Abstract
Lipids are highly structurally diverse molecules involved in a wide variety of biological processes. The involvement of lipids is even more pronounced in mycobacteria, including the human pathogen Mycobacterium tuberculosis, which produces a highly complex and diverse set of lipids in the cell envelope. These lipids include mycolic acids, which are among the longest fatty acids in nature and can contain up to 90 carbon atoms. Mycolic acids are ubiquitously found in mycobacteria and are alpha branched and beta hydroxylated lipids. Discrete modifications, such as alpha, alpha’, epoxy, methoxy, keto, and carboxy, characterize mycolic acids at the species level. Here, we used high precision ion mobility-mass spectrometry to build a database including 206 mass-resolved collision cross sections (CCSs) of mycolic acids originating from the strict human pathogen M. tuberculosis, the opportunistic strains M. abscessus, M. marinum and M. avium, and the nonpathogenic strain M. smegmatis. Primary differences between the mycolic acid profiles could be observed between mycobacterial species. Acyl tail length and modifications were the primary structural descriptors determining CCS magnitude. As a resource for researchers, this work provides a detailed catalogue of the mass-resolved collision cross sections for mycolic acids along with a workflow to generate and analyse the dataset generated.
Similar content being viewed by others
Introduction
Mycolic acids are long-chain fatty acids and comprise the inner leaflet of mycobacterial outer membranes in cell envelopes1,2,3,4,5. They occur mostly in association with sugar moieties and are either covalently linked to arabinogalactan or esterifying trehalose and glycerol; however, they may also exist freely3,4,6,7. Mycolic acids are important not only for mycobacterial cell envelope architecture but also for virulence and antibiotic resistance2,8,9. Mycolic acids determine cell envelope fluidity and create a hydrophobic layer in association with sugar moieties and lipids; this layer acts as a barrier for drug penetration, thereby conferring intrinsic resistance to antibiotics10. The synthesis of mycolic acids is targeted by the first-line anti-TB drug isoniazid through inhibiting the activities of InhA and KasA. Moreover, mycolic acids manipulate the host immune response7 and induce foamy macrophage formation characteristic of Mycobacterium spp.11.
The role of mycolic acids in virulence is determined by their structure. All mycolic acids share a main acyl chain, called the meromycolic chain, and an alpha branch4,5. Mycolic acids are divided into different classes according to the functional groups attached to the main meromycolic chain4,5,6,9,12. However, mycolic acid classes are species-specific and not present in all Mycobacterium species8,9,13. For example, the nonpathogenic mycobacterium Mycobacterium smegmatis mc2155 does not possess keto and methoxy groups, as found in the pathogenic mycobacterium strain Mycobacterium tuberculosis H37Rv9,14,15. Additionally, although epoxy mycolic acids are found in M. smegmatis mc2155, they are not present in M. bovis BCG, the vaccine strain9,14,16.
Recent studies have shown that mycolic acid chain length, functional groups and structures can influence the fluidity of the mycobacterial cell envelope, which in turn affects drug susceptibility10. Furthermore, the chemical functions on the meromycolic chain determine virulence1,9. For example, only mycobacteria with oxygenated mycolic acids can induce formation of foamy macrophages, which facilitate mycobacterial persistence within host macrophages11. Specifically, keto-mycolic acids are able to interact with the host lipid sensor TR4 and induce this foamy phenotype17. Methoxy and keto mycolic acids are also more antigenic than alpha mycolic acids18.
It is important to characterize the mycolic acid profiles of different mycobacterial species at a fine structural level to clarify their roles during infection and antimicrobial resistance between different species. Mycolic acids of the same classification may differ in the position and geometry of their chemical functions, adding to their structural complexity. Numerous studies have reported on the identification of mycolic acid classes in different mycobacterial species12,15,19 and investigated the roles of mycolic acid classes in virulence11,17,18. These approaches involve mass spectrometry techniques, such as matrix-assisted laser desorption ionization15 or electrospray ionization20,21,22 mass spectrometry.
However, none of these studies have investigated the use of ion mobility coupled to mass spectrometry to gain deeper insight into mycolic acids. Ion mobility-mass spectrometry (IM-MS) is a technique used to achieve molecular ion separation based on ionic mass, charge, and conformations in the gas phase23,24,25. In contrast to conventional mass spectrometry, the additional information on conformation, which is measured in terms of collisional cross section (CCS), improves the accuracy of identification of molecules with similar structures. Indeed, through ion mobility, the rotationally averaged CCS of an ion can be measured. CCS is an important distinguishing characteristic of an ion in the gas phase. It is related to its chemical structure and three-dimensional conformation.
This method has been previously used to identify lipid isomers, which are lipids that exhibit the same mass but different structures (hence, different CCS values), and classify them according to lipid head group, chain length, and degree of unsaturation26,27,28,29.
In this study, we focus on the relationship between lipid structure and gas-phase conformation via IM-MS analysis to generate an atlas of mycolic acid methyl ester (MAME) isomers in relevant mycobacterial species.
Results and discussion
Data processing workflow
A subset database from online sources (https://www.lipidmaps.org/) and the literature was created in comma-separated values (CSV) format15,30,31,32. Analysis of fragmentation data from the pooled quality control samples performed in Mass Hunter Qualitative Analysis (Agilent Technologies) was used to extract compounds with corresponding spectral information. Data were placed into Sirius 4 (Lehrstuhl Bioinformatik Jena) for molecular formula assignment. Fragment formulas were added to the database33,34. Finally, all compounds in the database were annotated with CCS using Mass Profiler (Agilent Technologies).
A screening workflow was set up in Skyline (MacCoss Lab), in which all mycolic acids were extracted from IM-QToF35. All Ions MS/MS data were collected with accurate mass, retention time and CCS (Fig. 1). Data were filtered to exclude matches with an isotope dot product score lower than 0.8 and a mass error of > 10 ppm.
Mycolic acid distribution and ion forms
MAMEs were identified and assigned primarily based on exact mass measurements and MS/MS profiles. The CCS measurements and corresponding mobility-mass correlations were utilized to provide additional confidence in the lipid assignment in conjunction with the mass measurement accuracy (± 10 ppm observed in this work).
This work represents 206 CCS values for uniquely identified mycolic acids from 5 mycobacterial species and 6 MAME families analysed in the positive ion mode and obtained on a drift tube instrument operated with nitrogen drift gas (DTCCSN2). Using the solvents required for chromatography, across all 5 mycobacterial species, ~ 76% and ~ 24% of the total ion abundance of corresponded to mycolic acid methyl ester species with proton [M + H]+ and sodium [M + Na]+ adducts, respectively (Fig. 2A). In previous literature, most MAMEs are reported as sodium or ammonium adducts36. In contrast, we observe that the large majority of adducts are [M + H]+, which can be explained by the preconditioning of the LC system by phosphoric acid; this was performed to reduce the presence of ions, such as Na+ and K+.
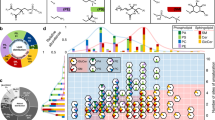
Mycolic acid methyl ester population observations. (a) The distribution of adducts observed resulting in the species of quasimolecular ion detected. (b) Mycolic acid methyl ester modification distributions across 5 mycobacterial species, M. tuberculosis, M. abscessus, M. avium, M. marinum and M. smegmatis.
In terms of mycolic acids categories, carboxy; methoxy; keto; epoxy/w1-methoxy; alpha’ and alpha were annotated. More particularly, in Mtb, alpha mycolic acid methyl esters represent 55%, keto mycolic acid methyl esters represent 39% and methoxy mycolic acid methyl esters represent 6% of the total MAME ion abundance measured. In M. abscessus, alpha mycolic acid methyl esters represent 55%, alpha’ mycolic acid methyl esters represent 32%, epoxy/w1 methoxy mycolic acid methyl esters represent 9% and carboxy mycolic acid methyl esters represent 5% of the total MAME ion abundance measured. In M. avium, alpha mycolic acid methyl esters represent 44%, epoxy/w1 methoxy mycolic acid methyl esters represent 6% and carboxy mycolic acid methyl esters represent 50% of the total MAME ion abundance measured. In M. marinum, alpha mycolic acid methyl esters represent 40%, keto mycolic acid methyl esters represent 52%, methoxy mycolic acid methyl esters represent 9% and carboxy mycolic acid methyl esters represent 8% of the total MAME ion abundance measured. In M. smegmatis, alpha mycolic acid methyl esters represent 58%, alpha’ mycolic acid methyl esters represent 14%, and epoxy/w1-methoxy mycolic acid methyl esters represent 28% of the total MAME ion abundance measured. These observations are summarized in the histogram in Fig. 2B and are in agreement with the thin layer chromatography from growing mycobacteria (Figs. S1 and S2). These distributions correlate well with those found in the literature using conventional methods14,37,38,39,40,41.
IM-MS correlation
In this work, all MAME classes exhibit a positive mobility-mass correlation in conformational space analyses, as typically observed with lipid classes in other studies. Within the MAME lipid trendline, unique families could be further differentiated by their respective DTCCSN2 information, as each family exhibited an average CCS increase in 0.1–0.2 Å2 per mass unit within the investigated mass range of 850–1400 m/z (Fig. 3, Supplementary Table 1). A closer examination of the IM-MS data shows a regular increase in size for each of the mycolic acid methyl ester families with a corresponding slope, DTCCSN2 vs. mass, ranging from 0.1–0.2 Å2 per mass unit.
Quantitative correlations within the cation data for mycolic acid methyl esters for (a) M. smegmatis, (b) M. abscessus, (c) M. avium, (d) M. marinum, and (e) M. tuberculosis. The colours correspond to mycolic acid methyl ester modifications, whereas the shapes correspond to cation type, as specified in the corresponding panel legends.
The CCS/mass relationship presented here describes MAMEs differing only in chain length, decorations, or adduct type and can be used to identify lipid features that are ambiguous to characterize by mass alone or conformations.
For protonated adducts, the same mass m/z can correlate to more than one CCS. These findings can also be observed for the alpha MAMEs in M. marinum at m/z 1152.202 correlated to CCS at 388.17 Å2 and 386.96 Å2, at m/z 1166.18 correlated to CCS at 389.36 Å2 and 392.10 Å2, at m/z 1180.232 correlated to CCS at 395.7 Å2 and 394.27 Å2, and for keto MAMEs at m/z 1224.257 correlated to CCS at 406.75 Å2, 408.12 Å2, 405.77 Å2, 407.19 Å2, 408.17 Å2, 405.32 Å2. In M. tuberculosis, alpha MAMEs at m/z 1152.201 correlated to CCS at 385.49 Å2 and 388.17 Å2, and m/z 1180.232 correlated to CCS at 395.7 Å2 and 394.27 Å2.
As in M. avium, these data indicate that for some carboxy MAMEs, a m/z can correlate to more than one CCS, such as for m/z 1001.987 with CCS at 340.01 Å2 and CCS at 348.89 Å2 (Fig. 4), suggesting that this lipid can adopt a different conformation in the gas phase. This information may indicate that the carboxy position is different in the meromycolic chain. A similar conclusion can be drawn for alpha’ mycolic acids, in which the double bond found in the meromycolic acid chain could be in different positions. As a result, the conformation of the meromycolic chain and therefore the CCS changes; however, the m/z remains the same, giving new insights into the complexity of mycolic acids.
In this study, 206 DTCCN2 measurements are presented for the MAMEs for 5 mycobacterial species representing 6 classes of decorations. This correlation m/z to CCS is unique to MAMEs and provides resources for future investigations. In addition, using this approach, we observed that MAMEs can adopt multiple CCS values, which could result from different conformations in the gas phase. The majority of isomeric mycolic acids belong to the alpha, keto and carboxy classes. Compared to the acyl chain length, the degree of unsaturation is four times more impactful on CCS26,28; thus, some MAMEs are more likely to form isomers, resulting in the same m/z but different CCS values. In conclusion, this is the first study reporting on a workflow and the use of IM-MS on MAMEs from 5 different mycobacterial lipids, providing a resource for researchers.
Overall, this study demonstrates that IM-MS improves separation of mycolic acids, especially between the different isomeric forms that share the same molecular weight. Considering the important roles of mycolic acids in mycobacterial virulence and their differential distribution across mycobacterial species, accurate separation and characterisation of mycolic acids would greatly help the research in this field. As demonstrated in this study, IM-MS could be a powerful tool for this purpose based on its accuracy and specificity compared with conventional mass spectrometry or ion mobility spectrometry alone. Specifically, in LC-IM-MS studies CCS values are being increasingly leveraged as molecular identifiers that provide an orthogonal dimension of information to mass and retention time, and many software workflows such as the screening workflow shown here are being adapted to accommodate filtering on CCS values for increased specificity for both targeted compound detection and unknown compound identification. To enable the mycobacterial research community, this study presents a database of mycolic acids from various mycobacterial species and an example workflow of data analysis for such purposes. We hope to encourage future studies on the species-specific mycolic acid profiles and their functions in mycobacterial species. However, validation of CCS values is restricted to the limited number of lipid standards that are commercially available for mycolic acids, making one of the limitations for this study.
Material and methods
Bacterial strains and growth conditions
Mycobacterium tuberculosis reference strain H37Rv, M. abscessus subsp. abscessus ATCC 19977, M. avium subsp. avium Chester ATCC 25291, M. marinum M and M. smegmatis mc2155 were grown in Middlebrook 7H9 liquid medium supplemented with 0.2% glycerol (w/v) and 10% enrichments containing BSA (Sigma) 5% (w/v), glucose (Sigma) 2% (w/v) and NaCl (Sigma) 0.85% (w/v) and left at + 37 °C and + 30 °C for M. marinum M to reach OD ~ 0.8–1, representing the mid-logarithmic phase of growth. To monitor the growth of the bacteria, control flasks were inoculated at an OD600 of 0.01 for the Mycobacterium tuberculosis reference strain H37Rv and an OD600 of 0.001 for M. abscessus subsp. abscessus ATCC 19977, M. avium subsp. avium Chester ATCC 25291, M. marinum M and M. smegmatis mc2155 and grown in 7H9 liquid medium supplemented with 0.2% glycerol (w/v) and 10% enrichments containing BSA (Sigma) 5% (w/v), glucose (Sigma) 2% (w/v), NaCl (Sigma) 0.85% (w/v) and 0.05% tyloxapol (Sigma).
Preparation of mycolic acid methyl esters
Mycolic acid methyl esters (MAMEs) were prepared as described by Bottai et al.42. Briefly, heat-killed mycobacteria were first washed 3 times with PBS. The pellets were then delipidated by CHCl3/MeOH 1:2 (v/v) extraction for 12 h at room temperature followed by one CHCl3/MeOH 1:1 (v/v) extraction and one CHCl3/MeOH 2:1 (v/v) extraction for 3 h at room temperature. To analyse the cell wall mycolic acids, the delipidated bacterial pellets were mixed with 2 ml 15% tetrabutyl ammonium hydroxide (TBAH) solution and incubated at 100 °C for 5 h. After cooling to room temperature for 1 h, the samples were incubated with 2 ml of CHCl3 and 100 μl of iodomethane for 1 h for methylation of the released mycolic acids. The organic lower layer after centrifugation at 2000×g at 4 °C for 10 min was collected and dried under a stream of N2. Dried methylated mycolic acids were resuspended in 3 ml diethyl ether and mixed thoroughly. All supernatants were collected after centrifugation at 2000×g at 4 °C for 10 min, transferred into a new glass tube and dried under a stream of nitrogen. Pellets containing MAMEs were resuspended in 50 μl of CHCl3 and 5 ml of MeOH and left at 4 °C overnight for precipitation. MAMEs were collected after centrifugation at 2000×g at 4 °C for 10 min and dried under a stream of nitrogen. The purified MAMEs were suspended in CHCl3 at a final concentration of 10 mg/ml and analysed by IM-MS in the positive ion mode.
Instrumentation
The instrumentation and experimental procedures are based on the work from Leaptrot et al.26. Briefly, a low-field drift-tube-based 6560 Ion Mobility LC/Q-TOF mass spectrometer (Agilent Technologies) was utilized to acquire accurate mass and CCS measurements from MAME samples. The instrument consists of an orthogonal electrospray ionization (ESI) source (Agilent Jet Stream) with a heated sheath gas nebulizer for desolvation and focusing of ions at atmospheric pressure. A single bore, resistively coated, glass capillary was used to transfer ions into the vacuum system. As ions exit the transfer capillary, they are directed by a high-pressure ion funnel into a trapping funnel with two wire grids for ion trapping and gating. Ions accumulate in the trapping funnel and are subsequently pulsed into the uniform field drift tube, which is approximately 78 cm in length. An additional rear ion funnel refocuses ions as they exit the drift tube and pass to a lower pressure region via a hexapole ion guide. Ions passed through a quadrupole mass filter and collision cell before mass measurement was performed in an orthogonal time-of-flight (TOF) mass spectrometer.
Experimental parameters
Data were acquired across all samples using UHPLC-IM-QToF. Chromatographic separation was performed using an Agilent 1290 Infinity II system (Agilent Technologies). Reversed-phase chromatography was performed using an InfinityLab RRHD Eclipse Plus C18 column, 2.1 × 50 mm, 1.8 µm (Agilent Technologies). The LC system was first preconditioned with phosphoric acid before sample acquisition. For this procedure, the LC was disconnected from the mass spectrometer, and the LC system and column were washed overnight with 0.5% (v/v) phosphoric acid at 0.1 mL/min in water/acetonitrile 20:80 (v/v), followed by a wash with LC/MS grade water for 4 h until the pH was neutral and finally reconnected to the MS system. The column temperature was set at 65 °C. Mobile phase A consisted of water/acetonitrile 40/60 + 0.1% formic acid + 10 mM ammonium formate, and mobile phase B consisted of isopropanol/acetonitrile 90/10 + 0.1% formic acid + 10 mM ammonium formate. The following gradient was applied at a flow rate of 0.4 ml/min: 0 min, 43% B; 0–0.55 min, 43% B; 0.55–3 min, 54% B; 3–3.2 min, 70% B; 3.2–4.5 min, 100% B; 4.5–10 min, 100% B; 10–10.5 min, 43% B; 10.5–12 min, 43% B. Mass axis calibration was achieved by continuous infusion after the chromatography of a reference mass solution using an isocratic pump connected to an ESI ionization source operated in positive-ion mode.
For all experiments, source conditions remained constant to ensure that the single field calibration could be used to measure CCS across all acquired data43. The following parameters were used: sheath gas temperature, 300 °C; nebulizer pressure, 35 psig; sheath gas flow, 11 l min−1; capillary voltage, 3500 V; nozzle voltage, 1000 V; and fragmentor voltage, 400 V. The data were collected in profile 2 GHz (extended dynamic range) mode.
IM-QToF All Ions MS/MS data acquisition
With the chromatographic conditions described fragmentation data were acquired using an “All Ions MS/MS” data independent acquisition approach with two alternating experiments. The first experiment was used to acquire full spectrum ion mobility data of precursors using no collision energy. The second experiment was then employed to collect fragmentation information across the same mass range as the previous experiment by applying a collision energy of 42 eV. Ion mobility acquisition settings were as follows: mass range 800–1400 m/z, frame rate 2.5 frames per second, maximum drift time 100 ms, trap full time 20,000 ms and trap release time 150 ms.
QToF iterative MS/MS
MS/MS was performed only on a pooled sample to acquire data-dependent fragmentation data. Fragmentation was performed in the collision cell following quadrupole isolation of precursor ions with a preference for mycolic acids using a preferred inclusion list. Alternate scans were performed first in TOF-only mode to collect and analyse full-spectrum MS precursor data and for mass axis calibration, a second scan with a collision energy of 20 eV and a third scan with an increased collision energy of 42 eV. Data acquisition speed was optimized for collision-induced dissociation (CID) experiments based on chromatographic peak and number of potential mycolic acid coeluting in the chromatographic space. Thus, 10 spectra/second was used to ensure sufficient MS/MS scans were acquired. In this mode, the pooled sample was injected multiple times, thereby significantly improving the coverage of precursors being selected for fragmentation.
Data availability
The datasets used and analysed during the current study are available from the corresponding author on reasonable request.
References
Batt, S. M., Minnikin, D. E. & Besra, G. S. The thick waxy coat of mycobacteria, a protective layer against antibiotics and the host’s immune system. Biochem. J. 477, 1983–2006. https://doi.org/10.1042/BCJ20200194 (2020).
Abrahams, K. A. & Besra, G. S. Mycobacterial cell wall biosynthesis: A multifaceted antibiotic target. Parasitology 145, 116–133. https://doi.org/10.1017/S0031182016002377 (2018).
Jackson, M. The mycobacterial cell envelope-lipids. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a021105 (2014).
Brennan, P. J. & Nikaido, H. The envelope of mycobacteria. Annu. Rev. Biochem. 64, 29–63. https://doi.org/10.1146/annurev.bi.64.070195.000333 (1995).
Minnikin, D. E., Minnikin, S. M., Parlett, J. H., Goodfellow, M. & Magnusson, M. Mycolic acid patterns of some species of Mycobacterium. Arch. Microbiol. 139, 225–231. https://doi.org/10.1007/BF00402005 (1984).
Brennan, P. J. Structure, function, and biogenesis of the cell wall of Mycobacterium tuberculosis. Tuberculosis (Edinb) 83, 91–97 (2003).
Verschoor, J. A., Baird, M. S. & Grooten, J. Towards understanding the functional diversity of cell wall mycolic acids of Mycobacterium tuberculosis. Prog. Lipid Res. 51, 325–339. https://doi.org/10.1016/j.plipres.2012.05.002 (2012).
Nataraj, V. et al. Mycolic acids: Deciphering and targeting the Achilles’ heel of the tubercle bacillus. Mol. Microbiol. 98, 7–16. https://doi.org/10.1111/mmi.13101 (2015).
Marrakchi, H., Laneelle, M. A. & Daffe, M. Mycolic acids: Structures, biosynthesis, and beyond. Chem Biol 21, 67–85. https://doi.org/10.1016/j.chembiol.2013.11.011 (2014).
Singh, R. et al. Recent updates on drug resistance in Mycobacterium tuberculosis. J. Appl. Microbiol. 128, 1547–1567. https://doi.org/10.1111/jam.14478 (2020).
Peyron, P. et al. Foamy macrophages from tuberculous patients’ granulomas constitute a nutrient-rich reservoir for M. tuberculosis persistence. PLoS Pathog. 4, e1000204. https://doi.org/10.1371/journal.ppat.1000204 (2008).
Daffe, M., Laneelle, M. A. & Lacave, C. Structure and stereochemistry of mycolic acids of Mycobacterium marinum and Mycobacterium ulcerans. Res. Microbiol. 142, 397–403. https://doi.org/10.1016/0923-2508(91)90109-n (1991).
Barry, C. E. 3rd. et al. Mycolic acids: Structure, biosynthesis and physiological functions. Prog. Lipid Res. 37, 143–179. https://doi.org/10.1016/s0163-7827(98)00008-3 (1998).
Daffé, M., Quémard, A. & Marrakchi, H. In Biogenesis of Fatty Acids, Lipids and Membranes (ed. Geiger, O.) 1–36 (Springer International Publishing, 2017).
Laval, F., Laneelle, M. A., Deon, C., Monsarrat, B. & Daffe, M. Accurate molecular mass determination of mycolic acids by MALDI-TOF mass spectrometry. Anal. Chem. 73, 4537–4544. https://doi.org/10.1021/ac0105181 (2001).
Barkan, D., Liu, Z., Sacchettini, J. C. & Glickman, M. S. Mycolic acid cyclopropanation is essential for viability, drug resistance, and cell wall integrity of Mycobacterium tuberculosis. Chem. Biol. 16, 499–509. https://doi.org/10.1016/j.chembiol.2009.04.001 (2009).
Dkhar, H. K. et al. Mycobacterium tuberculosis keto-mycolic acid and macrophage nuclear receptor TR4 modulate foamy biogenesis in granulomas: A case of a heterologous and noncanonical ligand–receptor pair. J. Immunol. 193, 295–305. https://doi.org/10.4049/jimmunol.1400092 (2014).
Beukes, M. et al. Structure-function relationships of the antigenicity of mycolic acids in tuberculosis patients. Chem. Phys. Lipids 163, 800–808. https://doi.org/10.1016/j.chemphyslip.2010.09.006 (2010).
Song, S. H. et al. Electrospray ionization-tandem mass spectrometry analysis of the mycolic acid profiles for the identification of common clinical isolates of mycobacterial species. J. Microbiol. Methods 77, 165–177. https://doi.org/10.1016/j.mimet.2009.01.023 (2009).
Szewczyk, R., Kowalski, K., Janiszewska-Drobinska, B. & Druszczynska, M. Rapid method for Mycobacterium tuberculosis identification using electrospray ionization tandem mass spectrometry analysis of mycolic acids. Diagn. Microbiol. Infect. Dis. 76, 298–305. https://doi.org/10.1016/j.diagmicrobio.2013.03.025 (2013).
Frankfater, C., Fujiwara, H., Williams, S. J., Minnaard, A. & Hsu, F. F. Characterization of Mycobacterium tuberculosis mycolic acids by multiple-stage linear ion-trap mass spectrometry. J. Am. Soc. Mass Spectrom. 33, 149–159. https://doi.org/10.1021/jasms.1c00310 (2022).
Shui, G. et al. Mycolic acids as diagnostic markers for tuberculosis case detection in humans and drug efficacy in mice. EMBO Mol. Med. 4, 27–37. https://doi.org/10.1002/emmm.201100185 (2012).
Hinz, C., Liggi, S. & Griffin, J. L. The potential of ion mobility mass spectrometry for high-throughput and high-resolution lipidomics. Curr. Opin. Chem. Biol. 42, 42–50. https://doi.org/10.1016/j.cbpa.2017.10.018 (2018).
Dodds, J. N. & Baker, E. S. Ion mobility spectrometry: Fundamental concepts, instrumentation, applications, and the road ahead. J. Am. Soc. Mass Spectrom. 30, 2185–2195. https://doi.org/10.1007/s13361-019-02288-2 (2019).
Kanu, A. B., Dwivedi, P., Tam, M., Matz, L. & Hill, H. H. Jr. Ion mobility-mass spectrometry. J. Mass Spectrom. 43, 1–22. https://doi.org/10.1002/jms.1383 (2008).
Leaptrot, K. L., May, J. C., Dodds, J. N. & McLean, J. A. Ion mobility conformational lipid atlas for high confidence lipidomics. Nat. Commun. 10, 985. https://doi.org/10.1038/s41467-019-08897-5 (2019).
Paglia, G. et al. Ion mobility-derived collision cross section as an additional measure for lipid fingerprinting and identification. Anal. Chem. 87, 1137–1144. https://doi.org/10.1021/ac503715v (2015).
Rose, B. S. et al. High confidence shotgun lipidomics using structurally selective ion mobility-mass spectrometry. Methods Mol. Biol. 2306, 11–37. https://doi.org/10.1007/978-1-0716-1410-5_2 (2021).
Paglia, G., Smith, A. J. & Astarita, G. Ion mobility mass spectrometry in the omics era: Challenges and opportunities for metabolomics and lipidomics. Mass Spectrom. Rev. 41, 722–765. https://doi.org/10.1002/mas.21686 (2022).
Layre, E. et al. A comparative lipidomics platform for chemotaxonomic analysis of Mycobacterium tuberculosis. Chem. Biol. 18, 1537–1549 (2011).
Sartain, M. J., Dick, D. L., Rithner, C. D., Crick, D. C. & Belisle, J. T. Lipidomic analyses of Mycobacterium tuberculosis based on accurate mass measurements and the novel “Mtb LipidDB”. J. Lipid Res. 52, 861–872. https://doi.org/10.1194/jlr.M010363 (2011).
Fahy, E., Sud, M., Cotter, D. & Subramaniam, S. LIPID MAPS online tools for lipid research. Nucleic Acids Res. 35, W606-612. https://doi.org/10.1093/nar/gkm324 (2007).
Ludwig, M. et al. Database-independent molecular formula annotation using Gibbs sampling through ZODIAC. Nat. Mach. Intell. 2, 629–641. https://doi.org/10.1038/s42256-020-00234-6 (2020).
Duhrkop, K. et al. SIRIUS 4: A rapid tool for turning tandem mass spectra into metabolite structure information. Nat. Methods 16, 299–302. https://doi.org/10.1038/s41592-019-0344-8 (2019).
MacLean, B. X. et al. Using skyline to analyze data-containing liquid chromatography, ion mobility spectrometry, and mass spectrometry dimensions. J. Am. Soc. Mass Spectrom. 29, 2182–2188. https://doi.org/10.1007/s13361-018-2028-5 (2018).
Hong, S. et al. Ultralong C100 mycolic acids support the assignment of Segniliparus as a new bacterial genus. PLoS One 7, e39017. https://doi.org/10.1371/journal.pone.0039017 (2012).
Minnikin, D. E. & Brennan, P. J. In Health Consequences of Microbial Interactions with Hydrocarbons, Oils, and Lipids (ed. Goldfine, H.) 1–76 (Springer International Publishing, 2020).
Singh, A. et al. Identification of a desaturase involved in mycolic acid biosynthesis in Mycobacterium smegmatis. PLoS One 11, e0164253. https://doi.org/10.1371/journal.pone.0164253 (2016).
Watanabe, M., Aoyagi, Y., Ridell, M. & Minnikin, D. E. Separation and characterization of individual mycolic acids in representative mycobacteria. Microbiology (Reading) 147, 1825–1837. https://doi.org/10.1099/00221287-147-7-1825 (2001).
Watanabe, M. et al. Location of functional groups in mycobacterial meromycolate chains; the recognition of new structural principles in mycolic acids. Microbiology (Reading) 148, 1881–1902. https://doi.org/10.1099/00221287-148-6-1881 (2002).
Sambandan, D. et al. Keto-mycolic acid-dependent pellicle formation confers tolerance to drug-sensitive Mycobacterium tuberculosis. MBio 4, e0022200213. https://doi.org/10.1128/mBio.00222-13 (2013).
Bottai, D. et al. TbD1 deletion as a driver of the evolutionary success of modern epidemic Mycobacterium tuberculosis lineages. Nat. Commun. 11, 684. https://doi.org/10.1038/s41467-020-14508-5 (2020).
Stow, S. M. et al. An interlaboratory evaluation of drift tube ion mobility-mass spectrometry collision cross section measurements. Anal. Chem. 89, 9048–9055. https://doi.org/10.1021/acs.analchem.7b01729 (2017).
Acknowledgements
G.L-M. received funding from the Medical Research Council MR/W018756/1.
Author information
Authors and Affiliations
Contributions
G.L.M. had full access to all of the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Study concept and design: Y.L., N.K., S.P., G.L.M. Acquisition, analysis, or interpretation of data: All authors. Drafting of the manuscript: Y.L., M.J.S., G.L.M. Critical revision of the manuscript for important intellectual content: All authors.
Corresponding author
Ethics declarations
Competing interests
S.P. and M.J.S. are employees of Agilent technology from which data acquisition was generated. Y.L., N.K. and G.L.-M. do not have conflict of interest to declare.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Y., Kaffah, N., Pandor, S. et al. Ion mobility mass spectrometry for the study of mycobacterial mycolic acids. Sci Rep 13, 10390 (2023). https://doi.org/10.1038/s41598-023-37641-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-37641-9
- Springer Nature Limited