Abstract
Sixty genotypes with different seed coat colour and seed sizes were evaluated for seed longevity under both natural and accelerated ageing over seasons. The genotypes were grown during rabi, 2018, and summer, 2021, and freshly harvested seeds were used. For natural ageing, seeds were stored in a cloth bag in ambient condition and were removed at bimonthly intervals till 20 months. Accelerated ageing was carried out as per International Seed Testing Association (ISTA) guidelines. The germination percentage after natural and accelerated ageing over two seasons was determined. The correlation between two seasons of accelerated ageing and month-wise natural ageing was highly significant. The principal component analysis (PCA) using seed longevity grouped the majority of black genotypes into a separate cluster. Higher seed longevity was associated with black seed coat colour and small seed size. Microsatellite marker-based clustering also produced a separate cluster for majority of black genotypes and grouped the genotypes into a large number of clusters suggesting high diversity in the plant material. Two black seed coat colour genotypes, ACC No.369 and ACC No.39 consistently showed higher longevity under natural and both the years of accelerated ageing and serve as a source of alleles for higher seed longevity in soybean.
Similar content being viewed by others
Introduction
Soybean (Glycine max (L.) Merrill) is one of the major oilseed crops belonging to the family Fabaceae, subfamily Papilionoidea, and genus Glycine. The crop originated in Southeast Asia and was primarily domesticated by China around 1100 BC. Later it was introduced to several countries including India in the first century AD1. Soybean seeds are composed of 20% oil and 40% protein and contribute 70% to world’s protein and 28% to world’s oil consumption2. It is grown in an area of 119.0 million hectares with a production of 339.0 million metric tonnes worldwide and India produces 9.3 million metric tonnes from an area of 11.64 million hectares3. Due to its importance as a staple source of protein and oil for the growing population of the world, resources were allocated in the breeding effort to improve the grain yield with wide adaptability. A large number of varieties with high yield, early maturity, wide adaptability, and suitable for diverse agro-climatic conditions are developed in India and elsewhere4. Soybean seeds are very sensitive to production5,6, storage conditions7,8, seed handling, and mechanical damage9,10,11. The seed embryo and embryonic axis are located under a very thin seed coat and the surface of seed coat has presence of pores12,13. Hence, the seed viability and longevity are low14,15.
Soybean seed longevity is a matter of high concern for farmers, traders, and seed producers14. Seed viability is a serious concern worldwide7,16,17. Soybean seeds deteriorate faster and lose viability during storage18,19,20. Lipid auto-oxidation and subsequent increase of free fatty acid content are the most mentioned reasons for ageing damage of soybean seeds21,22. Remarkable genotypic diversity for a decline in seed longevity during storage has been well documented in soybean23,24,25,26. Seed longevity is a polygenic trait and the identification of genotypes with high seed longevity is very important. Earlier studies indicated that soybean genotypes with dark and hard seed coat27,28,29, smaller seed size30, lesser space between seed coat and cotyledons13, and wild type accessions17,23 possess higher seed longevity as compared to other seed coat colour genotypes. Such soybean genetic resources provide a valuable source for the identification of genotypes suitable for seed longevity improvement in soybean. It will broaden the genetic base for breeding programmes to combine seed longevity with seed yield per plant.
An efficient screening technique to identify the genotypes with higher longevity in a short time to select the lines in breeding is imperative in crop improvement programmes. Natural ageing not only takes longer time17,24,26 but also influenced by storage conditions31,32. Alternatively accelerated ageing technique has been proposed for testing seed germination in general33,34,35 and soybean in particular24,28,36,37,38. The protocol for accelerated ageing has been standardized for soybean39 which may be employed for predicting the viability of soybean seeds.
Genetic diversity information in the source germplasm is important to develop new cultivars with seed longevity. Simple sequence repeats (SSR) markers which are ubiquitously distributed within genomes are used to identify genetic diversity between genotypes in a germplasm collection40,41,42. The SSR markers have been applied in various aspects of molecular research such as genetic diversity assessment, fingerprinting, marker assisted selection, gene flow, and genetic linkage mapping43.
The identification and development of high seed longevity genotypes in soybean is a requirement to be used as a source of germplasm for seed longevity in breeding. With this background, the present investigation was undertaken to elucidate the information on genetic and molecular diversity among the selected germplasm and identify genotypes with higher seed longevity through accelerated and natural ageing techniques and their association with seed coat colour and seed size.
Materials and methods
Plant material
Sixty diverse genotypes including 46 yellow, three green (104-31, JS 90-41, and BNS-5), and nine black seed coat colour (Pune 14, Pune 30, Kalitur, ACC Nos. 37, 369, 39, 101, 109, and LB-5) were collected from author affiliated government institute, viz., All India Co-ordinated Research Project (AICRP)-Soybean scheme, Zonal Agricultural Research Station, GKVK, Bangalore, for the study and no permission is required for the collection. The genotypes, their name, and seed characteristics are given in Supplementary Table S1 and can be obtained by contacting the first author. The genotypes also differed for seed size, growth habit, and other quantitative traits related to crop growth and seed yield. The genotypes were grown and multiplied before the start of the experiment at AICRP-National Seed Project, University of Agricultural Sciences, Bangalore, during Rabi, 2018, and in Summer, 2021. Harvesting was done during January, 2019 and April, 2021, respectively for Rabi, 2018 and Summer, 2021. The seeds were immediately dried to a safe moisture content of 9% after manual threshing and the germination percent of the seeds were recorded before the start of the seed storage experiment.
Seed longevity
The seeds obtained from Rabi, 2018 experiment were tested for seed longevity under both natural and accelerated ageing, while the seeds harvested in Summer, 2021 were tested for longevity using accelerated ageing at AICRP-National Seed Project, University of Agricultural Sciences, Bangalore.
Natural ageing
Freshly harvested seeds were dried uniformly to 9% moisture content and stored in a cloth bag from February, 2019 to September, 2020 under ambient conditions (25–30 °C and 60–65% RH). At bimonthly intervals from the 8th month onwards till the 20th month after harvest, the seed samples were drawn from the stored cloth bags and used for testing the germination. The germination was recorded till the 20th month after harvest. The laboratory germination test was carried out as per the International Seed Testing Association (ISTA) guidelines44 using between paper method. One hundred seeds in four replicates were randomly drawn from the cloth bag for each genotype at bimonthly intervals and kept for germination following between paper method. The seeds were kept in an incubator and the constant temperature of 30 °C and relative humidity of 90 ± 2% were maintained. The seeds were allowed to germinate and grow for 5 days. On the 5th day, the germination papers were removed from the incubator and the germination count was recorded following ISTA guidelines and expressed in percentage.
Accelerated ageing
Freshly harvested seeds after drying to 9% moisture content were used for the study in both seasons. Forty-two gram seeds for each genotype were placed in an ageing box with a wire mesh screen by following the guidelines specified by ISTA39 for soybean. Four varieties, DSB 32, JS 95-60, Hardee, and DSB 33, were used as checks for the experiment. The box is filled with 40 ml of distilled water and it was sealed all around to maintain more than 95% relative humidity. The box was kept in an ageing chamber which maintain a constant temperature of 41 ± 0.3 °C for 72 h. After 72 h, the treated seeds were removed from the box, weighted, and kept for germination as mentioned above in four replications immediately (within an hour). The experiment was replicated twice.
Genetic diversity analysis
DNA isolation
The genotypes were grown in paper cups and the genomic DNA was isolated from young leaf tissues using cetyl trimethylammonium bromide (CTAB) method45. The quality and quantity of DNA were estimated using 0.8% agarose gel electrophoresis with ethidium bromide staining and the DNA was uniformity diluted to 50 ng μl−1 and stored.
Microsatellite marker genotyping
Primers
Fifteen microsatellite primer pairs were chosen for the study. A list of selected primers, primer sequence, and standardized annealing temperatures are given in Supplementary Table S2.
Genotyping
The isolated DNA was subjected to a polymerase chain reaction (PCR) using a reaction mixture of 10 μl and 38 reaction cycles. Three percent agarose gel with 0.5 μg ml−1 ethidium bromide was used to visualize the marker alleles along with a 1000 bp ladder to score the fragment size.
Statistical analysis
Seed longevity
The germination percentage values were transformed using arcsine transformation to stabilize the variance. The analysis of variance (ANOVA) for the observed germination values for all the months and accelerated ageing was done by following a completely randomized design using SPSS software. PAST 4.03 software was used for plotting box plots and principal component analysis (PCA). Box plot was developed using the range, median, first, and third quartile values. PCA was undertaken based on scores of germination values from different storage months of natural ageing and accelerated ageing over two seasons. Eigen values and principal components were estimated using the germination values based on non-rotated loadings.
The correlation between two seasons accelerated ageing data and month-wise natural ageing data was determined using Pearson’s correlation coefficient. Further, the genotypes were classified into different longevity groups46 based on their germination values in natural and accelerated ageing as given below,
Classes of genotype | Seed longevity rank (SR) |
|---|---|
\(\left(\mu +2\sigma \right) \,and\, above\) | SR 1 |
\((\mu +\sigma )\text{ to }\left(\mu +2\sigma \right)\) | SR 2 |
\((\mu -\sigma )\text{ to }\left(\mu +\sigma \right)\) | SR 3 |
\((\mu -2\sigma )\text{ to }\left(\mu -\sigma \right)\) | SR 4 |
\(\left(\mu -2\sigma \right)\, and\, less\) | SR 5 |
Here, ‘μ’ and ‘σ’ are the mean and standard deviation of the overall germination of genotypes, respectively.
The seed longevity ranks ‘SR 1’ and ‘SR 5’ reveals “highest” and “lowest” seed longevity, respectively. The genotypes which showed consistently higher ranking in both natural and accelerated ageing were considered as genotypes with higher longevity.
Molecular data analysis
Marker scoring was done manually as 0/1 matrix. The presence of an allele was denoted as ‘1’ and absence as ‘0’. The band size matrix was used for the microsatellite genotypic data analysis. Allele frequency, genetic diversity and Polymorphism Information Content (PIC) were estimated using PowerMarker v 3.25 software. A number of different alleles (Na), number of effective alleles (Ne), Shannon’s information index (I), observed heterozygosity (Ho), expected heterozygosity (He), unbiased expected heterozygosity (uHe), and fixation index (F) was computed in GeneAlEx v 6.547. Dissimilarity index was calculated using Euclidean distance with 1000 bootstraps and with this, hierarchical clustering of genotypes was done based on the unweighted pair group method with arithmetic mean (UPGMA) in DARwin 5.0 software.
Results
Seed longevity
The longevity of a seed is measured using its germination ability after a period of dry storage. A reduction in germination percentage of the genotypes was observed throughout natural storage under ambient conditions. The average germination % of genotypes after 8 months of ambient storage was 82.34 and it gradually declined to 3.35% after 20 months (Table 1). By 14th month, CAT-44 and 104-31 genotypes lost their viability completely, whereas, ACC Nos.369, 39, and 101 maintained 100% germination. The germination of genotypes after 20 months of storage was ranging from 0.00 to 54.54%. The accelerated ageing also significantly reduced the germination of genotypes and the mean germination across genotypes was 48.16% in 2019, and 44.92% in 2021.
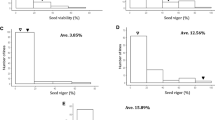
The analysis of variance suggested highly significant variation among genotypes for germination after ageing (Table 2). The box and jitter plot drawn for germination indicated high diversity among soybean genotypes for seed longevity (Fig. 1). The genotypes in the 12th, 14th, 16th and 18th month as well as accelerated ageing in 2019 were more varied toward the positive quartile, on the other hand, in 8th and 10th month of natural ageing and accelerated ageing in 2021, the variation between the genotypes were distributed towards first and third quartiles. In all the ageing methods, the black seed coat colour genotypes (indicated as black dots in the box plot) had comparatively higher seed longevity along with a few yellow and one green colour genotypes (BNS 5), which are depicted as yellow and green colour dots, respectively.
Principal component analysis (PCA)
PCA is an alternative method of population structure analysis and it gives the explicit pattern of combination between the genotypes in the factor plane. In the PCA analysis of 60 soybean genotypes, the eigenvalues were higher for components one and two explaining 66.046 and 10.761% variance of the population, respectively (Table 3, Fig. 2). A scatter plot was made based on the loadings of each genotype in principal components one and two (Fig. 3). Based on the similarity of the genotypes for the seed longevity, different clusters were formed in the scatter plot. ACC No. 369 genotype had a higher loading value and was found to have higher longevity. Other black genotypes such as, ACC Nos. 37, 101, 39, 109, Kalitur, and LB-5 were closely formed in a single cluster with higher scores in component 1. The genotypes with higher seed longevity are spotted in quadrants one and four, having the majority of the black seed coat colour genotypes. One green colour genotype (BNS-5) and a few yellow seed coat colour genotypes such as, MAUS-71, MACS 1410, and RKS-18 were also placed in the same quadrant along with black seeded genotypes. A wide cluster was formed in the second and third quadrant including a majority of the yellow seeded genotypes along with two green (104-31 and JS 90-41) and two black (Pune 14 and Pune 30) seed coat colour genotypes.
Association between natural and accelerated ageing and identification of genotypes with higher longevity
Pearson’s correlation was performed to study the association between natural and accelerated ageing methods to test the reliability of accelerated ageing for screening the soybean genotypes for seed longevity. A significant positive correlation was observed between germination after natural ageing with germination after accelerated ageing in both the years (Fig. 4). The correlation was significant at 8th and 10th month of natural storage, on the other hand, it was found to be highly significant after 12, 14, 16, 18, and 20 months of natural ageing with accelerated ageing in both the years. During the year 2019, the highest correlation was observed after 20th month of natural ageing (0.506) followed by 16th month of natural ageing (0.496). Similarly, during the year 2021, a higher correlation was found in 12th month (0.433) followed by 14th (0.432), and 16th (0.429) months.
The genotypes were grouped into five longevity ranks using the empirical rule which is also called as three sigma rule. The number and list of genotypes in each of the seed longevity classes are given in Table 4. The highest number of genotypes fell into SR 3 rank in both natural and accelerated ageing. More than 40 genotypes were grouped in SR 3 rank in all the months of testing under natural ageing. Forty two genotypes in accelerated ageing during the year 2019 and 38 genotypes during 2021 were observed in SR 3. None of the genotypes were found in SR 5 rank except 104-31 and KHSB 2 genotype in natural ageing after 10 months. The SR 1 rank consists of genotypes with the highest longevity. Two genotypes, ACC Nos. 39 and 369 were consistently found in SR 1 rank in most of the months of natural ageing. Under accelerated ageing also, the genotype ACC No. 369 in 2019, and the genotype ACC No. 39 in 2021 were found in SR 1 rank. Both of them are black seed coat colour genotypes. Genotypes viz., ACC No. 37, 109, LB-5, MAUS 71, and RKS 18 were consistently found in SR 2 rank of seed longevity. SR 1 and SR 2 rank classes are considered better for identifying genotypes for higher seed longevity. All the black seed coat genotypes, i.e., ACC Nos. 369, 39, 37, 109, and LB-5 were consistently found under either SR 1 or SR 2 rank. It is interesting to note that the seed size of the black seed coat colour genotypes which were grouped in higher longevity was small compared to other black seeded genotypes (Pune 14 and Pune 30) which were grouped in lower seed longevity rank (SR 3) (Supplementary Table S1). None of the green seed coat colour genotypes were consistently found in higher longevity rank classes. SR 1 rank has only consisted of black seed coat colour genotypes and none of the yellow colour genotypes were found in that ground except MAUS 71 in 8th month of natural storage. Even in SR 2 class, no single yellow seed coat colour genotypes was found constant in all the natural ageing months and accelerated ageing method of testing for seed longevity.
Microsatellite marker diversity of genotypes
Genetic analysis of the sixty soybean genotypes was done using fifteen microsatellite markers. The presence and absence of alleles were scored as ‘1’ and ‘0’, respectively (Fig. 5, Supplementary Fig. S1). Totally, 38 alleles were observed from all the loci studied with a range of 1 to 5 and a mean of 2.533 per locus. The highest number of alleles per loci was observed for the marker BSOY 43 (5). The percentage of polymorphic loci among the genotypes was 93.33. Three alleles had allele frequency of less than 5% which are considered as rare. The mean frequency of minor and major alleles was 0.153 and 0.692, respectively (Table 5). Major allele frequency ranged from 0.350 to 1.000 and the minor allele frequency ranged from 0.000 to 0.417. BSOY 23 loci was monomorphic for the studied genotypes. The 60 genotypes were genetically diverse with a mean genetic diversity of 0.414. The genotypes were highly diverse for the BSOY 43 loci with a diversity of 0.769. The PIC value of the primers ranged from 0.000 to 0.735 with a mean of 0.359. BSOY 43 had the highest PIC value (0.735) as well as a greater number of alleles. A highly positive correlation of r = 0.861 (p < 0.00001) was found between number of alleles and PIC value (Fig. 6).
Banding pattern of BSOY 29 marker alleles in 60 genotypes (L Ladder) (The gel with serial number 49–60 was cropped from different gel and the full-length image is included in Supplementary Fig. S1).
The average number of effective alleles, Shannon’s information index, observed heterozygosity, expected heterozygosity and unbiased heterozygosity of studied soybean genotypes were 1.949, 0.679, 0.136, 0.414, and 0.418 respectively (Table 5). Four out of 15 loci had multiple alleles in all the genotypes, with observed heterozygosity of 0.133, 1.00, 0.017 and 0.883, respectively for primers BSOY 34, BSOY 43, BSOY 47 and BSOY 45. The Shannon information index was higher for BSOY 43 marker with a value of 1.524 followed by BSOY 45 (0.948) predicting high diversity of the population and it is highly correlated with the genetic diversity of the population with a correlation coefficient of 0.962 (Fig. 6).
The UPGMA based hierarchical clustering of the soybean genotypes based on the dissimilarity index grouped 30 genotypes into ten clusters with more than one genotype per cluster and the remaining 30 genotypes formed solitary clusters with only one genotype each (Fig. 7). One cluster with 6 genotypes solely consisted of black seed coat genotypes. Among the remaining three black seed coat colour genotypes, LB-5 and Pune 14 formed independent solitary clusters and Pune 30 was solitary in a sub-cluster. Three green seeded genotypes were grouped into different clusters, The yellow seed coat colour genotypes were also grouped in to different clusters and sub clusters.
Discussion
The main purpose of this experiment in soybean is to identify the superior genotypes for seed longevity. Given the difficulty and time required for ageing under natural environmental conditions to assess the seed longevity of a genotype, accelerated ageing was proposed to test the longevity of a genotype or a segregant37. In soybean, an increase in seed yield combined with seed longevity in a single genotype is valuable for a breeder. Therefore, the identification of donors for seed longevity in soybean is more meaningful. Attempts have been made in the past to understand the genetic diversity for seed longevity in soybean17,36,38 and its influence on various seed characteristics12,13,30,48. Resources have also been allocated to understand the genetics of this complex polygenic trait26,28,49. However, the genetic resources identified for seed longevity for use in crop improvement programme are scanty.
This calls for screening genotypes for seed longevity and 60 genotypes representing different seed coat colour, seed size, and growth habit were evaluated for seed longevity using natural and accelerated ageing techniques. The germination percentage after ageing treatment has been frequently followed to estimate seed longevity50,51. The present study confirmed significant variability existing in tested soybean genotypes for seed longevity during seed dry storage. A similar observation was reported by Hosamani et al.24 in soybean, wherein, the viability of black genotypes had higher longevity than yellow seeded genotypes under natural and artificial ageing methods. Natural ageing for 3 years in three soybean species including cultivated and wild accessions exhibits significant variation for seed viability reduction between species, and wild accessions recorded higher seed longevity17.
Another objective of this study is a classification of diverse soybean genotypes with different seed coat colour and seed size based on their longevity which was tested after natural and accelerated ageing. In the PCA scatter diagram using seed germination values after both ageing, 11 genotypes with higher seed longevity were found in the first and fourth quadrants with higher loadings. Among them, seven were black, three with yellow, and one was green (BNS-5) seed coat colour genotypes. The PCA scatter plot has been used to identify stress tolerant, high productivity genotypes in different crops52,53. The genotypes were also grouped based on molecular markers. The molecular marker system has become an essential part of the genetic diversity analysis and for crop improvement programme54. The discriminatory power of a marker is well explained by PIC value by using the number of alleles and their frequencies55. Among the studied microsatellite markers, BSOY 43 was highly polymorphic having higher number of alleles and PIC value. PIC values above 0.5 are said to be consistent with the usefulness of the marker56. The informativeness of the markers on genetic diversity was being estimated using PIC value by many researchers55,57,58 and made a positive correlation with number of alleles. The high percentage of polymorphic loci among the genotypes suggests that the genotypes used for the study were diverse57. Further, the rare alleles with the occurrence of less than 5% allele frequency are also observed, hence, the genotypes used in the study represent both recently cultivated genotypes as well as rare genotypes55. A positive correlation between genetic diversity, PIC, and Shannon’s information index was also observed in this study. The Shannon information index is a measure of the degree of uncertainty in predicting the species of a sample, which is related to the diversity of a population59.
The molecular marker clustering also supports the results observed in PCA that the genotypes used for the present study are highly diverse at the genome level also. Like in PCA, one of the clusters formed using molecular markers contain majority of black seed coat colour genotypes. The clustering of genotypes using genotypic and phenotypic data are largely in agreement. The observations are in line with the finding made by Jeong et al.60 and Kachare et al.57 wherein, the clusters obtained using phenotypic diversity was in accordance with the cluster found after performing SSR marker diversity.
The major objective of the present study is to identify the genotypes with higher seed longevity. After different duration of ageing of seeds under natural conditions and accelerated ageing, the seeds were tested for viability using germination percent. A significant positive correlation was found between natural ageing after 8th month till 20th month of storage under ambient conditions and the accelerated ageing method. Hosamani et al.24 and Matera et al.38 also studied the association between natural and accelerated ageing in soybean and reported highly significant correlation between them in detecting seed longevity. The method has been widely used in crops like rice61, wheat62, chickpea33, Brassica63, soybean26,28,36, and palm64 for seed longevity. Accelerated ageing method is based on the principle of seed survival curve, that explains the relationship between seed viability and storage period. Accelerated ageing reduces the lag phase of a seed in declining its viability during storage37 by subjecting the seed to conditions that hasten the deterioration process. Such artificial ageing method is reliable, time saving, and useful for the quick assessment of segregants or genotypes in breeding. In the present study we have adapted both natural and accelerated ageing techniques to identify genotypes with higher seed longevity.
Based on germination after natural and accelerated ageing methods, the genotypes were classified into different longevity rank groups. Majority of the genotypes were found in low seed longevity rank group of SR 3 and SR 4. Only a few genotypes were consistently found in the highest seed longevity group of SR 1. The two genotypes, i.e., ACC No. 369 and ACC No. 39 were frequently found in SR 1 group in both ageing methods and hence, the highest seed longevity among the genotypes. Both of them were found to be black seed coat colour genotypes. The black seed coat soybean genotypes were found to have relatively higher longevity13,28,65. In addition to these two black seed coat genotypes, other black seed genotypes, ACC No. 37, ACC No. 109, and LB-5 were found either in SR 1 or in SR 2 group. Seed longevity was also found to be associated with seed coat colour. The seed coat colour association with seed longevity observed in this study is in harmony with the earlier findings13,17,24,28,65. Kuchlan et al.13 reported that, black seeded genotypes have a minor gap between the seed coat and cotyledon, lesser pores on the surface of the seed coat, and greater lignin content in the seed coat, which makes them less susceptible to mechanical damage and deteriorative changes during ageing, than other seed coat colours. All the black seeded genotypes with high seed longevity had small seed size and lower 100 seed weight. The 100 seed weight ranged from 9.95 to 13.07 g. The other two black genotypes, Pune 14 and Pune 30 which did not show higher longevity had bigger seed size (15.20 and 17.10, respectively). It can be concluded that along with black seed coat colour, seed size are the determining factors for seed longevity in soybean30.
Finally, in the study, contrasting accessions have been identified, which can be used for additional experiments to determine QTLs controlling the trait and also for breeding commercial lines. For this objective, the contrasting, yellow genotype with low longevity and black genotype with higher longevity have been crossed, and the true F1 plants have been identified using molecular and morphological markers and the F2 generation is being grown to develop RILs suitable for mapping seed longevity-related traits.
Data availability
The datasets generated during and/or analysed during the current study are available from the BioStudies repository of EMBL-EBI. The Accession Number is S-BSST814. The below link can be used to access the data https://www.ebi.ac.uk/biostudies/studies/S-BSST814?key=e287dfc1-07ae-4491-b44f-25d6f75cf80c.
References
Singh, B. B. Success of soybean in India: The early challenges and pioneer promoters. Asian Agric. Hist. 10(1), 43–53 (2006).
Jo, H. et al. Genetic diversity of soybeans (Glycine max (L.) Merr.) with black seed coats and green cotyledons in Korean germplasm. Agronomy 11, 581. https://doi.org/10.3390/agronomy11030581 (2021).
SoyStats®. http://soystats.com (2021).
Carter, T. E. J. R., Nelson, R. L., Sneller, C. H. & Cui, Z. Genetic diversity in soybean. In Soybeans: Improvement, Production, and Uses, Agronomy Monograph No. 3rd edn, Vol. 16 (American Soc Agronomy Crop Sci Soc America, 2004).
Bhatia, V. S. Seed longevity as affected by field weathering and its association with seed coat and pod characters in soybean. Seed Res. 24, 82–87 (1996).
Gillen, A. M., Smith, J. R., Mengistu, A. & Bellaloui, N. Effects of maturity and Phomopsis longicolla on germination and vigor of soybean seed of near-isogenic lines. Crop Sci. 52, 2757–2766. https://doi.org/10.2135/cropsci2011.10.0566 (2012).
Shelar, V. R., Shaikh, R. S. & Nikam, A. S. Soybean seed quality during storage: A review. Agric. Rev. 29(2), 125–131 (2008).
Nik, S. M. M., Tilebeni, H. G., Zeinali, E. & Tavassoli, A. Effects of seed ageing on heterotrophic seedling growth in cotton. Am. Eurasian J. Agric. Environ. Sci. 10(4), 653–657 (2011).
Shelar, V. R. Effect of mechanical damage on physiological and biochemical deterioration of soybean (Glycine max) seeds during storage. Ph.D. Thesis, MPKV, Rahuri (2002).
Shelar, V. R. Role of mechanical damage in deterioration of soybean seed quality during storage—A review. Agric. Rev. 29(3), 177–184 (2008).
Zihad, M. Physiological phenotyping of soybean (Glycine max L. Merrill) genotypes for seed longevity. M.Sc. Thesis. (Division of Seed Science and Technology, ICAR-IARI, 2013)
Shao, S., Meyer, C. J., Ma, F., Peterson, C. A. & Bernards, M. A. The outermost cuticle of soybean seeds: Chemical composition and function during imbibition. J. Exp. Bot. 58(5), 1071–1082. https://doi.org/10.1093/jxb/erl268 (2007).
Kuchlan, M. K., Dadlani, M. & Samuel, V. K. Seeds coat properties and longevity of soybean seeds. J. New Seeds 11, 239–249. https://doi.org/10.1080/1522886X.2010.497960 (2010).
Delouche, J. C. Soybean seed storage beyond one year. In Proc. 7th Soybean Res. Conf. ASTA 60–73 (1977).
De Souza, F. H. D. & Filho, J. M. The seed coat as a modulator of seed-environment relationships in Fabaceae. Rev. Braz. Bot. 24(4), 365–375 (2001).
Annonymous. Considerations for Making Carry-Over Soybean Seed Decisions. https://www.canr.msu.edu/news/considerations-for-making-carry-over-soybean-seed-decisions (Michigan State University Extension, 2019).
Chandra, S. et al. Seed longevity studies in wild type, cultivated and inter-specific recombination inbred lines (RILs) of soybean. Genet. Resour. Crop Evol. https://doi.org/10.1007/s10722-021-01240-2 (2021).
Brumm, T. J., Hurbugh, C. R. Jr. & Johnson, L. A. Cracking and dehulling shriveled and wrinkled soybeans. J. Am. Oil Chem. Soc. 67(11), 750–756 (1990).
Ha, B. K., Kim, H. K. & Kang, S. T. Mapping QTLs with epistatic effects and QTL-by-environment interactions for seed coat cracking in soybeans. Euphytica 186, 933–942. https://doi.org/10.1007/s10681-012-0719-8 (2012).
Bellaloui, N., Smith, J. R. & Mengistu, A. Seed nutrition and quality, seed coat boron and lignin are influenced by delayed harvest in exotically-derived soybean breeding lines under high heat. Front. Plant Sci. 8, 1563. https://doi.org/10.3389/fpls.2017.01563 (2017).
Sharma, S., Kaur, A., Bansal, A. & Gill, B. S. Positional effects of soybean seed composition during storage. J. Food Sci. Technol. 50, 353–359. https://doi.org/10.1007/s13197-011-0341-0 (2013).
Indiarto, R. & Qonit, M. A. H. A review of soybean oil lipid oxidation and its prevention techniques. Int. J. Adv. Sci. Technol. 29(6), 5030–5037 (2020).
Zhou, S., Sekizaki, H., Yang, Z., Sawa, S. & Pan, J. Phenolics in the seed coat of wild soybean (Glycine soja) and their significance for seed hardness and seed germination. J. Agric. Food Chem. 58(20), 10972–10978. https://doi.org/10.1021/jf102694k (2010).
Hosamani, J., Kumar, M. B., Talukdar, A., Lal, S. K. & Dadlani, M. Molecular characterization and identification of candidate markers for seed longevity in soybean [Glycine max (L.) Merill]. Indian J. Genet. 73(1), 64–71. https://doi.org/10.5958/j.0019-5200.73.1.009 (2013).
Chandra, S. et al. Seed coat permeability studies in wild and cultivated species of soybean. Int. J. Curr. Microbiol. Appl. Sci. 6(7), 2358–2363. https://doi.org/10.20546/ijcmas.2017.607.279 (2017).
Zhang, X. et al. Whole-genome mapping identified novel “QTL hotspots regions” for seed storability in soybean (Glycine max L.). BMC Genom. 20, 499. https://doi.org/10.1186/s12864-019-5897-5 (2019).
Liu, J. et al. Metabolism variation and better storability of dark-versus light-coloured soybean (Glycine max L. Merr.) seeds. Food Chem. 223, 104–113. https://doi.org/10.1016/j.foodchem.2016.12.036 (2017).
Adsul, A. T., Chimote, V. P. & Deshmukh, A. P. Inheritance of seed longevity and its association with other seed-related traits in soybean (Glycine max). Agric. Res. https://doi.org/10.1007/s40003-018-0297-7 (2018).
Kumar, A. et al. Genetic studies on seed coat permeability and viability in RILs derived from an inter-specific cross of soybean [Glycine max (L.) Merrill]. Indian J. Genet. 79(1), 48–55. https://doi.org/10.31742/IJGPB.79.1.7 (2019).
Hosamani, J., Dadlani, M., Santha, I. M., Kumar, M. B. A. & Jacob, S. R. Biochemical phenotyping of soybean (Glycine max (L) Merrill) genotypes to establish the role of lipid peroxidation and antioxidant enzymes in seed longevity. Agric. Res. 2(2), 119–126. https://doi.org/10.1007/s40003-013-0058-6 (2013).
Selvi, D. T. & Saraswathy, S. Seed viability, seed deterioration and seed quality improvements in stored onion seeds: A review. J. Hortic. Sci. Biotechnol. https://doi.org/10.1080/14620316.2017.1343103 (2017).
Wang, W. et al. The effect of storage conditions and duration on the deterioration of primed rice seeds. Front. Plant Sci. 9, 172. https://doi.org/10.3389/fpls.2018.00172 (2018).
Kapoor, N., Arya, A., Siddiqui, M. A., Amir, A. & Kumar, H. Seed deterioration in chickpea (Cicer arietinum L.) under accelerated aging. Asian J. Plant Sci. 9(3), 158–162 (2010).
Fenollosa, E., Jene, L. & Bosch, S. M. A rapid and sensitive method to assess seed longevity through accelerated ageing in an invasive plant species. Plant Methods 16, 64–75. https://doi.org/10.1186/s13007-020-00607-3 (2020).
Zhang, J., Xiang, S. & Wan, H. Negative association between seed dormancy and seed longevity in bread wheat. Am. J. Plant Sci. 12, 347–365. https://doi.org/10.4236/ajps.2021.123022 (2021).
Patil, S. S., Doddagoudar, S. R., Kurunalliker, V. J., Mathad, R. C. & Patil, R. P. Prediction of storability in soybean seeds through accelerated ageing technique (Glycine max (L.) Merrill). Legume Res. 41(4), 572–577. https://doi.org/10.18805/LR-3760 (2018).
Hay, F. R., Valdez, R., Lee, J. S. & Cruz, P. C. S. Seed longevity phenotyping: Recommendations on research methodology. J. Exp. Bot. 70(2), 425–434. https://doi.org/10.1093/jxb/ery358 (2019).
Matera, T. C. et al. Accelerated aging test and its relationship to physiological potential of soybean seeds. J. Seed Sci. 41(3), 301–308. https://doi.org/10.1590/2317-1545v41n3212746 (2019).
ISTA. Accelerated Ageing Method for Soybean (International Seed Testing Association, 2010).
Kaga, A. et al. Evaluation of soybean germplasm conserved in NIAS genebank and development of mini core collections. Breed. Sci. 61, 566–592. https://doi.org/10.1270/jsbbs.61.566 (2012).
Nachimuthu, V. V. et al. Analysis of population structure and genetic diversity in rice germplasm using SSR markers: An initiative towards association mapping of agronomic traits in Oryza sativa. Rice 8, 30. https://doi.org/10.1186/s12284-015-0062-5 (2015).
Zhao, H. et al. The genetic diversity and geographic differentiation of the wild soybean in Northeast China based on nuclear microsatellite variation. Int. J. Genet. https://doi.org/10.1155/2018/8561458 (2018).
Powell, W., Machray, G. C. & Provan, J. Polymorphism revealed by simple sequence repeats. Trends Plant Sci. 1, 215–222. https://doi.org/10.1016/1360-1385(96)86898-1 (1996).
Anonymous. International Rules for Seed Testing. ISTA (International Seed Testing Association), International rules for seed testing. (Bassersdorf, Switzerland, 2013).
Doyle, J. J. & Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12, 13–15 (1990).
Wheeler, D. J. & Chambers, D. S. Understanding Statistical Process Control 2nd edn. (SPC Press, 1992).
Peakall, R. & Smouse, P. E. GenALEx6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 28(19), 2537–2539. https://doi.org/10.1111/j.1471-8286.2005.01155.x (2012).
Kang, B. K. et al. High-density linkage map reveals QTL for type-1 seed coat cracking in RIL population of soybean (Glycine max (L.) Merr.). Euphytica 216, 153. https://doi.org/10.1007/s10681-020-02684-w (2020).
Renald, J. et al. Identification of novel seed longevity genes related to oxidative stress and seed coat by genome-wide association studies and reverse genetics. Plant Cell Environ. 43, 2523–2539. https://doi.org/10.1111/pce.13822 (2020).
Righetti, K. et al. Inference of longevity-related genes from a robust co-expression network of seed maturation identifies regulators linking seed storability to biotic defense-related pathways. Plant Cell 27, 2692–2708. https://doi.org/10.1105/tpc.15.00632 (2015).
Vidigal, D. S. et al. Galactinol as marker for seed longevity. Plant Sci. 246, 112–118. https://doi.org/10.1016/j.plantsci.2016.02.015 (2016).
Dutta, S. S. et al. Marker-trait association for low-light intensity tolerance in rice genotypes from Eastern India. Mol. Genet. Genom. 293, 1493–1506. https://doi.org/10.1007/s00438-018-1478-6 (2018).
Aleem, M. et al. Comprehensive RNA-seq analysis revealed molecular pathways and genes associated with drought tolerance in wild soybean (Glycine soja Sieb. And Zucc). Physiol. Plant. 172, 707–732. https://doi.org/10.1111/ppl.13219 (2020).
Rao, V. R. & Hodgkin, T. Genetic diversity and conservation and utilization of plant genetic resources. Plant Cell Tissue Organ Cult. 68, 1–19 (2002).
Guzman, F. A., Moore, S., de Vicente, M. D. & Jahn, M. M. Microsatellites to enhance characterization, conservation and breeding value of Capsicum germplasm. Genet. Resour. Crop Evol. 67, 569–585. https://doi.org/10.1007/s10722-019-00801-w (2020).
Botstein, D., White, R. L., Skolnick, M. & Davis, R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32(3), 314–331 (1980).
Kachare, S., Tiwari, S., Tripathi, N. & Takur, V. V. Assessment of genetic diversity of soybean (Glycine max) genotypes using qualitative traits and microsatellite markers. Agric. Res. 9(1), 23–34. https://doi.org/10.1007/s40003-019-00412-y (2020).
Abebe, A. T. et al. Assessment of diversity in tropical soybean (Glycine max (L.) Merr.) varieties and elite breeding lines using single nucleotide polymorphism markers. Plant Genet. Resour. Charact. Util. 19(1), 20–28. https://doi.org/10.1017/S1479262121000034 (2021).
Barnes, B. V., Zak, D. R., Denlon, S. R. & Spurr, S. H. Forest Ecology 4th edn. (Wiley, 1998).
Jeong, N. et al. Korean soybean core collection: Genotypic and phenotypic diversity population structure and genome-wide association study. PLoS ONE 14(10), e0224074. https://doi.org/10.1371/journal.pone.0224074 (2019).
Lin, Q. et al. Genetic dissection of seed storability using two different populations with a same parent rice cultivar N22. Breed. Sci. 65, 411–419. https://doi.org/10.1270/jsbbs.65.411 (2015).
Arif, M. A. R. et al. Genetic studies of seed longevity in hexaploid wheat using segregation and association mapping approaches. Euphytica 186, 1–13. https://doi.org/10.1007/s10681-011-0471-5 (2012).
Nagel, M. et al. Seed longevity in oilseed rape (Brassica napus L.)—Genetic variation and QTL mapping. Plant Genet. Resour. 9, 260–263. https://doi.org/10.1017/S1479262111000372 (2011).
Barreto, L. C. & Gracia, Q. S. Accelerated ageing and subsequent imbibition affect seed viability and the efficiency of antioxidant system in macaw palm seeds. Acta Physiol. Plant. 39, 72. https://doi.org/10.1007/s11738-017-2367-z (2017).
Pawar, P. V., Naik, R. M., Deshmukh, M. P., Satbhai, R. D. & Mohite, S. G. Biochemical and molecular marker-based screening of seed longevity in soybean (Glycine max (L.) Merrill). Legume Res. https://doi.org/10.18805/LR-3915 (2017).
Acknowledgements
The authors would like to extend thanks to Department of Science and Technology, Government of India, for the INSPIRE fellowship to Naflath, All India Co-ordinated Research Project (AICRP) on Soybean, University of Agricultural Sciences, Bangalore, for providing soybean germplasm and AICRP-National Seed Project, DST-FIST funded Department of Plant Biotechnology, University of Agricultural Sciences, Bangalore, Karnataka, India, for extending the facilities to carry-out the work.
Author information
Authors and Affiliations
Contributions
Conception or design of the work (R.S., R.L.R, and N.T.V.), development of experimental material, data collection (N.T.V.), data analysis and interpretation R.L R., N.T.V.), Drafting of the article (N.T.V.), Critical revision of the article (R.L.R.).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
T. V., N., S., R. & R. L., R. Evaluation of diverse soybean genotypes for seed longevity and its association with seed coat colour. Sci Rep 13, 4313 (2023). https://doi.org/10.1038/s41598-023-31071-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31071-3
- Springer Nature Limited