Abstract
microRNAs (miRNAs) are endogenous small RNAs that are key regulatory factors participating in various biological activities such as the signaling of phosphorus deficiency in the plant. Previous studies have shown that miR156 expression was modulated by phosphorus starvation in Arabidopsis and soybean. However, it is not clear whether the over-expression of soybean miR156b (GmmiR156b) can improve a plant’s tolerance to phosphorus deficiency and affect yield component traits. In this study, we generated Arabidopsis transgenic lines overexpressing GmmiR156b and investigated the plant’s response to phosphorus deficiency. Compared with the wild type, the transgenic Arabidopsis seedlings had longer primary roots and higher phosphorus contents in roots under phosphorus-deficit conditions, but lower fresh weight root/shoot ratios under either phosphorus-deficient or sufficient conditions. Moreover, the GmmiR156b overexpression transgenic lines had higher phosphorus content in shoots of adult plants and grew better than the wide type under phosphorus-deficient conditions, and exhibited increased seed yields as well as strong pleiotropic developmental morphology such as dwarfness, prolonged growth period, bushy shoot/branching, and shorter silique length, suggesting that the transgenic lines were more tolerant to phosphorus deficiency. In addition, the expression level of four SQUAMOSA PROMOTER BINDING PROTEIN LIKE (SPL) genes (i.e., AtSPL4/5/6/15) were markedly suppressed in transgenic plants, indicating that they were the main targets negatively regulated by GmmiR156b (especially AtSPL15) and that the enhanced tolerance to phosphorus deficiency and seed yield is conferred mainly by the miR156-mediated downregulation of AtSPL15.
Similar content being viewed by others
Introduction
Phosphorus (P) is an essential major nutrient required for plant growth and crop production1. It is a constituent element of many important molecules such as sugar phosphates, adenosine triphosphate, ribonucleic acids, deoxyribonucleic acid, and biomembrane phospholipids2. In addition, P also participates in many metabolic processes3. Although P is rich in the soils, only a small proportion (< 30%), in form of orthophosphate, can be uptake by plants4,5. Therefore, P deficiency is not uncommon and is a constraint for crop production in > 30% of the arable land that can be treated with P fertilizers6,7. The demand for P fertilizers worldwide in 2022 was predicted as high as 49.1 million tons8; however excessive application of P fertilizer also caused severe problems such as environmental pollution7. Therefore, the development of a new crop variety with efficient P use is required for agricultural production in P-deficit soils9,10.
Several sophisticated strategies have been evolved in plants to cope with low P stress11, such as growing more roots to promote P absorption, secreting acid phosphatase to hydrolyze P in the soil, and activating a cassette of phosphate stress-responsive genes to enhance the transport and absorption of P12. Several metabolic pathway genes, regulatory factors, membrane transporter proteins, and endogenous phytohormones are involved in P deficiency responses or P signal transduction13. Among these important factors, microRNAs (miRNAs) were regarded as key regulators in plant P starvation response14,15,16.
miRNAs are defined as a group of regulatory noncoding RNAs with 20–24 nucleotides in length, which modulate gene expression at the posttranscriptional level in plants. Among these, miR156 is the first kind of miRNA identified, which mainly targets genes that encode SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) proteins17. By interacting with a subset of SPLs, miR156 exerts its function in a variety of developmental processes such as stress response17,18. For instance, miR156 together with its target SPL3 is a key regulator in response to P starvation by directly modulating the expression of three genes (i.e., PLDZ2, Pht1;5, and miR399f) related to P uptake in the model plant Arabidopsis19. The role of miRNAs in P-deficiency signal transduction was also investigated in rapeseed20 and common beans21. Apart from miR156, many other miRNAs were also detected in response to P starvation. For instance, miR156/778/827/2111 were induced upon P deprivation, while miR169/395/398 were suppressed20,21,22,23. In soybean, at least 57 miRNAs were significantly up- or down-regulated under P deficiency24.
Soybean is the major oil crop worldwide providing oil for humans and protein for livestock, with a total production of 362 million tons in 2020 (https://www.fao.org/faostat/en/#search/soybean). Under low P conditions, the production and quality of soybean are seriously affected. Recently, miR156 was found to be up-regulated in Arabidopsis root and soybean leaf in response to low P stress23,24. Moreover, we also observed increased expression of miR156b in the shoots of soybean at 0.5–12 h after P-deficiency treatment (unpublished). However, very little is known about the consequence of overexpressing miR156 on plant architecture and seed yield under P stress conditions. To gain a deeper understanding of the function of miR156 in the phosphorus signaling pathway, we developed Arabidopsis lines overexpressing the soybean homolog of miR156b (termed GmmiR156b hereafter) and investigated their responses to P deficiency. The transgenic Arabidopsis lines overexpressing GmmiR156b were found to be more tolerant to P deficiency and have a higher seed yield. In addition, pleiotropy was also observed in the transgenic lines such as dwarfness, longer growth period, bushy shoot, and shorter silique length. Our study will shed new light on molecular mechanisms of P starvation adaption in soybean and will help find new ways to increase P use efficiency.
Plant materials and methods
GmmiR156b gene isolation, vector construction, and plant transformation
A genomic fragment comprising the whole sequence of GmmiR156b was amplified from soybean variety ‘BX10’ using the primer pair 156b-F1 and 156b-R1, which were designed based on the upstream and downstream of GmmiR156b (MI0001790, chr14: 990334-990453, http://www.mirbase.org), respectively (Supplementary table 1), and deposited in NCBI under accession No. OP784762. PCR was performed using the following parameters: one cycle of denaturing at 94 ℃ for 2 min; then 35 cycles of denaturing at 94 ℃ for 30 s, annealing at 58 ℃ for 30 s, and extension at 72 ℃ for 30 s; followed by inoculation at 72 ℃ for 5 min. The resultant PCR product was purified and inserted into the pCAMBIA1300-based T-DNA vector pJG045 driven by two 35S cauliflower mosaic virus promoters25. The vector was then induced into Agrobacterium strain GV3101 before transforming into Arabidopsis thaliana ecotype Columbia (Col-0) by the floral dipping method26. Seeds harvested from the transgenic generation 1 (T1) plants were screened for hygromycin resistance (25 µg mL−1) on Murashige and Skoog (MS) medium, and the survived plants were grown in separate pots to harvest T2 seeds, from which T2 lines exhibiting a 3:1 segregation ratio of hygromycin resistance were selected and further confirmed by PCR to be homozygous for insertion. The confirmed homozygous T3 lines were used for further analyses. The transcript abundance in transgenic lines was quantified by RT-PCR using the specific primer pairs 156b-F1 and 156-R1.
Plant materials and growth conditions
The seeds of Col-0 and the GmmiR156b overexpression (OE) lines were sterilized with 10% sodium hypochlorite (v/v) for 10 min and washed 5 times with ddH2O, then germinated in Petri dishes containing MS solid medium (pH 5.8), 0.8% (w/v) agar, and 1% (w/v) sucrose. The seeds in Petri dishes were placed at 4 ℃ for 2 days for stratification and then moved to a plant growth chamber for germination 22 ℃ under long-day conditions (16 h light/8 h dark; 95 μmol m−2 s−1). After 5 days of growth, the seedlings were transferred to P-sufficient (+ P; 1 mmol L−1 KH2PO4) or P-deficient (−P; 0.5 mmol L−1 K2SO4) medium27. For seedlings analysis, seeds were germinated and then vertically grown on the surface of the agar. For adult plant analysis, the 5-day-old seedlings were transplanted from MS medium to vermiculite-filled pots supplied with + P or − P liquid medium, and grown in a greenhouse at a constant temperature of 22 ℃, photoperiod of 16 h light/8 h dark, and relative humidity of 70%.
RT-PCR and qPCR analysis
Total RNA was prepared from fresh tissue samples using a Trizol reagent kit (Invitrogen, Carlsbad, CA, USA) and treated with DNaseI (Ambion, USA) to remove any genomic DNA residual. For RT-PCR analysis, 2 µg of total RNA was reversely transcribed by MMLV (Promega, USA) and oligo-d(T) primer to synthesize the first-strand cDNA following the manufacturer’s instruction. Then the ten-fold diluted cDNA was used as the template for the performance of RT-PCR with gene-specific primer pairs 156b-F1 and 156b-R1. Quantitative real-time PCR was carried out using primer combination 156b-F2/156b-R2 as previously reported27. The sequences of all primers for RT-PCR and quantitative real-time PCR in this study were given in Additional File 1. The relative gene expression levels were normalized by using Tublin as an internal reference gene following the method reported by Livak and Schmittgen28,29.
Quantification of phosphorus content
The P content was measured by the malachite green method30. The shoots and roots of 10-day-old Arabidopsis seedlings, as well as shoots of adult plants at the bolting stage for Col-0, OE1, and OE2 lines growing in + P or − P treatments were excised, weighed, and dehydrated at 80 ℃ for 24 h. After digestion with sulfuric acid and hydrogen peroxide, approximately 40 µL of 5 mol L−1 H2SO4 per 20 mg dried samples were mixed with 3 mL of water, then first filtered by Whatman No. 4 filter paper, and then the supernatant aliquot was brought to a final volume of 1.5 mL with water, followed by the addition of 0.5 mL malachite green solution and mixed thoroughly. After inoculation at room temperature for at least 30 min, the phosphorus content was quantified from the optical absorbance values of the above mixture detected at a wavelength of 650 nm in a spectrophotometer (METASH, Shanghai, China).
Measurement of biological traits
Fifteen seedlings per line were selected as one replicate for each treatment and four biological replicates were carried out. The root length of 10-day-old seedlings of Col-0, OE1, and OE2 lines growing in + P and − P conditions were investigated. Then the roots and shoots were dissected, weighted, and measured for P content. The fresh weight shoot/root ratio was calculated (Sha et al.27). For the investigation of yield component traits, each of the ten plants of Col-0 and OE1, and OE2 growing in either + P or − P conditions were collected with four biological replicates. Silique length was measured on five siliques from the medial inflorescence of a plant. Comparisons of means for all traits with replicates were statistically analyzed in IBM SPSS Version 16 (SPSS, Chicago, IL, USA).
Ethics declaration
All authors agree to the ethics and consent to participate in this article and declare that this submission follows the policies of Scientific Reports. The paper is not being considered for publication elsewhere. All authors confirmed that experimental research on plants, including gene transformation and sample collection activities, complying with relevant institutional, national, and international guidelines and legislations. Furthermore, the handling of genetically modified organisms was carried out according to the local relevant guidelines and regulations of China.
Results
Construction and overexpression of GmmiR156b
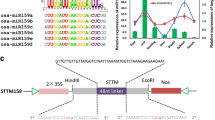
A genomic fragment of pri-miR156b (GmmiR156b) was successfully amplified from soybean by PCR, which forms a typical stem-root structure showing 45% sequence similarity with Arabidopsis pri-miR156b but nearly identical in the mature gene sequences (Fig. 1). The DNA fragment was then inserted into the expression vector pJG045. After verification by DNA sequencing, the construct was transformed into Arabidopsis, and eight independent transgenic events were obtained. The T1 plants showing hygromycin resistance could be verified by PCR using gene-specific primers (Fig. 2a), then the expression level of pri-GmmiR156b was detected by RT-PCR, which indicated that all of the eight transgenic overexpression lines (named OE1 to OE8) exhibited a very high expression level of GmmiR156b (Fig. 2b). Next, quantitative real-time PCR (qPCR) further revealed that, if compared to the wild type (WT), the mature GmmiR156b was highly expressed in the eight transgenic lines, with a relative gene expression level varying from 70.0 to 756.3 folds increase and a mean of 298.9 folds for all OE lines (Fig. 2c). Of these, only two lines (namely OE1 and OE2) showed an overexpression higher than 600 folds, which were thus chosen for further analysis.
Verification of transgenic events and quantification of GmmiR156b expression in Arabidopsis seedlings. (a) PCR confirmation of positive transgenic Arabidopsis plants carrying GmmiR156b by gene-specific primers. (b) RT-PCR analysis of pri-GmmiR156b in the leaf of the wild type (WT) and eight independent transgenic lines. (c) Quantitative RT-PCR analysis of mature GmmiR156b in 8 overexpression (OE) lines. The relative expression level of mature GmmiR156b was normalized to that of the inner reference gene Tublin. Lanes 1–8, transgenic lines; lane 9, the WT. M, DNA marker DL2000. ** Indicates statistical difference between GmmiR156b and reference gene at P < 0.01. Original gels are presented in Supplementary Fig. 1.
Overexpression of GmmiR156b confers enhanced tolerance to P deficiency
At the seedling stage under + P condition, the Arabidopsis WT, OE1, and OE2 plants grew normally, all having primary roots with a similar length of 3 cm. However, under − P condition, the primary root length was reduced by nearly 50% for all lines, suggesting that P starvation limited plant growth substantially. Nevertheless, the roots of OE1 and OE2 (~ 1.5 cm) were significantly longer than that of WT (1.1 cm) under P nutritional stress (Fig. 3a), implying that plant growth could be partially rescued by overexpressing GmmiR156b. Meanwhile, the root/shoot ratios ranged from 0.15 to 0.24 under the + P condition but decreased markedly (0.12–0.15) under the − P condition, with the lowest value observed in OE1 and the highest in WT, under both situations (Fig. 3b). Interestingly, the higher root/shoot ratios in WT seedlings were found to be caused by higher fresh weight of roots. In addition, the total P contents of OE1 and OE2 lines were higher than that of WT in the roots but similar in the shoots under − P condition (Fig. 3c), showing that overexpressing GmmiR156b could promote the uptake and accumulation of P in the root rather than the shoot. When grown in normal condition, the total P contents of a given tissue (root or shoot) was similar among the OE lines and WT but varied between tissues, i.e., higher content in the roots and lower in the shoots at the seedling stage (Fig. 3c).
Root length, root/shoot ratio, and phosphorus contents for 10-day-old Arabidopsis seedlings and adult plants. (a) Root length of wild-type (WT) and transgenic plants overexpressing GmmiR156b (OE1, OE2) under + P (Pi-sufficient) and − P (Pi-deficient) conditions. (b) Fresh weight root/shoot ratio of 10-day-old seedlings of OE1, OE2, and WT under + P and − P conditions. (c) Phosphorus content in root and shoot of OE1, OE2, and WT under + P and − P conditions. (d) Phosphorate content in roots of adult plants of OE and WT lines. (e) The phenotype of WT and OE seedlings grown under + P and − P conditions. Different letters above the bars indicate statistical significance (P < 0.05). Shoot-, shoots from low P grown plant; Root-, roots from low Pi grown plant; Shoot +, shoots from high P grown plant; Root +, roots from high P grown plant. Scale bar = 1 cm.
At the adult plant stage, the OE1 and OE2 lines still maintained a better tolerance to P deficiency than WT. Under − P condition, the WT plants grew poorly and the first senescent leaf appeared 55 days after emergence (DAE), whereas the OE1 and OE2 lines still grew vigorously and the leaves maintained green at least 70 DAE. Under normal conditions, the WT plants grew well until 70 DAE when all leaves became fully senescent, whereas the leaves of OE1 and OE2 were still green (Fig. 4a). Moreover, the total P content in roots of OE1 and OE2 was higher than that in WT under − P condition; however, there was no obvious difference under + P condition (Fig. 3d).
Phenotype of adult Arabidopsis plants grown under P-sufficient and P-deficient conditions. (a) The morphological phenotype of GmmiR156b overexpressing lines (OE1, OE2) and the wild type (WT) on 55 days and 70 days after emergence (DAE) under + P (P sufficient) and − P (P deficient conditions). (b) Siliques of WT and OE lines, bar = 1 cm. (c) Mature seeds of WT and OE lines, bar = 1 mm.
Seed yield increased by GmmiR156b overexpression
In normal growth conditions, the average number of rosette leaves in WT was 23.4 ± 3.5, which was much lower than the OE lines (44.4 ± 8.7) showing dwarfness; however, an opposite trend was found for the number of primary shoots (7.1 ± 1.3 versus 5.3 ± 1.2). Under − P condition, the WT plants were affected remarkedly but the OE lines were affected slightly (Fig. 4); the average seed yield per plant was 42–48 mg for the OE lines but only 33 mg for the WT (Fig. 5a), since the OE lines could grow for a much longer period (> 70 days) thus a better seed setting. It was noted that, although silique length was ~ 35% shorter in OE lines (Fig. 5c), the width of silique was much larger (Fig. 5d) and allowed for a bigger seed size (Figs. 4c and 5b).
Yield traits in transgenic and wild-type Arabidopsis lines. (a) Seed yield of GmmiR156b overexpression (OE) and wild type (WT) plants under + P and − P conditions. (b) Seed weight of OE and WT plants under + P and − P conditions. (c) Fruit length of OE and WT plants under + P and − P conditions. (d) Fruit width of OE and WT plants under + P and − P conditions. Different letters above the bars indicate statistical significance (P < 0.05).
Expression of genes targeted by GmmiR156b in Arabidopsis
There are 10 members in the SPL gene family targeted by miR156 in Arabidopsis, seven of which can be grouped into three major clades based on their functions in the regulation of plant morphology and development, namely AtSPL2/10/11, AtSPL9/15, and AtSPL6/1331. To investigate which SPL genes (i.e., AtSPL2/3/4/5/6/9/10/11/13/15) are regulated by GmmiR156b in Arabidopsis transgenic lines, we detected the expression of the above ten AtSPLs in two GmmiR156b OE lines and WT plants by qRT-PCR. Six genes (AtSPL4/5/6/9/11/15) were down-regulated in at least one of the transgenic lines. Of these, the expression of AtSPL15 was most affected and decreased by more than 90% (Fig. 6), suggesting that AtSPL15 is the major target of GmmiR156b. In contrast, the expression of the other four AtSPLs (i.e., AtSPL2/3/10/13) in transgenic lines was equal to or higher than that in wild-type plants (Fig. 5), indicating that they are not the direct targets of GmmiR156b.
Transcriptional analysis of AtSPLs targeted by GmmiR156b in overexpression lines. Quantitative RT-PCR analysis was performed using RNA extracted from wild-type (WT) and overexpression lines (OE1 and OE2) for putative targets of GmmiR156b (i.e., AtSPL2/3/4/5/6/9/10/11/13/15). The relative expression level of AtSPLs in OE lines was normalized to that of the WT. Tublin was used as inner control for gene expression analysis. ** indicates a statistical difference between AtSPLs and Tublin at P < 0.01.
Discussion
miR156 is a pivotal regulator for plant growth and development with pleiotropic effects. Several studies reported that miR156 is not only a key player in modulating a series of plant developmental processes such as branching, early flowering, fruit ripening, and cell proliferation but also in responses to environmental nutritional stresses by interaction with its targets, the SPL gene family32,33,34,35. Here we demonstrated for the first time, that overexpression of soybean homolog of miR156 in Arabidopsis could directly enhance the tolerance to P deficiency and seed yield under P deficit conditions, which is valuable for the development of efficient P-use soybean varieties in the future.
Plants usually cope with P deficiency by changing root architecture such as alteration of the root-to-shoot ratio, an increase of total root length, as well as lateral root lengths/numbers to improve the absorbance of P33,34,35. Lei et al.19 reported that miRNA156, together with its target SPL3, plays a pivotal role in responses to low P stress. They found that the transcript level of miR156 was significantly increased under low P condition but SPL3 was suppressed. Transgenic lines constantly expressing SPL3 grown under P-deficient conditions exhibited decreased rhizosphere acidification (thus closer to the optimal pH facilitating the release of insoluble P), which promoted P uptake. Moreover, a P transporter (i.e., Pht1;5) was found to be significantly activated via SPL3 and hence increased P uptake in the root19. In this study, however, we found that overexpression of GmmiR156 didn’t alter the level of AatSPL3 significantly, suggesting that AatSPL3 is not the main target of GmmiR156b (Fig. 6). Instead, some other SPL genes (i.e., AtSPL4/5/615) were substantially suppressed, resulting in more efficient P uptake in the root, delayed flowering, and prolonged juvenal period. Therefore, we conclude that these four SPL genes (especially AtSPL15) are the main targets for GmmiR156b. In Arabidopsis, SPL2/910/11/13/15 modulate a variety of biological processes in root and shoot development whereas SPL3/4/5 predominately boosts floral induction and/or floral meristem identity32. miR156 represses SPL gene expression by cleaving SPL transcripts and by promoting their translational repression35. So far, there are no reports demonstrating the importance of SPL15 in P metabolite, and it is worth to further investigate the mechanism of the miR156-SPL15 module in the regulation of P uptake in soybean. Apart from miR156, many other miRNAs have been shown to take part in the P starvation response in soybean. For instance, Sha et al.27 identified 60 miRNAs in soybean via deep sequencing, which exhibited an altered gene expression pattern at least in roots or shoots under P stress conditions. Moreover, miRNA399-PHO2 modules are found to orchestrate the P homeostasis in Arabidopsis by mediating the loading of P in the xylem by the breakdown of PHO136,37.
Rhizosphere acidification is frequently observed in response to low P stress, which expedites the discharge of P from soil and apoplasts38. SPL3 overexpression plants displayed an increased level of P uptake because of decreased rhizosphere acidification19. In the present study, the adult plant of GmmiR156 overexpression lines showed more tolerance to P starvation (Fig. 4). Higher P content was accumulated in shoots of transgenic lines which resulted in better growth of the transgenic lines than WT under P-deficient conditions. The senescence process of transgenic lines largely lagged behind the WT under both P deficient and P sufficient conditions (Fig. 4). One reason is likely that overexpression of GmmiR156b delayed the vegetative-to-reproductive phase transition19. Another explanation is that the continuous growing shoots, referred to as bushy architecture39, resulted in inconsistent maturity of the old and young shoots (Fig. 4).
We also observed that the primary root lengths of OE1 and OE2 seedlings are longer than that of WT, and P content is higher in shoots of OE1 and OE2 than in WT under low P stress conditions, suggesting that the overexpression of GmmiR156b might increase P uptake by promoting primary root growth of seedlings (Fig. 3). The result is similar to the report in maize that the length of 12-day-old seedlings was increased under low P conditions40. However, the ratio of root-to-shoot in OE1 and OE2 seedlings lines was decreased under both P deficient and P sufficient conditions. It might be attributed to the promoted shoot growth by overexpressing GmmiR156, which has been reported in torenia39, Arabidopsis41, tomato42, switchgrass43, and alfalfa44.
A group of genes control the two important transitions in shoot development and are coordinated by miR156 and its target SPL gene families45. Several studies reported that the pleiotropic effect of overexpressing miR156 includes the delay of juvenile-to-adult phase transition, more branches, smaller leaf size, and altered metabolism in plants46,47,48,49. Highly overexpressing of miR156b in transgenic poplar lines exhibited a globulous phenotype with many branches and leaves, markedly smaller leaf size, and the accumulation of reddish pigment50. Similar to the above reports, in the present study, overexpression of GmmiR156b showed increased seed yields and strong pleiotropic developmental phenotypes such as dwarfness, prolonged growth period, bushy shoot, and shorter silique length (Figs. 4 and 5). Our observation is also in agreement with the increase of soybean seed yield in GmmiR156 overexpressed lines, by suppression of its target GmSPL951. We also found that the length of silique was shorter in transgenic lines than in WT, which is reminiscent of the Arabidopsis sk156 mutant in which expression of miR156b was enhanced31. Moreover, six SPL genes, namely AtSPL4/5/6/11/15 were down-regulated in GmmiR156b overexpression lines (Fig. 6). Wei et al.31 reported that suppression of SPL15 causes decreased silique size; SPL4 and SPL5 mainly control plant juvenile-to-adult transition; SPL9 and SPL15 regulate shoot maturation and leaf initiation, while SPL11 modulate shoot maturation in the reproductive phase. Intriguingly, we found that SPL15 was most suppressed in GmmiR156b OE lines, resulting in a longer growth period and delayed senescent progress than WT (Fig. 6). This information collectively indicates that the developmental morphology and increased P deficiency tolerance of GmmiR156b OE lines might be the consequence of the suppression of these target SPL genes.
In conclusion, here we showed that suppression of SPL genes, especially AtSPL15 by GmmiR156b resulted in longer primary roots and higher P contents in the roots of seedlings. Overexpression of GmmiR156b also increased seed yields and exhibited strong pleiotropic developmental phenotypes under P-deficient conditions. These findings will deepen our understanding of the post-transcriptional regulation of gene expression under low P in higher plants and highlight the pivotal role of the miR156b-SPL15 module in responses to P starvation.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request. The DNA fragment containing soybean miR156b is deposited in NCBI under accession No. OP784762 (https://www.ncbi.nlm.nih.gov/).
References
Vance, C. P. Symbiotic nitrogen fixation and phosphorus acquisition. Plant nutrition in a world of declining renewable resources. Plant Physiol. 127, 390–397 (2001).
Schachtman, D. P., Reid, R. J. & Ayling, S. M. Phosphorus uptake by plants: From soil to cell. Plant Physiol. 116, 447–453 (1998).
Yuan, H. & Liu, D. Signaling components involved in plant responses to phosphate starvation. J. Integr. Plant Biol. 50, 849–859 (2008).
Raghothama, K. G. Phosphate acquisition. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 665–693. https://doi.org/10.1146/annurev.arplant.50.1.665 (1999).
Nadeem, M. et al. Understanding the adaptive mechanisms of plants to enhance phosphorus use efficiency on podzolic soils in boreal agroecosystems. Front. Plant Sci. 13, 804058. https://doi.org/10.3389/fpls.2022.804058 (2022).
Abel, S., Ticconi, C. A. & Delatorre, C. A. Phosphate sensing in higher plants. Physiol. Plantarum 115, 1–8 (2002).
Vance, C. P., Uhde-Stone, C. & Allan, D. L. Phosphorus acquisition and use: Critical adaptations by plants for securing a nonrenewable resource. New Phytol. 157, 423–447 (2003).
FAO. World Fertilizer Trends and Outlook to 2022. Rome. https://www.fao.org/3/ca6746en/CA6746EN.pdf (2019).
Jamla, M., Joshi, S., Patil, S., Tripathi, B. N. & Kumar, V. MicroRNAs modulating nutrient homeostasis: A sustainable approach for developing biofortified crops. Protoplasma https://doi.org/10.1007/s00709-022-01775-w (2022).
Begum, Y. Regulatory role of microRNAs (miRNAs) in the recent development of abiotic stress tolerance of plants. Gene 821, 146283. https://doi.org/10.1016/j.gene.2022.146283 (2022).
Islam, W., Tauqeer, A., Waheed, A. & Zeng, F. MicroRNA mediated plant responses to nutrient stress. Int. J. Mol. Sci. 23, 2562. https://doi.org/10.3390/ijms23052562 (2022).
Prathap, V., Kumar, A., Maheshwari, C. & Tyagi, A. Phosphorus homeostasis: Acquisition, sensing, and long-distance signaling in plants. Mol. Biol. Rep. 49, 8071–8086. https://doi.org/10.1007/s11033-022-07354-9 (2022).
Jodder, J. miRNA-mediated regulation of auxin signaling pathway during plant development and stress responses. J. Biosci. 45, 91 (2020).
Zhang, B., Pan, X., Cobb, G. P. & Anderson, T. A. Plant microRNA: A small regulatory molecule with big impact. Dev. Biol. 289, 3–16 (2006).
Kuo, H. F. & Chiou, T. J. The role of microRNAs in phosphorus deficiency signaling. Plant Physiol. 156, 1016–1024 (2011).
Paz-Ares, J. et al. Plant adaptation to low phosphorus availability: Core signaling, crosstalks, and applied implications. Mol. Plant 15, 104–124. https://doi.org/10.1016/j.molp.2021.12.005 (2022).
Stief, A. et al. Arabidopsis miR156 regulates tolerance to recurring environmental stress through SPL transcription factors. Plant Cell 26, 1792–1807 (2014).
Jerome-Jeyakumar, J. M., Ali, A., Wang, W. M. & Thiruvengadam, M. Characterizing the role of the miR156-SPL network in plant development and stress response. Plants (Basel) 9, 1206. https://doi.org/10.3390/plants9091206 (2020).
Lei, K. J. et al. Modulation of the phosphate-deficient responses by microRNA156 and its targeted SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 3 in Arabidopsis. Plant Cell Physiol. 57, 192–203. https://doi.org/10.1093/pcp/pcv197 (2016).
Pant, B. D. et al. Identification of nutrient-responsive Arabidopsis and rapeseed MicroRNAs by comprehensive realtime polymerase chain reaction profiling and small RNA sequencing. Plant Physiol. 150, 1541–1555 (2009).
Valdes-Lopez, O. et al. Essential role of MYB transcription factor: PvPHR1 and microRNA: PvmiR399 in phosphorus-deficiency signalling in common bean roots. Plant Cell Environ. 31, 1834–1843 (2008).
Pant, B. D., Buhtz, A., Kehr, J. & Scheible, W. R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 53, 731–738 (2008).
Hsieh, L. C. et al. Uncovering small RNA-Mediated responses to phosphate deficiency in Arabidopsis by deep sequencing. Plant Physiol. 151, 2120–2132 (2009).
Zeng, H. Q., Zhu, Y. Y., Huang, S. Q. & Yang, Z. M. Analysis of phosphorus-deficient responsive miRNAs and cis-elements from soybean (Glycine max L.). J. Plant Physiol. 167, 1289–1297 (2010).
Zhao, J. et al. The rubisco small subunit is involved in tobamovirus movement and Tm-22-mediated extreme resistance. Plant Physiol. 161, 374–383 (2013).
Clough, S. J. & Bent, A. F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Sha, A. H. et al. Identification of Glycine Max microRNAs in response to phosphorus deficiency. J. Plant Biol. 55, 268–280 (2012).
Kulcheski, F. R. et al. Identification of novel soybean microRNAs involved in abiotic and biotic stresses. BMC Genom. 12, 307 (2011).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Xu, L. et al. Overexpression of GbWRKY1 positively regulates the Pi starvation response by alteration of auxin sensitivity in Arabidopsis. Plant Cell Rep. 31, 2177–2188 (2012).
Wei, S. et al. Arabidopsis mutant sk156 reveals complex regulation of SPL15 in a miR156-controlled gene network. BMC Plant Biol. 12, 169 (2012).
Wang, H. & Wang, H. The miR156/SPL module, a regulatory hub and versatile toolbox, gears up crops for enhanced agronomic traits. Mol. Plant 8, 677–688. https://doi.org/10.1016/j.molp.2015.01.008 (2015).
Li, Z., Tian, P., Huang, T. & Huang, J. Noncoding-RNA-mediated regulation in response to macronutrient stress in plants. Int. J. Mol. Sci. 22, 11205. https://doi.org/10.3390/ijms222011205 (2021).
Li, Z. et al. Phosphate starvation of maize inhibits lateral root formation and alters gene expression in the lateral root primordium zone. BMC Plant Biol. 12, 89 (2012).
Dong, Q., Hu, B. & Zhang, C. microRNAs and their roles in plant development. Front. Plant Sci. 13, 824240. https://doi.org/10.3389/fpls.2022.824240 (2022).
Liu, T. Y. et al. PHO2-dependent degradation of PHO1 modulates phosphate homeostasis in Arabidopsis. Plant Cell 24, 2168–2183. https://doi.org/10.1105/tpc.112.096636 (2012).
Stefanovic, A. et al. Over-expression of PHO1 in Arabidopsis leaves reveals its role in mediating phosphate efflux. Plant J. 66, 689–699. https://doi.org/10.1111/j.1365-313X.2011.04532.x (2011).
Neumann, G., Massonneau, A., Martinoia, E. & Römheld, V. Physiological adaptations to phosphorus deficiency during proteoid root development in white lupin. Planta 208, 373–382 (1999).
Shikata, M., Yamaguchi, H., Sasaki, K. & Ohtsubo, N. Overexpression of Arabidopsis miR157b induces bushy architecture and delayed phase transition in Torenia fournieri. Planta 236, 1027–1035 (2012).
Anghinoni, I. & Barber, S. A. Phosphorus influx and growth characteristics of corn roots as influenced by phosphorus supply. Agron. J. 172, 655–668 (1980).
Schwab, R. et al. Specific effects of microRNAs on the plant transcriptome. Dev. Cell 8, 517–527 (2005).
Zhang, X. et al. Over-expression of sly-miR156a in tomato results in multiple vegetative and reproductive trait alterations and partial phenocopy of the sft mutant. FEBS Lett. 585, 435–439 (2011).
Fu, C. et al. Overexpression of miR156 in switchgrass (Panicumvirgatum L.) results in various morphological alterations and leads to improved biomass production. Plant Biotech. J. 10, 443–452 (2012).
Aung, B. et al. microRNA156 as a promising tool for alfalfa improvement. Plant Biotech. J. 13, 779–790 (2015).
Xu, M. et al. Developmental functions of miR156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet. 12, e1006263. https://doi.org/10.1371/journal.pgen.1006263 (2016).
Luo, P. et al. microRNAs are involved in regulating plant development and stress response through fine-tuning of TIR1/AFB-dependent auxin signaling. Int. J. Mol. Sci. 23, 510. https://doi.org/10.3390/ijms23010510 (2022).
Manuela, D. & Xu, M. Juvenile leaves or adult leaves: Determinants for vegetative phase change in flowering plants. Int. J. Mol. Sci. 21, 9753. https://doi.org/10.3390/ijms21249753 (2020).
Chand-Jha, U., Nayyar, H., Mantri, N. & Siddique, K. H. M. Non-coding RNAs in legumes: Their emerging roles in regulating biotic/abiotic stress responses and plant growth and development. Cells 10, 1674. https://doi.org/10.3390/cells10071674 (2021).
Wu, G. & Poethig, R. S. Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 133, 3539–3547. https://doi.org/10.1242/dev.02521 (2006).
Wang, Y. et al. miR156 regulates anthocyanin biosynthesis through SPL targets and other microRNAs in poplar. Hortic. Res. 7, 118. https://doi.org/10.1038/s41438-020-00341-w (2020).
Sun, Z. et al. Genetic improvement of the shoot architecture and yield in soya bean plants via the manipulation of GmmiR156b. Plant Biotechnol. J. 17, 50–62. https://doi.org/10.1111/pbi.12946 (2019).
Acknowledgements
This work was supported by the Guangdong University of Petrochemical Technology (Talent Fund), Science and Technology Project of Guangdong Laboratory Maoming Branch Center of Lingnan Modern Agricultural Science and Technology (2022KF015), Maoming City Science and Technology Planning Project (2021KJZXZJGSPDX004, 2021N002).
Author information
Authors and Affiliations
Contributions
G.L. and Y.Z. designed the study. Z.T., Y.H., M.X., Y.L., and J.W. conducted the experimental works. G.L. and Z.T. analyzed the data and wrote the paper. Y.Z. modified the language and approved the final manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, G., Tian, Z., Hao, Y. et al. Overexpression of soybean microRNA156b enhanced tolerance to phosphorus deficiency and seed yield in Arabidopsis. Sci Rep 13, 652 (2023). https://doi.org/10.1038/s41598-023-27847-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-27847-2
- Springer Nature Limited