex chromosomes synapsis of C. mitratus after immunolocalization of SYCP3 (red), MLH1 (green) and centromeric proteins (green). Centromeres are indicated by larger and diffuse signals, whereas the MLH1 foci are small and round. (e–h) Schematic drawings of the sex trivalents. Arrowheads indicate centromeres at sex trivalents. Bar: 2 µm.
Numbers and distributions of the MLH1 foci on sex chromosome trivalents and two autosomal bivalents (chromosomes 5 and 6) in C. mitratus. The x-axis shows the positions of MLH1 foci along the SCs in relation to the centromere (black arrowheads). One scale division represents a segment of the average length of each SC equal to 1 µm. The y-axis shows the proportion of MLH1 foci in each interval. Different colors show SCs with different MLH1 numbers, from 1 to 3.
Genetic content of the sex chromosomes
According to gene content, the Y chromosome of C. elegans is homologous to A. carolinensis chromosomes 3p, 6q, 8 and 12 (ACA3p, ACA6q, ACA8, ACA12), or E. macularius chromosomes 6, except for the terminal 12.5 Mb (EMA6, homologous to ACA3p), chromosome 12 (EMA12, homologous to ACA6q and ACA12), and distal part of chromosome 16, except the first 10.7 Mb (EMA16, the aligned part is homologous to ACA8) (Fig. 5). According to the synteny analysis, the proximal part (approximately 12 Mb) of EMA16, not found in the Y chromosome of C. elegans, is homologous to a part of ACA5. The X1 chromosome was homologous to ACA6q, ACA8 and ACA12, or EMA12 and the distal part of EMA16, except for the first 4.9 Mb. The X2 chromosome library contained DNA homologous to ACA3 and ACA4q, or EMA6 and EMA5 (Fig. 5). The detailed DOPseq results are presented in Supplementary File 1.
Homology between the sex chromosomes of C. mitratus and C. elegans and the chromosomes of reference species: G. gallus (GGA), A. carolinensis (ACA), E. macularius (EMA). Purple shading shows homology. The homologues of the proximal segments of the X chromosomes are absent in the Y chromosome due to its degeneration24.
Discussion
Sex chromosome synapsis and recombination
We demonstrate recombination suppression in the pericentromeric regions of the sex trivalent of C. mitratus, which corresponds to the earlier reported degenerate part of the Y chromosome in C. elegans25. The recombination suppression of these sex chromosomes can be attributed to several mechanisms. First, the centromere has generally a strong suppressive effect on recombination as has been demonstrated in most plants and animals examined so far55, although the underlying mechanisms remain unclear. Most chromosomes examined in reptiles, birds and mammals show a polarized distribution of recombination events, with peaks at chromosome ends and valleys in the middle56,57. In C. mitratus, the proximal crossover peaks in acrocentric autosomal SCs are much weaker than those in Trapelus sanguinolentus (Agamidae) and in birds36,58, indicating a strong centromeric suppression of recombination in the gecko.
Delayed synapsis, observed in the sex trivalent of C. mitratus, is typical for autosomal Robertsonian trivalents in many mammalian species and it is hence not special for sex chromosomes59,60,61. Metacentric SCs emerged by Robertsonian fusions in shrews, and in most mouse models, have significantly lower recombination in the centromeric region than the same chromosomes in the acrocentric form, but heterozygous metacentric trivalents do not have lower recombination in the centromeric region than homozygous metacentric bivalents59,61,62. Thus, we suggest that the Robertsonian heterozygosity probably has minor, if any, influence on the recombination suppression.
The total number of crossing over events on the sex chromosome trivalent in C. mitratus could be also the same as in the ancestral all-acrocentric states, but they can differ in distribution being localized more towards the end of the Y chromosome. Overall, the already low recombination during male meiosis near centromeres also in autosomes, and low recombination in central parts of large chromosomes could all result in the complete cessation of recombination in the median part of the Y chromosome in C. mitratus. In female meiosis, such large metacentric chromosomes may have more even recombination distribution due to heterochiasmy63. However, for the male-specific Y chromosome this means a complete cessation of recombination in the median part.
According to the classical model of sex chromosome differentiation, recombination between heterologous sex chromosomes is blocked by inversions or specific epigenetic modifications6. Here, the sex chromosomes probably followed another pathway: recombination in the Y chromosome was very likely suppressed due to neutral mechanistic reasons connected with the Robertsonian fusion of ancestral acrocentric Y and an acrocentric autosome, and to the general recombination patterns in males of this species11,13. Nevertheless, changes in gene presence and expression around the sex-determining locus before and after Robertsonian fusion can contribute to fixation of this multiple sex chromosome system in the common ancestor of C. elegans and C. mitratus.
Sex chromosome contents and homology
The karyotype of C. mitratus was identical to the previously known karyotype of C. elegans23, in accordance with their sex chromosome identity25 and similarity of flow-sorted karyotypes42. The degenerate parts of C. mitratus and C. elegans Y chromosomes are homologous to a part of chicken chromosome 11 (GGA11, homologous to ACA8), small proximal part of chromosome 6 (GGA6), and small median part of chromosome 1 (GGA1) as was shown in the previous study based on a different technique, i.e. the comparison of coverage between sexes25. In Podarcis muralis, both latter parts are homologous to the proximal part of chromosome 5 (homologous to ACA3p).
By sequencing the chromosome-specific DNA libraries of C. elegans, we were able to reveal the remaining fragments that did not experience degeneration and were therefore not detected by the previous coverage analysis. We assigned these fragments to chromosomes X1 and X2. Moreover, data on Y chromosome degeneration, synapsis data and crossover map allow us to reconstruct the orientation of the syntenic regions inside the sex chromosomes. First, the synaptic configuration of the sex trivalent shows that the formation of the Y chromosome resulted from a centromere-to-centromere, but not centromere-to-telomere, or telomere-to-telomere fusion. Second, the suppression of recombination in the median part of the trivalent shows that the Y-degenerate parts are in the proximal segments of the X chromosomes. The schematic reconstruction is shown in Fig. 5. The only uncertain element is the order of the ACA6q-homologous and ACA12-homologous fragments.
The association between the homolog of ACA8 and a part of ACA5, found in EMA16, was not detected in C. elegans. This may be caused by either an E. macularius-specific translocation or a genome assembly error. The terminal part of EMA6, showing very low coverage with Y chromosome reads, probably corresponds to the degenerate part. The degenerate part of the ACA8-homologous segment was not detected by DOPseq either due to low coverage in this region or lower level of degeneration. The presence of DNA homologous to EMA5 (ACA4q) in the X2 probe indicates contamination with a similar-size autosome.
The recruitment of autosomes for the role of sex chromosomes is non-random, and some genomic regions are more frequently involved in sex chromosome formation due to their genetic content64. Several squamate species have been found to use the synthetic regions found on the C. elegans and C. mitratus sex chromosomes as part of their sex chromosomes. The involvement of ACA3p/GGA6 and ACA8/GGA11 was discussed previously25. ACA6q/GGA27 is a conserved element of ZZ/ZW sex chromosomes of caenophidian snakes65 and XX/XY sex chromosomes of Python bivittatus66. The syntenic block homologous to ACA12/GGA14 has been reported as part of the ZZ/ZW sex chromosomes of the marbled gecko (Christinus marmoratus)67, and as the pseudoautosomal part of the XX/XY sex chromosomes of the brown anole (Norops sagrei)68.
Conclusion
In the current study we revealed a possible mechanism of recombination suppression in the sex chromosomes of C. mitratus via Robertsonian fusion of sex chromosomes and reduced overall recombination in pericentromeric regions in male meiosis, as reported for other cases in which Robertsonian translocations are present. This finding is consistent with the neutral hypotheses of sex chromosome recombination suppression12,13, although further tests of gene content and gene expression are needed to determine the involvement of sexually antagonistic selection. Accurate fine-scale assemblies of the three sex chromosomes of these geckos and a study of female meiosis are required to understand the recombination between them in further detail. The identification of all syntenic regions involved in the formation of this complex sex chromosome system, as performed in this study, will help to understand the possibility of parallel co-option of different syntenic regions during sex chromosome formation in vertebrates.
Data availability
All raw sequence data are deposited in the NCBI SRA database under accession number PRJNA945407.
References
PokornÁ, M. & Kratochvíl, L. Phylogeny of sex-determining mechanisms in squamate reptiles: Are sex chromosomes an evolutionary trap?. Zool. J. Linn. Soc. 156, 168–183 (2009).
Alam, S. M. I., Sarre, S. D., Gleeson, D., Georges, A. & Ezaz, T. Did lizards follow unique pathways in sex chromosome evolution?. Genes (Basel) 9, 20–28 (2018).
Bachtrog, D. et al. Sex determination: Why so many ways of doing it?. PLoS Biol. 12, 1–13 (2014).
Kamiya, T. et al. A trans-species missense SNP in Amhr2 is associated with sex determination in the tiger Pufferfish, Takifugu rubripes (Fugu). PLoS Genet. 8, e1002798 (2012).
Torgasheva, A. et al. Highly conservative pattern of sex chromosome synapsis and recombination in neognathae birds. Genes 12, 1358 (2021).
Charlesworth, D. Evolution of recombination rates between sex chromosomes. Philos. Trans. R. Soc. B Biol. Sci. 372, 1736 (2017).
Charlesworth, D. The timing of genetic degeneration of sex chromosomes. Philos. Trans. R. Soc. B Biol. Sci. 376, 20200093 (2021).
Borodin, P. M. et al. Multiple independent evolutionary losses of XY pairing at meiosis in the grey voles. Chromosom. Res. 20, 259–268 (2012).
Rice, W. R. The accumulation of sexually antagonistic genes as a selective agent promoting the evolution of reduced recombination between primitive sex chromosomes. Evolution (N. Y.) 41, 911–914 (1987).
Wright, A. E. et al. Convergent recombination suppression suggests role of sexual selection in guppy sex chromosome formation. Nat. Commun. 8, 14251 (2017).
Bergero, R., Gardner, J., Bader, B., Yong, L. & Charlesworth, D. Exaggerated heterochiasmy in a fish with sex-linked male coloration polymorphisms. Proc. Natl. Acad. Sci. U. S. A. 116, 6924–6931 (2019).
Jeffries, D. L., Gerchen, J. F., Scharmann, M. & Pannell, J. R. A neutral model for the loss of recombination on sex chromosomes. Philos. Trans. R. Soc. B Biol. Sci. 376, 20200096 (2021).
Perrin, N. Sex-chromosome evolution in frogs: What role for sex-antagonistic genes?. Philos. Trans. R. Soc. B Biol. Sci. 376, 20200094 (2021).
Ponnikas, S., Sigeman, H., Abbott, J. K. & Hansson, B. Why do sex chromosomes stop recombining?. Trends Genet. 34, 492–503 (2018).
Augstenova, B., Pensabene, E., Vesel, M., Kratochvıl, L. & Rovatsos, M. Are Geckos special in sex determination? Independently evolved differentiated ZZ/ZW sex chromosomes in carphodactylid geckos. Genome Biol. Evol. 13, evab119 (2021).
Gamble, T. et al. XX/XY sex chromosomes in the South American Dwarf Gecko (Gonatodes humeralis). J. Hered. 109, 462–468 (2018).
Pensabene, E., Yurchenko, A., Kratochvíl, L. & Rovatsos, M. Madagascar leaf-tail geckos (Uroplatus spp.) share independently evolved differentiated ZZ/ZW sex chromosomes. Cells 12, 260 (2023).
Rovatsos, M., Farkačová, K., Altmanová, M., Johnson-Pokorná, M. & Kratochvíl, L. The rise and fall of differentiated sex chromosomes in geckos. Mol. Ecol. 28, 3042–3052 (2019).
Gamble, T. et al. Restriction site-associated DNA sequencing (RAD-seq) reveals an extraordinary number of transitions among gecko sex-determining systems. Mol. Biol. Evol. 32, 1296–1309 (2015).
Agarwal, I. et al. The evolutionary history of an accidental model organism, the leopard gecko Eublepharis macularius (Squamata: Eublepharidae). Mol. Phylogenet. Evol. 168, 107414 (2022).
Keating, S. E., Greenbaum, E., Johnson, J. D. & Gamble, T. Identification of a cis-sex chromosome transition in banded geckos (Coleonyx, Eublepharidae, Gekkota). J. Evol. Biol. 35, 1675–1682 (2022).
Butler, B. O., Smith, L. L. & Flores-Villela, O. Phylogeography and taxonomy of Coleonyx elegans Gray 1845 (Squamata: Eublepharidae) in Mesoamerica: The Isthmus of Tehuantepec as an environmental barrier. Mol. Phylogenet. Evol. 178, 107632 (2023).
Pokorná, M. et al. Differentiation of sex chromosomes and karyotypic evolution in the eye-lid geckos (Squamata: Gekkota: Eublepharidae), a group with different modes of sex determination. Chromosom. Res. 18, 809–820 (2010).
Pokorná, M., Kratochvíl, L. & Kejnovský, E. Microsatellite distribution on sex chromosomes at different stages of heteromorphism and heterochromatinization in two lizard species (Squamata: Eublepharidae: Coleonyx elegans and Lacertidae: Eremias velox). BMC Genet. 12, 1–7 (2011).
Pensabene, E., Kratochvíl, L. & Rovatsos, M. Independent evolution of sex chromosomes in eublepharid geckos, a lineage with environmental and genotypic sex determination. Life 10, 1–11 (2020).
Dedukh, D., Altmanova, M., Klima, J. & Kratochvil, L. Premeiotic endoreplication is essential for obligate parthenogenesis in geckos. Development 149, dev200345 (2022).
Seufer, H., Kaverkin, Y. & Kirschner, A. The Eyelash Geckos: Care, Breeding and Natural History (Kirschner & Seufer, 2005).
Klauber, L. M. The Geckos of the Genus Coleonyx: With descriptions of new subspecies. Trans. San Diego Soc. Nat. Hist. 10, 133–216 (1945).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular cloning: A laboratory manual. Mol. Cloning Lab. Manual. 1989, 852 (1989).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Jin, J.-J. et al. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes Germplasm Bank of Wild Species. Genome Biol. 21, 1–31 (2020).
Donath, A. et al. Improved annotation of protein-coding genes boundaries in metazoan mitochondrial genomes. Nucleic Acids Res. 47, 10543–10552 (2019).
Ratnasingham, S. & Hebert, P. D. N. BOLD: The barcode of life data system (www.barcodinglife.org). Mol. Ecol. Notes 7, 355–364 (2007).
Peters, A., Plug, A., Van Vugt, M. & De Boer, P. A drying-down technique for the spreading of mammalian meiocytes from the male and female germline. Chromosom. Res. 5, 66–68 (1997).
Anderson, L. K., Reeves, A., Webb, L. M. & Ashley, T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151, 1569–1579 (1999).
Lisachov, A. P., Tishakova, K. V., Tsepilov, Y. A. & Borodin, P. M. Male meiotic recombination in the Steppe Agama, Trapelus sanguinolentus (Agamidae, Iguania, Reptilia). Cytogenet. Genome Res. 157, 107–114 (2019).
Stanyon, R. & Galleni, L. A rapid fibroblast culture technique for high resolution karyotypes A rapid fibroblast culture technique for high resolution karyotypes. Ital. J. Zool. 58, 81–83 (1991).
Romanenko, S. A. et al. Segmental paleotetraploidy revealed in sterlet (Acipenser ruthenus) genome by chromosome painting. Mol. Cytogenet. 8, 90 (2015).
Yang, F. et al. A complete comparative chromosome map for the dog, red fox, and human and its integration with canine genetic maps. Genomics 62, 189–202 (1999).
Graphodatsky, A. et al. Comparative cytogenetics of hamsters of the genus Calomyscus. Cytogenet. Cell Genet. 88, 296–304 (2000).
Graphodatsky, A. S., Yang, F., O’brien, P. C. M. & Perelman, P. Phylogenetic implications of the 38 putative ancestral chromosome segments for four canid species Create new project ‘Ultrasonic energy transport proved using femtosecond laser pulses’ View project Cytogenomics in Amazonian bats View project. Cytogenet. Genome Res. 92, 243–247 (2001).
Kasai, F., O’Brien, P. C. M. & Ferguson-Smith, M. A. Squamate chromosome size and GC content assessed by flow karyotyping. Cytogenet. Genome Res. 157, 46–52 (2019).
Telenius, H. et al. Cytogenetic analysis by chromosome painting using DOP-PCR amplified flow-sorted chromosomes. Genes Chromosom. Cancer 4, 251–263 (1992).
Yang, F., Carter, N. P., Shiu, L. & Ferguson-Smith, M. A. A comparative study of karyotypes of muntjacs by chromosome painting. Chromosoma 103, 642–652 (1995).
Maden, B. E. H. et al. Clones of human ribosomal DNA containing the complete 18 S-rRNA and 28 S-rRNA genes. Characterization, a detailed map of the human ribosomal transcription unit and diversity among clones. Biochem. J. 246, 519–527 (1987).
Ijdo, J. W., Wells, R. A., Baldini, A. & Reeders, S. T. Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucleic Acids Res. 19, 4780 (1991).
Yang, F. & Graphodatsky, A. S. Animal probes and ZOO-FISH. In Fluorescence In Situ Hybridization (FISH)—Application Guide 323–346 (Springer, 2009). https://doi.org/10.1007/978-3-540-70581-9_29.
Pinto, B. J. et al. The revised reference genome of the leopard gecko (Eublepharis macularius) provides insight into the considerations of genome phasing and assembly. J. Hered. 2023, esad016. https://doi.org/10.1093/jhered/esad016 (2023).
Alföldi, J. et al. The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature 477, 587–591 (2011).
Dudchenko, O. et al. The Juicebox Assembly Tools module facilitates de novo assembly of mammalian genomes with chromosome-length scaffolds for under $1000. bioRxiv https://doi.org/10.1101/254797 (2018).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science (80-) 356, 92–95 (2017).
Makunin, A. I. et al. Contrasting origin of B chromosomes in two cervids (Siberian roe deer and grey brocket deer) unravelled by chromosome-specific DNA sequencing. BMC Genom. 17, 618 (2016).
Cabanettes, F. & Klopp, C. D-GENIES: Dot plot large genomes in an interactive, efficient and simple way. PeerJ 2018, e4958 (2018).
Marín-Gual, L. et al. Meiotic chromosome dynamics and double strand break formation in reptiles. Front. Cell Dev. Biol. 10, 1009776 (2022).
Vincenten, N. et al. The kinetochore prevents centromere-proximal crossover recombination during meiosis. Elife 4, e10850 (2015).
Lisachov, A. P., Trifonov, V. A., Giovannotti, M., Ferguson-Smith, M. A. & Borodin, P. M. Immunocytological analysis of meiotic recombination in two anole lizards (Squamata, Dactyloidae). Comp. Cytogenet. 11, 129–141 (2017).
Basheva, E. A., Torgasheva, A. A., Sakaeva, G. R., Bidau, C. & Borodin, P. M. A- and B-chromosome pairing and recombination in male meiosis of the silver fox (Vulpes vulpes L., 1758, Carnivora, Canidae). Chromosom. Res. 18, 689–696 (2010).
Semenov, G. A., Basheva, E. A., Borodin, P. M. & Torgasheva, A. A. High rate of meiotic recombination and its implications for intricate speciation patterns in the white wagtail (Motacilla alba). Biol. J. Linn. Soc. 125, 600–612 (2018).
Capilla, L. et al. Genetic recombination variation in wild Robertsonian mice: On the role of chromosomal fusions and Prdm9 allelic background. Proc. R. Soc. B Biol. Sci. 281, 20140297 (2014).
Davisson, M. T. & Akeson, E. C. Recomnination suppression by heterozygous Robertsonian chromosomes in the mouse. Genetics 133, 649–667 (1993).
Kierszenbaum, A. L. et al. Meiotic recombination and spermatogenic impairment in Mus musculus domesticus carrying multiple simple Robertsonian translocations. Cytogenet. Genome Res. 103, 321–329 (2003).
Borodin, P. M. et al. Recombination map of the common shrew, Sorex araneus (Eulipotyphla, Mammalia). Genetics 178, 621–632 (2008).
Mank, J. E. The evolution of heterochiasmy: The role of sexual selection and sperm competition in determining sex-specific recombination rates in eutherian mammals. Genet. Res. (Camb.) 91, 355–363 (2009).
O’Meally, D., Ezaz, T., Georges, A., Sarre, S. D. & Graves, J. A. M. Are some chromosomes particularly good at sex? Insights from amniotes. Chromosom. Res. 20, 7–19 (2012).
Rovatsos, M., Vukić, J., Lymberakis, P. & Kratochvíl, L. Evolutionary stability of sex chromosomes in snakes. Proc. R. Soc. B Biol. Sci. 282, 20151992 (2015).
Gamble, T. et al. The discovery of XY sex chromosomes in a boa and python. Curr. Biol. 27, 2148-2153.e4 (2017).
Zhu, Z. X. et al. Diversity of reptile sex chromosome evolution revealed by cytogenetic and linked-read sequencing. Zool. Res. 43, 719–733 (2022).
Kichigin, I. G. et al. Evolutionary dynamics of Anolis sex chromosomes revealed by sequencing of flow sorting-derived microchromosome-specific DNA. Mol. Genet. Genom. 291, 1955–1966 (2016).
Acknowledgements
The research was completed using equipment (materials) of the Core Facilities Centre “Cryobank of cell cultures” Institute of Molecular and Cellular Biology Siberian Branch of Russian Academy of Sciences (Novosibirsk, Russia). The research was funded by the Ministry of Science and Higher Education of the Russian Federation grant number FWNR-2022-0015 (for PB), research grant of the Russian Science Foundation 19-14-00034-P (for KT and SR) and Czech Science Foundation project number 20-27236J (for LK).
Author information
Authors and Affiliations
Contributions
A.L. designed the research and wrote an initial draft of the manuscript. K.T., L.L. and S.R. prepared the cell cultures of C. mitratus and made mitotic chromosome preparation. K.T. carried out FISH and SC analysis. G.D. sequenced the flow-sorted chromosome libraries. D.P. sequenced the genomic DNA. L.K. provided the samples and cell cultures of C. elegans. P.O`.B. and M.F.S. performed flow sorting. P.B. and V.T. carried out general project administration and acquired funding. All authors participated in editing and revising the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lisachov, A., Tishakova, K., Romanenko, S. et al. Robertsonian fusion triggers recombination suppression on sex chromosomes in Coleonyx geckos. Sci Rep 13, 15502 (2023). https://doi.org/10.1038/s41598-023-39937-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-39937-2
- Springer Nature Limited