Abstract
The AmrZ/FleQ hub has been identified as a central node in the regulation of environmental adaption in the plant growth-promoting rhizobacterium and model for rhizosphere colonization Pseudomonas ogarae F113. AmrZ is involved in the regulation of motility, biofilm formation, and bis-(3′-5′)-cyclic dimeric guanosine monophosphate (c-di-GMP) turnover, among others, in this bacterium. The mutants in amrZ have a pleiotropic phenotype with distinguishable colony morphology, reduced biofilm formation, increased motility, and are severely impaired in competitive rhizosphere colonization. Here, RNA-Seq and qRT-PCR gene expression analyses revealed that AmrZ regulates many genes related to the production of extracellular matrix (ECM) components at the transcriptional level. Furthermore, overproduction of c-di-GMP in an amrZ mutant, by ectopic production of the Caulobacter crescentus constitutive diguanylate cyclase PleD*, resulted in increased expression of many genes implicated in the synthesis of ECM components. The overproduction of c-di-GMP in the amrZ mutant also suppressed the biofilm formation and motility phenotypes, but not the defect in competitive rhizosphere colonization. These results indicate that although biofilm formation and motility are mainly regulated indirectly by AmrZ, through the modulation of c-di-GMP levels, the implication of AmrZ in rhizosphere competitive colonization occurs in a c-di-GMP-independent manner.
Similar content being viewed by others
Introduction
Pseudomonas ogarae F113, formerly known as P. fluorescens F1131, was isolated from the sugar-beet rhizosphere and is considered a plant-growth-promoting and biocontrol rhizobacterium2. This bacterium can colonize the roots of many plants and is used as a model for rhizosphere colonization3,4,5. The adaption of F113 to the rhizosphere environment is partially dependent on two master transcription factors (TFs), AmrZ and FleQ, and a second messenger molecule, bis-(3′-5′)-cyclic dimeric guanosine monophosphate (c-di-GMP)6,7. The transition between motile and sessile lifestyles is also controlled by the levels of this second messenger, which are in turn controlled by the activity of diguanylate cyclases (DGCs), containing GGDEF domains, involved in c-di-GMP synthesis and phosphodiesterases (PDEs), which contain EAL or HD-GYP domains and degrade c-di-GMP8,9. High c-di-GMP levels are associated with a sessile lifestyle and biofilm formation, whereas low levels of this second messenger are responsible for motility and a planktonic lifestyle10. The role of this molecule in modulating rhizosphere colonization and attachment to the plant has been demonstrated in F113, Pseudomonas putida KT2440, and Pseudomonas syringae11,12,13. AmrZ has been described as a regulator of more than half of the DGCs and PDEs encoded in the F113 genome and its mutant is deficient in c-di-GMP production7.
AmrZ belongs to the Arc superfamily of proteins and possesses a ribbon-helix-helix DNA binding domain, a flexible N-terminus, and a C-terminal domain14. AmrZ is ubiquitously found in Pseudomonas and its activity depends on the formation of tetramers mediated by its C-terminal domain15. The AmrZ protein was first described as a regulator of the production of the exopolysaccharide (EPS) alginate14,16. Later, it has been shown to regulate several other functions, including motility and c-di-GMP metabolism in different pseudomonads7,15,17,18,19,20,21 functioning either as an activator or a repressor of the gene expression22 and with opposing roles in different strains23. In F113, AmrZ is involved in the regulation of different functions such as motility, biofilm formation, c-di-GMP production, or iron homeostasis7,24. Conversely, this protein has been associated with repression of c-di-GMP synthesis in P. aeruginosa19,20 and P. syringae pv. tomato DC3000, being the former a negative transcriptional regulator of cellulose synthesis and an activator of alginate production, motility, and virulence21. In P. aeruginosa and F113, AmrZ has been identified as a negative transcriptional regulator of FleQ, the master regulator for flagellar synthesis18,20,25,26. FleQ is another bifunctional TF, able to act as a transcriptional activator or repressor27. This protein is recognized as a c-di-GMP effector, able to bind c-di-GMP28. The TF FleQ can inversely regulate swimming motility and EPSs production in a c-di-GMP-dependent manner in several species of pseudomonads27,29,30,31. In F113, AmrZ and FleQ constitute a regulatory hub in which they mutually and transcriptionally repress each other while conversely regulating other functions: AmrZ down-regulates motility and up-regulates putative extracellular matrix (ECM) components production whereas FleQ up-regulates motility and down-regulates putative ECM components production6. A similar antagonistic role of both TFs has also been shown by RNA-Seq during rhizosphere colonization32.
The ECM is the structure that binds cells in biofilms, acting mainly as a scaffold of these three-dimensional structures. It is composed mostly of different EPSs, extracellular proteins, and DNA33. The ECM of F113 is predicted to contain alginate, levan, Pseudomonas acidic polysaccharide (Pap), and poly-β-1, 6-N-acetylglucosamine (PNAG) polysaccharides, functional amyloids in Pseudomonas (Fap), large adhesion protein A (LapA) and medium adhesion protein A (MapA) adhesins, the putative extracellular mannuronan C-5 epimerase PsmE, and the fimbrial low-molecular-weight protein/tight adherence (Flp/Tad) pili34. Since the amrZ mutant is defective in biofilm formation7 and AmrZ has been found as a regulator of some putative ECM components in liquid cultures and rhizosphere7,32, this work aimed to investigate the role of AmrZ in the regulation of the ECM components in F113. Additionally, we have analyzed the implication of c-di-GMP in this AmrZ-mediated regulation and the impact of this second messenger in the AmrZ-associated phenotypes of biofilm formation, motility, and rhizosphere colonization in F113. Finally, we have also studied the cross-regulation of AmrZ and FleQ in the regulation of ECM components and biofilm-related phenotypes.
Results
AmrZ regulates the expression of genes encoding ECM components in P. ogarae F113
As previously observed, amrZ mutants are dramatically impaired in the attachment to surfaces and present a different morphology compared to the wild-type strain in the presence of Congo Red (CR)7. Since these observations are consistent with a defect in the production of ECM components, we have studied the transcriptomic profile of F113 and its isogenic amrZ mutant using RNA-Seq approach32. The RNA-Seq analyses were done under different growth conditions: exponential and stationary phases in liquid cultures and after rhizosphere colonization. As observed in Fig. 1, the transcriptomic analyses have shown that the expression of several gene clusters encoding proteins implicated in polysaccharide synthesis and the production of extracellular structural proteins is downregulated in the amrZ mutant under all tested conditions compared to the wild-type strain. Differential gene expression was found for 59 genes putatively involved in the production of eight different ECM components, mostly observed during stationary phase, indicating a role for AmrZ in activating the expression of genes related to ECM during this phase (Fig. 1). Fold-Change and p-adjusted values for those genes are shown in Supplementary Table S1.
AmrZ is a transcriptional regulator of ECM components in P. ogarae F113. Heatmap displaying the differential gene expression found in the amrZ mutant taking P. ogarae F113 wild-type as the reference at different growth conditions: exponential (Exp) or stationary (St) liquid cultures or rhizosphere (Rhiz). The genes included in this analysis putatively encode ECM components34. The heatmap includes the log2 Fold-Change (amrZ-/wild type) values obtained in RNA-Seq analysis and are included in Supplementary Table S1. Color scale: green, negative regulation by AmrZ; white, no regulation; purple, positive regulation by AmrZ.
The AmrZ-regulated clusters include papA-P (PSF113_1970-1955) involved in the production of the putative Pap polysaccharide and pgaA-D (PSF113_0161-0164) encoding PNAG EPS. Some of the genes from the alg cluster (PSF113_4752-4763), involved in alginate synthesis, were differentially expressed in the amrZ mutant compared to the wild-type strain with log2 Fold-Change values close to the threshold under the tested conditions. The other F113-encoded polysaccharide, levan, is not subjected to AmrZ regulation under any of the tested conditions. Regarding the regulated-clusters related with the production of extracellular proteins or proteinaceous structures, they include fapA-F (PSF113_2680-2685), which are necessary to produce Fap; PSF113_0208, PSF113_1511 and PSF113_3004 encoding the extracellular proteins LapA, MapA, and PsmE, respectively; as well as their associated transport systems (PSF113_0209-0211, PSF113_1508-1510, and PSF113_3005-3007); and PSF113_4178-4192 involved in the Flp/Tad pilus formation.
c-di-GMP governs the AmrZ-dependent regulation of ECM components in P. ogarae F113
We have previously shown that AmrZ is a major regulator of c-di-GMP synthesis and that AmrZ-mediated transcriptional activation of several DGCs occurs mainly during stationary phase7. On the other hand, almost all the ECM-related clusters have a FleQ-binding site6 in their promoter regions and only some of them harbor an AmrZ-binding site24. Moreover, we have previously described the reciprocal regulation between AmrZ and FleQ TFs6. For these reasons, we hypothesized that the observed AmrZ regulation of EMCs could be done through the regulation of c-di-GMP levels or its interrelation with FleQ. In order to test this hypothesis, we first artificially increased the overall c-di-GMP concentration in the wild-type strain, the amrZ mutant, and a double mutant in amrZ and fleQ. The increase of c-di-GMP concentration was obtained through ectopic expression of the mutated version of the pleD* gene from Caulobacter crescentus encoding a constitutive DGC35,36. It has been shown that pleD* overexpression stimulates EPSs production, such as cellulose in P. syringae pv. tomato DC300013.
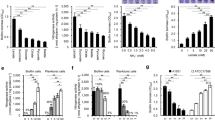
We have also used a qRT-PCR approach to test the expression of selected genes encoding putative ECM components in the wild-type strain and derivatives growing in SA medium at stationary phase either in the absence or presence of pleD* (Fig. 2). As shown in Fig. 2, gene expression was remarkably low, and in some cases, almost undetectable for most of the ECM-related genes tested in amrZ- and amrZ-fleQ- carrying the empty plasmid pJB3Tc19, validating RNA-Seq results. In the case of psmE (PSF113_3004), its gene expression is much higher in the double mutant than in the single mutant, suggesting that FleQ is repressing its expression. Conversely, the presence of pleD* in the amrZ mutant resulted in a significant increase in the expression levels of all tested genes. This increase in expression was several folds higher than in the wild-type for genes encoding Pap (papA, PSF113_1970), PNAG (pgaA; PSF113_0164), Fap (fapB; PSF113_2684), MapA (mapA, PSF113_1511), and Flp/Tad (flp-1, PSF113_4192), demonstrating that the role of AmrZ in the regulation of ECM components occurs indirectly through c-di-GMP. Interestingly, the expression of lapA (PSF113_0208), which encodes the large adhesin LapA was higher in the amrZ–pJBpleD* background than in the amrZ mutant, although expression was still lower than in the wild-type strain, suggesting additional elements involved in its regulation. However, the artificial increase in c-di-GMP levels was not enough to increase gene expression levels in the amrZ-fleQ- background in papA, alg8, fapB, lapA, mapA, and flp-1, also suggesting the dependency of FleQ in their regulation. These results confirm that AmrZ mainly regulates ECM production indirectly, through the increase of c-di-GMP levels and with the interplay of the TF FleQ.
AmrZ regulates ECM-related genes in a c-di-GMP- and FleQ-dependent manner. qRT-PCR analysis of selected extracellular matrix-related genes in stationary phase. Gene expression from amrZ- or amrZ-fleQ- carrying pJB3Tc19 or pJBpleD* was normalized to rpoZ and relativized to the wild-type strain carrying the empty plasmid pJB3Tc19. Averages and standard deviation (SD) from two biological replicates with three technical replicas are represented. Means not sharing any letter are significantly different by multiple t-tests with Bonferroni-Dunn method (p value < 0.05).
The second approach used in this work aims to decipher which DGCs and PDEs are involved in the transcriptional regulation of ECM components in F113. For this purpose, we used mutants affected in the production of DGCs and PDEs with defects in biofilm formation and that have been previously reported as AmrZ-regulated targets: dipA- (PSF113_0499), yfiN- (PSF113_0715), adrA- (PSF113_1982), PSF113_4827, PSF113_3553, and PSF113_46817. DipA and PSF113_4681 contain both GGDEF and EAL motifs, YfiN, AdrA, and PSF113_4827 contain GGDEF motifs, and PSF113_3553 contains an HD-GYP domain. AmrZ activated expression of all the genes encoding these enzymes except for the dipA gene, for which a negative regulation was observed7. As shown in Fig. 3, the gene expression of ECM-related genes is also affected by changes in the pool of c-di-GMP produced by the DGCs and PDEs studied. Our results show that gene expression for papA, pgaA and alg8 are decreased in PSF113_4681 mutant. Besides, papA and pgaA gene expression is increased in the dipA mutant; pgaA gene expression is increased in PSF113_4827 mutant and reduced in adrA mutant. Regarding protein components, psmE, fapB, lapA and flp-1 gene expression is decreased in the PSF113_4681 mutant. Moreover, fapB gene expression is increased in the mutant PSF113_3553; lapA gene expression is reduced in yfiN and adrA mutants; flp-1 gene expression is increased in dipA and adrA mutants and reduced in the PSF113_4827 and finally, mapA gene expression is exclusively increased in the dipA mutant (Fig. 3). These results show that different DGCs and PDEs participate in the regulation of the production of ECM components, by affecting the expression of genes involved in the synthesis of the ECM.
Local DGCs and PDEs are involved in the regulation of ECM-related genes. qRT-PCR analysis of selected extracellular matrix-related genes in stationary phase. Gene expression from mutants affected in the synthesis of proteins with GGDEF motifs (yfiN-, adrA-, and PSF113_4827), HD-GYP domain (PSF113_3553), or containing both GGDEF and EAL motifs (dipA- and PSF113_4681) was normalized to rpoZ and relativized to the wild-type strain. Averages and SD from two biological replicates with three technical replicas are represented. Significant differences in gene expression were determined with multiple t-tests with Bonferroni-Dunn method (*p value < 0.05; **p value < 0.01; ***p value < 0.001; ****p value < 0.0001; ns: not significant).
Ectopic expression of pleD* restores the c-di-GMP related phenotypes in the amrZ and amrZ-fleQ mutants
We have previously reported that amrZ mutants show alterations in CR colony morphology and staining, are hypermotile, and are defective in the attachment to surfaces7. All these phenotypes are consistent with the low levels of c-di-GMP. To test whether these phenotypes were caused by the low levels of c-di-GMP that are prevalent in the amrZ mutant7, we have used ectopic expression of pleD* from a plasmid in the wild-type and amrZ mutant backgrounds. Furthermore, since FleQ is a regulator of motility and biofilm-related genes in this bacterium6 and there is a cross-regulation by AmrZ and FleQ of ECM-related genes expression according to the findings of this work, we have also tested the suppression by c-di-GMP of the CR, biofilm and motility phenotypes in the double mutant amrZ-fleQ.
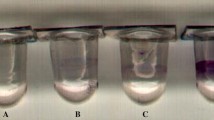
As shown in Fig. 4, the phenotypes associated with amrZ- and amrZ-fleQ- are substantially modified when increasing intracellular levels of c-di-GMP by overexpression of pleD*. We analyzed the colony morphology and CR-binding ability of amrZ- and amrZ-fleQ- on CR-supplemented plates in the absence or presence of pleD* (pJB3Tc19 and pJBpleD* respectively). As shown in Fig. 4a, amrZ- and amrZ-fleQ-, showed a light staining, a smooth patch, and an entire border. Conversely, the wild-type strain presented stronger staining, the patch had a rough texture, and its border was dented. In the wild-type, the overproduction of PleD* resulted in a rougher texture and slightly stronger staining. A similar phenotype was observed for all the mutants overexpressing pleD*: an increase in the color intensity, rougher texture, and dented borders. Furthermore, Fig. 4b shows that the defective biofilm formation phenotypes of the amrZ- and amrZ-fleQ- are reversed by increasing c-di-GMP intracellular levels, both after the initial attachment at 2 h and in a later development stage at 4 h. Indeed, not only attachment to solid surfaces was increased, but also cell–cell aggregates and pellicle formation (Supplementary Figures S1 and S2) were higher in the pleD* overexpressing strains. Finally, swimming motility analysis has shown the suppression of the hypermotile phenotype of the amrZ mutant caused by pleD* overexpression (Fig. 4c). Moreover, both F113-pJBpleD* and amrZ–pJBpleD* were non-motile after 24 h. As expected, amrZ-fleQ- is non-motile in the absence or presence of pleD*, due to the crucial role of FleQ in the synthesis of the flagellar apparatus in this bacterium4,6. Altogether, these results suggest that AmrZ regulates motility and biofilm formation mainly through the regulation of c-di-GMP levels.
c-di-GMP restores the biofilm and motility phenotypes in amrZ- and amrZ-fleQ-. (a) Morphology and Congo Red (CR)-binding pattern of Pseudomonas ogarae F113 and derivatives colonies carrying either the empty plasmid pJB3Tc19 or pJBpleD* grown for 48 h in Yeast-Mannitol-Broth (YMB) medium containing CR and Tc (zoom 1.25x). (b) Relative crystal violet-based biofilm formation assay of P. ogarae F113 and derivatives carrying pJB3Tc19 or pJBpleD* after 2 h and 4 h growth in microtiter plates with LB medium. The average and SD of four biological replicates with 16 technical replicates are represented; data was relativized to F113pJBTc19. (c) Swimming motility assay of P. ogarae F113 and derivatives carrying pJB3Tc19 or pJBpleD* at 24 h in SA medium containing Tc. Average and SD of three biological replicates with three technical replicates are shown. Images show swimming haloes formed after 24 h. Asterisks denote statistical significance of the data according to t-tests with Bonferroni-Dunn correction: *p value < 0.05; **p value < 0.01; ****p value < 0.0001.
Overproduction of c-di-GMP does not suppress the defect in rhizosphere competitive colonization in an amrZ mutant
Increased motility in P. ogarae F113 has been associated with enhanced rhizosphere competitive colonization ability11. However, despite amrZ- being a hypermotile mutant, it is dramatically impaired in competitive rhizosphere colonization assays7. Thus, a similar approach as previously described in this work was carried out to understand the role of c-di-GMP in the AmrZ regulation of the rhizosphere competitive colonization process. To avoid plasmid loss due to the lack of selection, alfalfa rhizosphere competitive colonization assays were performed using F113, the amrZ mutant, and the amrZ mutant harboring a mini-Tn7 based pleD* overexpression system (amrZ- pleD*). The overexpression system was tested for the expected swimming and biofilm formation phenotypes (Supplementary Figure S3). As shown in Fig. 5, amrZ- was totally displaced from the alfalfa rhizosphere by F113, consistent with our previous study7. Similarly, amrZ- pleD* was displaced entirely by F113, indicating that an increase in c-di-GMP levels was not enough to suppress the competitive colonization phenotype of the amrZ mutant. Furthermore, amrZ- pleD* is dramatically displaced even by the colonization-impaired amrZ mutant, suggesting that bacteria must tightly regulate c-di-GMP levels to be competitive during rizhosphere colonization.
c-di-GMP does not restore the defective rhizosphere competitive colonization phenotype of an amrZ mutant. Pseudomonas ogarae F113 wild-type, amrZ-, and the amrZ mutant containing a miniTn7-based pleD* integration were tested in competitive colonization assays. CFUs recovered from the alfalfa rhizosphere after seven days post-inoculation were tested for antibiotic resistance to distinguish the different strains. Percentages of colonies are represented. Each experiment was repeated twice with ten plants per experiment.
Discussion
Bacterial regulation of the switch between a planktonic lifestyle towards biofilm formation is an important trait for rhizosphere competence and persistence and, therefore, for the performance of rhizobacteria as plant growth promoters. When bacteria form biofilms, they are embedded in a self-produced ECM. The ECM components are subjected to fine control as they are needed not only in adhesion and immobilization of cells providing structure to biofilms but also play an essential role in signaling, genetic exchange, ion-sequestration, and migration37.
The second messenger c-di-GMP has been shown to play a crucial role in the transition from motile to sessile lifestyles10. We have previously shown that two pleiotropic regulatory proteins, AmrZ and FleQ, are involved in this transition in F1136. The involvement of c-di-GMP in this pathway is also evidenced by the regulation of DGC-coding gene expression exerted by AmrZ in this bacterium7, while FleQ is a c-di-GMP binding protein, whose activity depends on the binding of c-di-GMP as demonstrated in other pseudomonads38.
Here we have shown that in F113, AmrZ is a positive transcriptional regulator of genes related to ECM production, such as polysaccharides, Fap, extracellular adhesins, and the Flp/Tad pilus (Figs. 1 and 2) and that an increase in c-di-GMP levels in the amrZ mutant is enough to increase the expression to wild-type levels or higher (Fig. 2). Interestingly, only two of these genes, lapA and flp-1 were previously found as putative direct regulatory targets for AmrZ due to the presence of an AmrZ-binding site in their promoter region24. In fact, in the case of lapA, c-di-GMP levels do not affect at the transcription level, suggesting a direct regulation by AmrZ. Four of the putative ECM components, Pap, Fap, MapA and Flp/Tad pili show a common regulatory pattern. AmrZ appears to regulate all of them via c-di-GMP production. Interestingly, in all of them c-di-GMP effect is FleQ dependent, since PleD* overproduction had no effect in their transcription in the amrZ-fleQ double mutant (Fig. 2). It is interesting to note that we have previously identified FleQ-binding sites in the promoter regions of papA (PSF113_1970),), lapA (PSF113_0208), mapA (PSF113_1511) tadB (PSF113_4182), and rcpA (PSF113_4178)6. These results indicate complex AmrZ-FleQ-c-di-GMP interplay participating in the regulation of these ECM components. AmrZ and FleQ have already been described as regulators of the synthesis of different ECM components in other pseudomonads19,31,38,39,40. A role for FleQ has been also observed for lapA expression in P. putida KT24406. Conversely, we have not observed a role for FleQ in psmE transcription and the possible role of c-di-GMP on the regulation of pgaA appears independent of FleQ. Furthermore, in this work, we have shown that the expression of genes related to the production of ECM components responds to changes in the pool of c-di-GMP caused by mutation of distinct DGCs and PDEs that are regulated by AmrZ7 (Fig. 3). Genes dipA and PSF113_4681, encoding enzymes with both GGDEF and EAL domains appear to be the most important regulators for ECM components production. Mutants affected in either of these genes show an effect on the transcription of papA, pgaA and flp1, the dipA mutation affects mapA expression and the PSF113_4681 mutation has an effect on the transcription of alg8, lapA and psmE. In all cases, mutations in dipA resulted in an increased transcription, while mutations in PSF113_4681, resulted in a transcriptional decrease. Mutations affecting other genes encoding proteins with GGDEF domains (adrA, yfiN and PSF113_4827) and a HD-GYP domain (PSF113_3533) also affected the expression of several genes implicated in ECM components biosynthesis. In E. coli, PNAG production is also subjected to c-di-GMP-dependent transcriptional regulation via the c-di-GMP produced by the DGC YddV41. Similarly, c-di-GMP contribution to the alginate gene cluster expression has been previously observed in E. coli, in which the ectopic overexpression of the DGC YedQ led to increased alg8 and alg44 gene expression42. Although, to our knowledge, no previous data for the relation of c-di-GMP with the fap cluster in pseudomonads is available, regulation of their functional homologs in Salmonella sp. and E. coli, the amyloid curli fibers, is c-di-GMP dependent through the CsgD regulator43,44.
The results presented here have shown that modulation of c-di-GMP levels is the major pathway for the AmrZ regulation of biofilm formation and motility. In this regard, pleD* overexpression, leading to c-di-GMP overproduction, can suppress the biofilm-forming defects in the amrZ mutant, restoring attachment, Congo Red staining, and bacterial aggregation while suppressing the hypermotile phenotype (Fig. 4). However, the increase of c-di-GMP levels did not restore the rhizosphere colonization defect of the amrZ mutant (Fig. 5). We have previously shown that an amrZ mutant is totally displaced by the wild-type strain in competitive rhizosphere colonization experiments7. Here we show that the amrZ- strain overproducing c-di-GMP is also displaced from the rhizosphere by the wild-type strain and by the colonization-defective amrZ mutant. These results clearly show that rhizosphere competitive colonization phenotype of the amrZ mutant is not a consequence of c-di-GMP levels, in contrast to biofilm formation and motility in the amrZ mutant, and that high levels of c-di-GMP seem to be detrimental for competitive rhizosphere colonization.
Methods
Bacterial strains and growth conditions
Bacterial strains and plasmids used in this work are listed in Supplementary Table S2. Overnight cultures of Escherichia coli were routinely grown in Luria Bertani (LB) medium45 at 37 °C. Pseudomonas ogarae F113 and derivatives were grown in LB, Sucrose-Asparagine (SA)46, or Yeast-Mannitol Broth (YMB) pH 6.847 media, as indicated in each experiment, overnight at 28 °C with shaking. Purified agar (1.5% (w/v) for routine growth and Congo Red (CR) experiments or 0.3% (w/v) for swimming assays) or 1.5% (w/v) American Bacteriological Agar were used for solid medium. Antibiotics and dyes were added as required, at the following final concentrations: Rifampicin (Rif), 100 µg/mL, Cycloheximide (Chx), 10 μg/mL, Tetracycline (Tc), 70 μg/mL or 10 μg/mL for F113 or E. coli, respectively; Kanamycin (Km), 50 μg/mL for F113; Gentamicin (Gm), 10 μg/mL for E. coli or 5 μg/mL for F113; Chloramphenicol (Cm), 30 μg/mL and Ampicillin (Amp), 100 μg/mL for E. coli; and Congo Red (CR), 100 μg/mL.
Plasmid mobilization into F113 and derivatives was performed by triparental mating or electroporation. pJB3Tc1948 and pJBpleD*13 plasmids were introduced into F113 by triparental mating using E. coli JM109 as donor strain and using pRK60049. Similarly, miniTn7pleD*Tc50 was introduced in F113 by four-parental mating using pRK600 and pUX-BF1351.
Swimming motility
Motility of P. ogarae F113 and derivatives carrying either the empty vector pJB3Tc19 or the overproducing diguanylate cyclase PleD*: pJBpleD* or miniTn7pleD*Tc was tested in swimming motility assays. Strains were grown overnight in SA agar plates supplemented with Tc at 28 °C. Strains were inoculated in the center of SA agar plates (0.3%) supplemented with Tc. Swimming haloes diameters were measured after 24 h incubation at 28 °C. Experiments were performed at least in duplicate with three replicates in each experiment.
Congo red binding assay
pJB3Tc19 and pJBpleD* or miniTn7pleD*Tc-containing strains were grown overnight in YMB supplemented with Tc and 10 µL of culture were inoculated in triplicate in YMB agar plates supplemented with CR and Tc and incubated at 28 °C. Colony morphology and staining were recorded from day two to seven with a Leica MX125 stereoscope (zoom 1.25x). Every assay was performed three times.
Adherence and biofilm formation assays
Adherence and biofilm formation abilities were assayed following a procedure previously described52. Briefly, overnight cultures containing either pJB3Tc19 and pJBpleD* or miniTn7pleD*Tc grown in LB medium supplemented with Tc were adjusted to an optical density (OD)600 of 0.04 into fresh LB medium before inoculation of 100 µL into 96-well microtiter plates and incubated for 2 h or 4 h at 28 °C statically. Loosely attached cells and medium were removed (for 2 h experiments, attached cells were fixed with 99% (v/v) methanol for 15 min and air-dried). Cells were stained with 100 µL of 0.1% (w/v) crystal violet solution for 20 min. The excess stain was removed by rinsing with distilled water. Crystal violet staining was quantified by eluting with 150 µL of 33% (v/v) acetic acid for 10 min and measured at OD590 in a Synergy™ HT Multi-Mode Microplate reader (BioTek Instruments). Experiments were repeated at least twice with 16 technical replicates.
Competitive rhizosphere colonization assays
Alfalfa (Medicago sativa var. Resis) seeds were surface-sterilized with 70% (v/v) ethanol, 5% (v/v) sodium hypochlorite, washed gently with sterile distilled H2O, and germinated in H2O 1% (w/v) purified agar at 28 °C for 48 h. Seedlings were transferred and grown into 50 mL-tubes containing 25 mL of sterile pre-wetted medium-grain vermiculite and 10 mL Fåhraeus Plant (FP) medium53, in a controlled environment room (16/8 h, light/dark photoperiod and 25/18 ºC). Control and miniTn7pleD*Tc test strains were co-inoculated at 1·103 colony forming units (CFUs) at a 1:1 ratio to one-week-old seedlings. Pseudomonas ogarae F113 wild-type and the amrZ mutant were used as controls. After a week post-inoculation, shoots were removed, and rhizosphere bacteria were recovered by resuspension of soil in 20 mL of NaCl 0.75% (w/v) by vortex. Dilution series were plated onto SA agar plates supplemented with Rif and Chx and allowed to grow for 72 h at 28 °C. CFUs of each strain were plated onto fresh SA plates and distinguished by resistance to antibiotics (Km in the case of the amrZ mutant and KmTc in the amrZ mutant with miniTn7pleD*Tc integration). Assays were performed in duplicate with ten independent plants each time.
RNA isolation, cDNA synthesis, and qRT-PCR assay
Two independent cultures of F113 carrying the empty plasmid pJB3Tc19, amrZ-, amrZ-fleQ- containing pJB3Tc19 or pJBpleD* were grown in SA medium until stationary phase. Then, 1 mL of each culture was centrifuged for 5 min at 12,000 × g at RT. The supernatant was discarded and the cell pellet was resuspended in 100 µL of RNAlater® Solution (Ambion) before storage at 4 °C. RNA isolation, cDNA synthesis, and qRT-PCR assays were custom made by Plataforma de Genómica de la Fundación Parque Científico de Madrid (Madrid, Spain) with the primers listed in Supplementary Table S3. Gene expression was calculated using the Ct values. Data were normalized using rpoZ expression as housekeeping and relativized to F113 pJB3Tc19 following the 2−ΔΔCt method54.
RNA-Seq
RNA isolation, sequencing, and bioinformatic analysis from F113 and amrZ- cultures grown in SA medium at exponential (OD600 = 0.6) and stationary phase (OD600 = 1.2), or from rhizosphere colonization experiments were performed as described in32. F113 genes predicted to encode putative extracellular matrix components were selected. Differential gene expression was considered for values meeting a threshold of log2 Fold-Change (mutant/wild-type) ≥ 1 or ≤ -1 and a p-adjusted value ≤ 0.001. Heat-map was made using pheatmap R package version 1.0.1255.
Statistical analysis
GraphPad Prism version 7.00 for Windows (GraphPad Software, San Diego, California USA, www.graphpad.com) was used in the statistical analysis and representation of swimming motility, biofilm formation assays, and qRT-PCR data using multiple t-tests for independent samples with Bonferroni-Dunn method.
Data availability
The transcriptome sequences of Pseudomonas ogarae F113 and derivatives used in this study are available at the National Center for Biotechnology Information (NCBI) under the BioProject accession number PRJNA419480: P. ogarae F113 in exponential (SAMN17839758 and SAMN17839763) and stationary (SAMN17839759 and SAMN17839764) growth phases and under rhizosphere conditions (SAMN17839757 and SAMN17839762); amrZ mutant in exponential (SAMN26939580 and SAMN26939582) and stationary growth phases (SAMN26939581 and SAMN26939583), and under rhizosphere conditions (SAMN17839760 and SAMN17839765).
References
Garrido-Sanz, D., Redondo-Nieto, M., Martín, M. & Rivilla, R. Comparative genomics of the Pseudomonas corrugata subgroup reveals high species diversity and allows the description of Pseudomonas ogarae sp. nov. Microb. Genom. https://doi.org/10.1099/mgen.0.000593 (2021).
Shanahan, P., O’Sullivan, D. J., Simpson, P., Glennon, J. D. & O’Gara, F. Isolation of 2,4-diacetylphloroglucinol from a fluorescent pseudomonad and investigation of physiological parameters influencing its production. Appl. Environ. Microbiol. 58, 353–358. https://doi.org/10.1128/AEM.58.1.353-358.1992 (1992).
Sánchez-Contreras, M. et al. Phenotypic selection and phase variation occur during alfalfa root colonization by Pseudomonas fluorescens F113. J. Bacteriol. 184, 1587–1596 (2002).
Capdevila, S., Martínez-Granero, F. M., Sánchez-Contreras, M., Rivilla, R. & Martín, M. Analysis of Pseudomonas fluorescens F113 genes implicated in flagellar filament synthesis and their role in competitive root colonization. Microbiology (Reading) 150, 3889–3897. https://doi.org/10.1099/mic.0.27362-0 (2004).
Barahona, E. et al. Pseudomonas fluorescens F113 Can produce a second flagellar apparatus, which is important for plant root colonization. Front. Microbiol. 7, 1471. https://doi.org/10.3389/fmicb.2016.01471 (2016).
Blanco-Romero, E. et al. Genome-wide analysis of the FleQ direct regulon in Pseudomonas fluorescens F113 and Pseudomonas putida KT2440. Sci. Rep. 8, 13145. https://doi.org/10.1038/s41598-018-31371-z (2018).
Muriel, C. et al. AmrZ is a major determinant of c-di-GMP levels in Pseudomonas fluorescens F113. Sci. Rep. 8, 1979. https://doi.org/10.1038/s41598-018-20419-9 (2018).
Simm, R., Morr, M., Kader, A., Nimtz, M. & Romling, U. GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol. Microbiol. 53, 1123–1134. https://doi.org/10.1111/j.1365-2958.2004.04206.x (2004).
Ryan, R. P. et al. Cell-cell signaling in Xanthomonas campestris involves an HD-GYP domain protein that functions in cyclic di-GMP turnover. Proc. Natl. Acad. Sci. USA 103, 6712–6717. https://doi.org/10.1073/pnas.0600345103 (2006).
Hengge, R. Principles of c-di-GMP signalling in bacteria. Nat. Rev. Microbiol. 7, 263–273. https://doi.org/10.1038/nrmicro2109 (2009).
Barahona, E. et al. Pseudomonas fluorescens F113 mutant with enhanced competitive colonization ability and improved biocontrol activity against fungal root pathogens. Appl. Environ. Microbiol. 77, 5412–5419. https://doi.org/10.1128/AEM.00320-11 (2011).
Matilla, M. A., Travieso, M. L., Ramos, J. L. & Ramos-González, M. I. Cyclic diguanylate turnover mediated by the sole GGDEF/EAL response regulator in Pseudomonas putida: its role in the rhizosphere and an analysis of its target processes. Environ. Microbiol. 13, 1745–1766. https://doi.org/10.1111/j.1462-2920.2011.02499.x (2011).
Pérez-Mendoza, D. et al. Responses to elevated c-di-GMP levels in mutualistic and pathogenic plant-interacting bacteria. PLoS ONE 9, e91645. https://doi.org/10.1371/journal.pone.0091645 (2014).
Baynham, P. J. & Wozniak, D. J. Identification and characterization of AlgZ, an AlgT-dependent DNA-binding protein required for Pseudomonas aeruginosa algD transcription. Mol. Microbiol. 22, 97–108. https://doi.org/10.1111/j.1365-2958.1996.tb02659.x (1996).
Xu, B. & Wozniak, D. J. Development of a Novel Method for Analyzing Pseudomonas aeruginosa Twitching Motility and Its Application to Define the AmrZ Regulon. PLoS ONE 10, e0136426. https://doi.org/10.1371/journal.pone.0136426 (2015).
Baynham, P. J., Brown, A. L., Hall, L. L. & Wozniak, D. J. Pseudomonas aeruginosa AlgZ, a ribbon-helix-helix DNA-binding protein, is essential for alginate synthesis and algD transcriptional activation. Mol. Microbiol. 33, 1069–1080. https://doi.org/10.1046/j.1365-2958.1999.01550.x (1999).
Baynham, P. J., Ramsey, D. M., Gvozdyev, B. V., Cordonnier, E. M. & Wozniak, D. J. The Pseudomonas aeruginosa ribbon-helix-helix DNA-binding protein AlgZ (AmrZ) controls twitching motility and biogenesis of type IV pili. J. Bacteriol. 188, 132–140. https://doi.org/10.1128/JB.188.1.132-140.2006 (2006).
Tart, A. H., Blanks, M. J. & Wozniak, D. J. The AlgT-dependent transcriptional regulator AmrZ (AlgZ) inhibits flagellum biosynthesis in mucoid, nonmotile Pseudomonas aeruginosa cystic fibrosis isolates. J. Bacteriol. 188, 6483–6489. https://doi.org/10.1128/JB.00636-06 (2006).
Jones, C. J., Ryder, C. R., Mann, E. E. & Wozniak, D. J. AmrZ modulates Pseudomonas aeruginosa biofilm architecture by directly repressing transcription of the psl operon. J. Bacteriol. 195, 1637–1644. https://doi.org/10.1128/JB.02190-12 (2013).
Jones, C. J. et al. ChIP-Seq and RNA-Seq reveal an AmrZ-mediated mechanism for cyclic di-GMP synthesis and biofilm development by Pseudomonas aeruginosa. PLoS Pathog. 10, e1003984. https://doi.org/10.1371/journal.ppat.1003984 (2014).
Prada-Ramírez, H. A. et al. AmrZ regulates cellulose production in Pseudomonas syringae pv. tomato DC3000. Mol. Microbiol. 99, 960–977. https://doi.org/10.1111/mmi.13278 (2016).
Pryor, E. E. Jr. et al. The transcription factor AmrZ utilizes multiple DNA binding modes to recognize activator and repressor sequences of Pseudomonas aeruginosa virulence genes. PLoS Pathog. 8, e1002648. https://doi.org/10.1371/journal.ppat.1002648 (2012).
Baltrus, D. A., Dougherty, K., Díaz, B., Murillo, R. Evolutionary plasticity of AmrZ regulation in Pseudomonas. mSphere https://doi.org/10.1128/mSphere.00132-18 (2018).
Martínez-Granero, F., Redondo-Nieto, M., Vesga, P., Martín, M. & Rivilla, R. AmrZ is a global transcriptional regulator implicated in iron uptake and environmental adaption in P fluorescens F113. BMC Genomics 15, 237. https://doi.org/10.1186/1471-2164-15-237 (2014).
Tart, A. H., Wolfgang, M. C. & Wozniak, D. J. The alternative sigma factor AlgT represses Pseudomonas aeruginosa flagellum biosynthesis by inhibiting expression of fleQ. J. Bacteriol. 187, 7955–7962. https://doi.org/10.1128/JB.187.23.7955-7962.2005 (2005).
Martínez-Granero, F. et al. The Gac-Rsm and SadB signal transduction pathways converge on AlgU to downregulate motility in Pseudomonas fluorescens. PLoS ONE 7, e31765. https://doi.org/10.1371/journal.pone.0031765 (2012).
Baraquet, C., Murakami, K., Parsek, M. R. & Harwood, C. S. The FleQ protein from Pseudomonas aeruginosa functions as both a repressor and an activator to control gene expression from the pel operon promoter in response to c-di-GMP. Nucleic Acids Res. 40, 7207–7218. https://doi.org/10.1093/nar/gks384 (2012).
Hickman, J. W. & Harwood, C. S. Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol. Microbiol. 69, 376–389. https://doi.org/10.1111/j.1365-2958.2008.06281.x (2008).
Baraquet, C. & Harwood, C. S. Cyclic diguanosine monophosphate represses bacterial flagella synthesis by interacting with the Walker A motif of the enhancer-binding protein FleQ. Proc. Natl. Acad. Sci. USA 110, 18478–18483. https://doi.org/10.1073/pnas.1318972110 (2013).
Xiao, Y. et al. C-di-GMP regulates the expression of lapA and bcs operons via FleQ in Pseudomonas putida KT2440. Environ. Microbiol. Rep. 8, 659–666. https://doi.org/10.1111/1758-2229.12419 (2016).
Molina-Henares, M. A., Ramos-González, M. I., Daddaoua, A., Fernández-Escamilla, A. M. & Espinosa-Urgel, M. FleQ of Pseudomonas putida KT2440 is a multimeric cyclic diguanylate binding protein that differentially regulates expression of biofilm matrix components. Res. Microbiol. 168, 36–45. https://doi.org/10.1016/j.resmic.2016.07.005 (2017).
Blanco-Romero, E. et al. Transcriptomic analysis of Pseudomonas ogarae F113 reveals the antagonistic roles of AmrZ and FleQ during rhizosphere adaption. Microb. Genom. https://doi.org/10.1099/mgen.0.000750 (2022).
Payne, D. E. & Boles, B. R. Emerging interactions between matrix components during biofilm development. Curr. Genet. 62, 137–141. https://doi.org/10.1007/s00294-015-0527-5 (2016).
Blanco-Romero, E., Garrido-Sanz, D., Rivilla, R., Redondo-Nieto, M. & Martín, M. In silico characterization and phylogenetic distribution of extracellular matrix components in the model rhizobacteria Pseudomonas fluorescens F113 and Other Pseudomonads. Microorganisms 8, https://doi.org/10.3390/microorganisms8111740 (2020).
Aldridge, P., Paul, R., Goymer, P., Rainey, P. & Jenal, U. Role of the GGDEF regulator PleD in polar development of Caulobacter crescentus. Mol. Microbiol. 47, 1695–1708. https://doi.org/10.1046/j.1365-2958.2003.03401.x (2003).
Paul, R. et al. Cell cycle-dependent dynamic localization of a bacterial response regulator with a novel di-guanylate cyclase output domain. Genes Dev. 18, 715–727. https://doi.org/10.1101/gad.289504 (2004).
Dragos, A. & Kovacs, A. T. The peculiar functions of the bacterial extracellular matrix. Trends Microbiol. 25, 257–266. https://doi.org/10.1016/j.tim.2016.12.010 (2017).
Matsuyama, B. Y. et al. Mechanistic insights into c-di-GMP-dependent control of the biofilm regulator FleQ from Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 113, E209-218. https://doi.org/10.1073/pnas.1523148113 (2016).
Liu, H. et al. The exopolysaccharide gene cluster pea is transcriptionally controlled by RpoS and repressed by AmrZ in Pseudomonas putida KT2440. Microbiol. Res. 218, 1–11. https://doi.org/10.1016/j.micres.2018.09.004 (2019).
Pérez-Mendoza, D., Felipe, A., Ferreiro, M. D., Sanjuan, J. & Gallegos, M. T. AmrZ and FleQ Co-regulate cellulose production in Pseudomonas syringae pv. tomato DC3000. Front Microbiol 10, 746. https://doi.org/10.3389/fmicb.2019.00746 (2019).
Tagliabue, L. et al. The diguanylate cyclase YddV controls production of the exopolysaccharide poly-N-acetylglucosamine (PNAG) through regulation of the PNAG biosynthetic pgaABCD operon. Microbiology (Reading) 156, 2901–2911. https://doi.org/10.1099/mic.0.041350-0 (2010).
Wang, T. et al. Pleiotropic effects of c-di-GMP content in Pseudomonas syringae. Appl. Environ. Microbiol. https://doi.org/10.1128/AEM.00152-19 (2019).
Lindenberg, S., Klauck, G., Pesavento, C., Klauck, E. & Hengge, R. The EAL domain protein YciR acts as a trigger enzyme in a c-di-GMP signalling cascade in E. coli biofilm control. EMBO J. 32, 2001–2014. https://doi.org/10.1038/emboj.2013.120 (2013).
Zakikhany, K., Harrington, C. R., Nimtz, M., Hinton, J. C. & Romling, U. Unphosphorylated CsgD controls biofilm formation in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 77, 771–786. https://doi.org/10.1111/j.1365-2958.2010.07247.x (2010).
Bertani, G. Studies on lysogenesis I: the mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 62, 293–300 (1951).
Scher, F. M. & Baker, R. Effect of Pseudomonas putida and a synthetic iron chelator on induction of soil suppressiveness to Fusarium wilt pathogens. Phytopathology 72, 1567–1573 (1982).
Vincent, J. M. A manual for the practical study of the root-nodule bacteria. A manual for the practical study of the root-nodule bacteria. (1970).
Blatny, J. M., Brautaset, T., Winther-Larsen, H. C., Haugan, K. & Valla, S. Construction and use of a versatile set of broad-host-range cloning and expression vectors based on the RK2 replicon. Appl. Environ. Microbiol. 63, 370–379 (1997).
Finan, T. M., Kunkel, B., De Vos, G. F. & Signer, E. R. Second symbiotic megaplasmid in Rhizobium meliloti carrying exopolysaccharide and thiamine synthesis genes. J. Bacteriol. 167, 66–72 (1986).
Romero-Jiménez, L., Rodríguez-Carbonell, D., Gallegos, M. T., Sanjuán, J. & Pérez-Mendoza, D. Mini-Tn 7 vectors for stable expression of diguanylate cyclase PleD* in Gram-negative bacteria. BMC Microbiol. 15, 1–10 (2015).
Bao, Y., Lies, D. P., Fu, H. & Roberts, G. P. An improved Tn7-based system for the single-copy insertion of cloned genes into chromosomes of gram-negative bacteria. Gene 109, 167–168 (1991).
Peeters, E., Nelis, H. J. & Coenye, T. Comparison of multiple methods for quantification of microbial biofilms grown in microtiter plates. J. Microbiol. Methods 72, 157–165 (2008).
Fåhraeus, G. The infection of clover root hairs by nodule bacteria studied by a simple glass slide technique. Microbiology 16, 374–381 (1957).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25, 402–408 (2001).
Kolde, R. pheatmap: Pretty Heatmaps. R package version 1.0. 12. R Packag. version 1.0 8 (2019).
Acknowledgements
This work has been funded by Ministerio de Ciencia, Innovación y Universidades FEDER/EU Grant RTI2018-093991-B-I00. EB-R was granted by FPU-MECD program (FPU16/05513). D.G.-S. was funded by FPU-MECD fellowship program (FPU14/03965). The authors would like to thank Daniel Pérez Mendoza and Juan Sanjuán from EEZ-CSIC for the cession of PleD* constructs.
Author information
Authors and Affiliations
Contributions
M.M., M.R.-N and R.R. conceived the study; E.B.-R., D.G.-S. and D.D. designed and performed experiments; E.B.-R. and M.R.-N. performed the bioinformatic analysis; E.B.-R.; M.M., M.R.-N. and R.R. wrote the manuscript. All authors revised the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Blanco-Romero, E., Garrido-Sanz, D., Durán, D. et al. Regulation of extracellular matrix components by AmrZ is mediated by c-di-GMP in Pseudomonas ogarae F113. Sci Rep 12, 11914 (2022). https://doi.org/10.1038/s41598-022-16162-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16162-x
- Springer Nature Limited